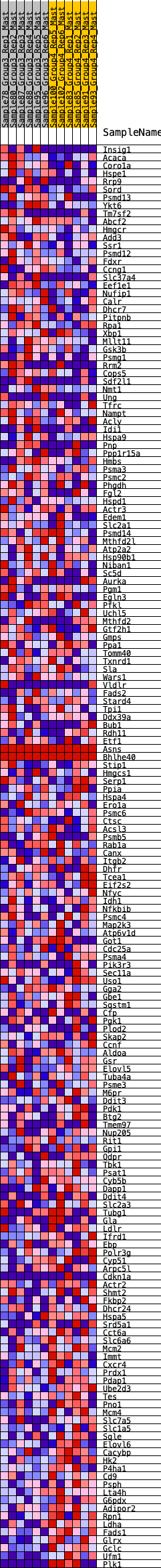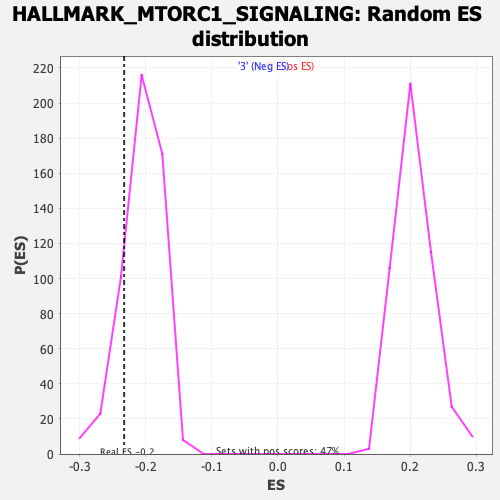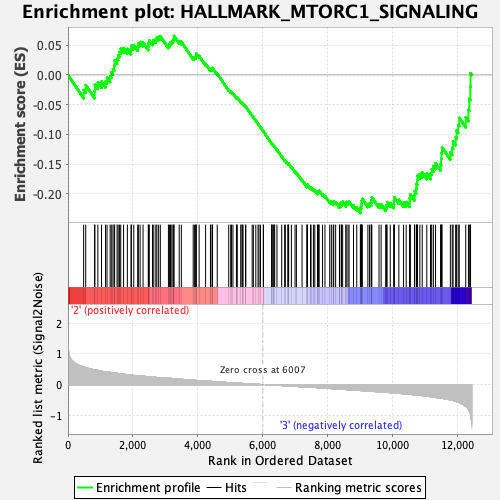
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Mast.Mast_Pheno.cls #Group3_versus_Group4.Mast_Pheno.cls #Group3_versus_Group4_repos |
| Phenotype | Mast_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_MTORC1_SIGNALING |
| Enrichment Score (ES) | -0.23262613 |
| Normalized Enrichment Score (NES) | -1.133904 |
| Nominal p-value | 0.15151516 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.988 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Insig1 | 482 | 0.577 | -0.0253 | No |
| 2 | Acaca | 546 | 0.550 | -0.0170 | No |
| 3 | Coro1a | 819 | 0.481 | -0.0275 | No |
| 4 | Hspe1 | 828 | 0.479 | -0.0164 | No |
| 5 | Rrp9 | 920 | 0.465 | -0.0125 | No |
| 6 | Sord | 1032 | 0.440 | -0.0109 | No |
| 7 | Psmd13 | 1157 | 0.418 | -0.0108 | No |
| 8 | Ykt6 | 1203 | 0.414 | -0.0044 | No |
| 9 | Tm7sf2 | 1298 | 0.400 | -0.0023 | No |
| 10 | Abcf2 | 1333 | 0.395 | 0.0046 | No |
| 11 | Hmgcr | 1384 | 0.387 | 0.0100 | No |
| 12 | Add3 | 1423 | 0.383 | 0.0162 | No |
| 13 | Ssr1 | 1436 | 0.381 | 0.0245 | No |
| 14 | Psmd12 | 1512 | 0.367 | 0.0273 | No |
| 15 | Fdxr | 1545 | 0.361 | 0.0335 | No |
| 16 | Ccng1 | 1589 | 0.354 | 0.0386 | No |
| 17 | Slc37a4 | 1625 | 0.350 | 0.0443 | No |
| 18 | Eef1e1 | 1712 | 0.340 | 0.0456 | No |
| 19 | Nufip1 | 1829 | 0.324 | 0.0440 | No |
| 20 | Calr | 1943 | 0.311 | 0.0424 | No |
| 21 | Dhcr7 | 1952 | 0.309 | 0.0492 | No |
| 22 | Pitpnb | 2026 | 0.299 | 0.0505 | No |
| 23 | Rpa1 | 2148 | 0.286 | 0.0477 | No |
| 24 | Xbp1 | 2167 | 0.284 | 0.0531 | No |
| 25 | Mllt11 | 2221 | 0.279 | 0.0556 | No |
| 26 | Gsk3b | 2312 | 0.272 | 0.0549 | No |
| 27 | Psmg1 | 2473 | 0.259 | 0.0481 | No |
| 28 | Rrm2 | 2480 | 0.259 | 0.0540 | No |
| 29 | Cops5 | 2505 | 0.256 | 0.0582 | No |
| 30 | Sdf2l1 | 2603 | 0.246 | 0.0563 | No |
| 31 | Nmt1 | 2643 | 0.242 | 0.0590 | No |
| 32 | Ung | 2706 | 0.235 | 0.0597 | No |
| 33 | Tfrc | 2738 | 0.233 | 0.0629 | No |
| 34 | Nampt | 2792 | 0.228 | 0.0641 | No |
| 35 | Acly | 2844 | 0.226 | 0.0655 | No |
| 36 | Idi1 | 3094 | 0.208 | 0.0502 | No |
| 37 | Hspa9 | 3129 | 0.206 | 0.0525 | No |
| 38 | Pnp | 3157 | 0.203 | 0.0552 | No |
| 39 | Ppp1r15a | 3199 | 0.199 | 0.0567 | No |
| 40 | Hmbs | 3236 | 0.195 | 0.0585 | No |
| 41 | Psma3 | 3259 | 0.194 | 0.0615 | No |
| 42 | Psmc2 | 3264 | 0.193 | 0.0658 | No |
| 43 | Phgdh | 3426 | 0.180 | 0.0571 | No |
| 44 | Fgl2 | 3492 | 0.173 | 0.0560 | No |
| 45 | Hspd1 | 3857 | 0.145 | 0.0299 | No |
| 46 | Actr3 | 3900 | 0.142 | 0.0299 | No |
| 47 | Edem1 | 3935 | 0.139 | 0.0305 | No |
| 48 | Slc2a1 | 3945 | 0.138 | 0.0332 | No |
| 49 | Psmd14 | 3950 | 0.138 | 0.0362 | No |
| 50 | Mthfd2l | 4039 | 0.131 | 0.0322 | No |
| 51 | Atp2a2 | 4239 | 0.116 | 0.0188 | No |
| 52 | Hsp90b1 | 4388 | 0.107 | 0.0093 | No |
| 53 | Niban1 | 4406 | 0.105 | 0.0105 | No |
| 54 | Sc5d | 4451 | 0.102 | 0.0094 | No |
| 55 | Aurka | 4453 | 0.102 | 0.0118 | No |
| 56 | Pgm1 | 4598 | 0.091 | 0.0023 | No |
| 57 | Egln3 | 4952 | 0.067 | -0.0249 | No |
| 58 | Pfkl | 5013 | 0.062 | -0.0282 | No |
| 59 | Uchl5 | 5039 | 0.061 | -0.0288 | No |
| 60 | Mthfd2 | 5082 | 0.058 | -0.0308 | No |
| 61 | Gtf2h1 | 5189 | 0.050 | -0.0382 | No |
| 62 | Gmps | 5203 | 0.049 | -0.0381 | No |
| 63 | Ppa1 | 5211 | 0.049 | -0.0375 | No |
| 64 | Tomm40 | 5321 | 0.042 | -0.0453 | No |
| 65 | Txnrd1 | 5364 | 0.039 | -0.0478 | No |
| 66 | Sla | 5367 | 0.039 | -0.0470 | No |
| 67 | Wars1 | 5393 | 0.037 | -0.0481 | No |
| 68 | Vldlr | 5470 | 0.033 | -0.0535 | No |
| 69 | Fads2 | 5478 | 0.033 | -0.0533 | No |
| 70 | Stard4 | 5674 | 0.020 | -0.0687 | No |
| 71 | Tpi1 | 5710 | 0.018 | -0.0711 | No |
| 72 | Ddx39a | 5786 | 0.014 | -0.0769 | No |
| 73 | Bub1 | 5855 | 0.010 | -0.0822 | No |
| 74 | Rdh11 | 5885 | 0.007 | -0.0844 | No |
| 75 | Etf1 | 5931 | 0.005 | -0.0879 | No |
| 76 | Asns | 6019 | 0.000 | -0.0950 | No |
| 77 | Bhlhe40 | 6020 | 0.000 | -0.0950 | No |
| 78 | Stip1 | 6274 | -0.011 | -0.1154 | No |
| 79 | Hmgcs1 | 6278 | -0.011 | -0.1154 | No |
| 80 | Serp1 | 6293 | -0.012 | -0.1162 | No |
| 81 | Ppia | 6341 | -0.016 | -0.1197 | No |
| 82 | Hspa4 | 6345 | -0.016 | -0.1195 | No |
| 83 | Ero1a | 6367 | -0.018 | -0.1208 | No |
| 84 | Psmc6 | 6438 | -0.023 | -0.1260 | No |
| 85 | Ctsc | 6584 | -0.030 | -0.1371 | No |
| 86 | Acsl3 | 6675 | -0.037 | -0.1435 | No |
| 87 | Psmb5 | 6695 | -0.038 | -0.1441 | No |
| 88 | Rab1a | 6773 | -0.042 | -0.1494 | No |
| 89 | Canx | 6788 | -0.043 | -0.1495 | No |
| 90 | Itgb2 | 6798 | -0.044 | -0.1491 | No |
| 91 | Dhfr | 6882 | -0.049 | -0.1547 | No |
| 92 | Tcea1 | 6995 | -0.056 | -0.1625 | No |
| 93 | Eif2s2 | 7037 | -0.058 | -0.1644 | No |
| 94 | Nfyc | 7211 | -0.069 | -0.1768 | No |
| 95 | Idh1 | 7362 | -0.078 | -0.1872 | No |
| 96 | Nfkbib | 7364 | -0.078 | -0.1854 | No |
| 97 | Psmc4 | 7378 | -0.079 | -0.1845 | No |
| 98 | Map2k3 | 7471 | -0.084 | -0.1900 | No |
| 99 | Atp6v1d | 7488 | -0.085 | -0.1892 | No |
| 100 | Got1 | 7561 | -0.090 | -0.1929 | No |
| 101 | Cdc25a | 7603 | -0.093 | -0.1939 | No |
| 102 | Psma4 | 7687 | -0.100 | -0.1983 | No |
| 103 | Pik3r3 | 7702 | -0.100 | -0.1970 | No |
| 104 | Sec11a | 7715 | -0.101 | -0.1955 | No |
| 105 | Uso1 | 7738 | -0.103 | -0.1948 | No |
| 106 | Gga2 | 7842 | -0.111 | -0.2005 | No |
| 107 | Gbe1 | 7917 | -0.116 | -0.2037 | No |
| 108 | Sqstm1 | 8064 | -0.126 | -0.2125 | No |
| 109 | Cfp | 8114 | -0.129 | -0.2134 | No |
| 110 | Pgk1 | 8171 | -0.132 | -0.2147 | No |
| 111 | Plod2 | 8187 | -0.133 | -0.2127 | No |
| 112 | Skap2 | 8246 | -0.137 | -0.2141 | No |
| 113 | Ccnf | 8363 | -0.145 | -0.2200 | No |
| 114 | Aldoa | 8369 | -0.146 | -0.2169 | No |
| 115 | Gsr | 8421 | -0.149 | -0.2174 | No |
| 116 | Elovl5 | 8427 | -0.149 | -0.2142 | No |
| 117 | Tuba4a | 8462 | -0.152 | -0.2133 | No |
| 118 | Psme3 | 8564 | -0.160 | -0.2176 | No |
| 119 | M6pr | 8565 | -0.160 | -0.2137 | No |
| 120 | Ddit3 | 8626 | -0.165 | -0.2146 | No |
| 121 | Pdk1 | 8659 | -0.168 | -0.2131 | No |
| 122 | Btg2 | 8796 | -0.177 | -0.2199 | No |
| 123 | Tmem97 | 8896 | -0.184 | -0.2235 | No |
| 124 | Nup205 | 9009 | -0.192 | -0.2279 | Yes |
| 125 | Rit1 | 9010 | -0.193 | -0.2232 | Yes |
| 126 | Gpi1 | 9034 | -0.194 | -0.2204 | Yes |
| 127 | Qdpr | 9046 | -0.195 | -0.2165 | Yes |
| 128 | Tbk1 | 9050 | -0.195 | -0.2120 | Yes |
| 129 | Psat1 | 9066 | -0.197 | -0.2084 | Yes |
| 130 | Cyb5b | 9239 | -0.210 | -0.2174 | Yes |
| 131 | Dapp1 | 9291 | -0.214 | -0.2163 | Yes |
| 132 | Ddit4 | 9340 | -0.217 | -0.2149 | Yes |
| 133 | Slc2a3 | 9350 | -0.217 | -0.2104 | Yes |
| 134 | Tubg1 | 9360 | -0.218 | -0.2058 | Yes |
| 135 | Gla | 9582 | -0.233 | -0.2181 | Yes |
| 136 | Ldlr | 9647 | -0.237 | -0.2176 | Yes |
| 137 | Ifrd1 | 9788 | -0.248 | -0.2229 | Yes |
| 138 | Ebp | 9815 | -0.251 | -0.2190 | Yes |
| 139 | Polr3g | 9829 | -0.252 | -0.2139 | Yes |
| 140 | Cyp51 | 9928 | -0.261 | -0.2155 | Yes |
| 141 | Arpc5l | 10037 | -0.270 | -0.2177 | Yes |
| 142 | Cdkn1a | 10046 | -0.270 | -0.2118 | Yes |
| 143 | Actr2 | 10057 | -0.271 | -0.2060 | Yes |
| 144 | Shmt2 | 10193 | -0.282 | -0.2101 | Yes |
| 145 | Fkbp2 | 10338 | -0.296 | -0.2147 | Yes |
| 146 | Dhcr24 | 10419 | -0.302 | -0.2138 | Yes |
| 147 | Hspa5 | 10524 | -0.313 | -0.2147 | Yes |
| 148 | Srd5a1 | 10528 | -0.313 | -0.2073 | Yes |
| 149 | Cct6a | 10547 | -0.316 | -0.2011 | Yes |
| 150 | Slc6a6 | 10675 | -0.331 | -0.2034 | Yes |
| 151 | Mcm2 | 10677 | -0.331 | -0.1954 | Yes |
| 152 | Immt | 10730 | -0.337 | -0.1914 | Yes |
| 153 | Cxcr4 | 10737 | -0.338 | -0.1837 | Yes |
| 154 | Prdx1 | 10758 | -0.341 | -0.1770 | Yes |
| 155 | Pdap1 | 10768 | -0.342 | -0.1694 | Yes |
| 156 | Ube2d3 | 10840 | -0.350 | -0.1667 | Yes |
| 157 | Tes | 10913 | -0.357 | -0.1638 | Yes |
| 158 | Pno1 | 11054 | -0.376 | -0.1661 | Yes |
| 159 | Mcm4 | 11168 | -0.392 | -0.1657 | Yes |
| 160 | Slc7a5 | 11204 | -0.396 | -0.1589 | Yes |
| 161 | Slc1a5 | 11256 | -0.405 | -0.1532 | Yes |
| 162 | Sqle | 11324 | -0.416 | -0.1486 | Yes |
| 163 | Elovl6 | 11478 | -0.436 | -0.1504 | Yes |
| 164 | Cacybp | 11497 | -0.440 | -0.1412 | Yes |
| 165 | Hk2 | 11507 | -0.442 | -0.1311 | Yes |
| 166 | P4ha1 | 11525 | -0.446 | -0.1217 | Yes |
| 167 | Cd9 | 11780 | -0.495 | -0.1303 | Yes |
| 168 | Psph | 11837 | -0.507 | -0.1225 | Yes |
| 169 | Lta4h | 11864 | -0.518 | -0.1120 | Yes |
| 170 | G6pdx | 11943 | -0.547 | -0.1051 | Yes |
| 171 | Adipor2 | 11972 | -0.556 | -0.0938 | Yes |
| 172 | Rpn1 | 12022 | -0.572 | -0.0838 | Yes |
| 173 | Ldha | 12054 | -0.585 | -0.0721 | Yes |
| 174 | Fads1 | 12254 | -0.700 | -0.0713 | Yes |
| 175 | Glrx | 12337 | -0.792 | -0.0587 | Yes |
| 176 | Gclc | 12363 | -0.852 | -0.0399 | Yes |
| 177 | Ufm1 | 12395 | -0.930 | -0.0198 | Yes |
| 178 | Plk1 | 12402 | -0.964 | 0.0032 | Yes |