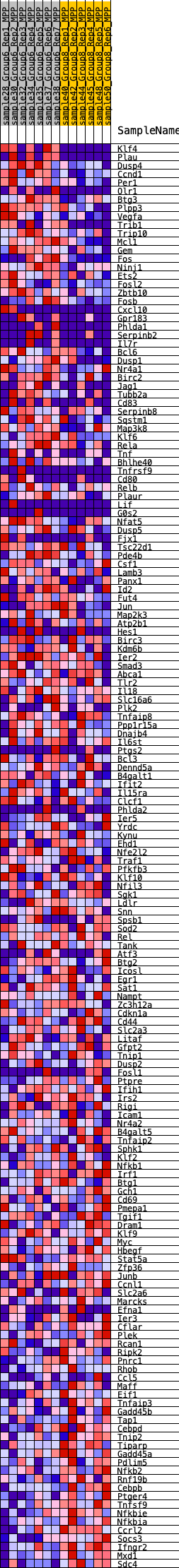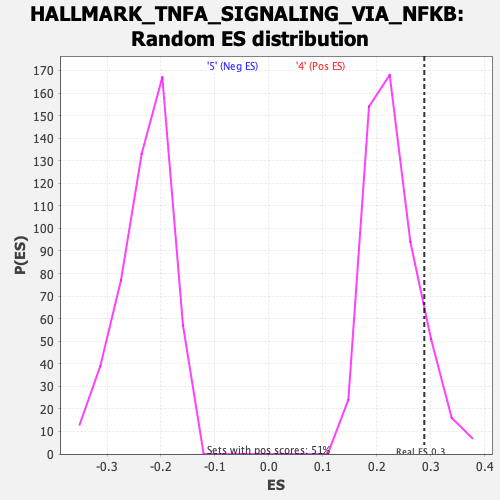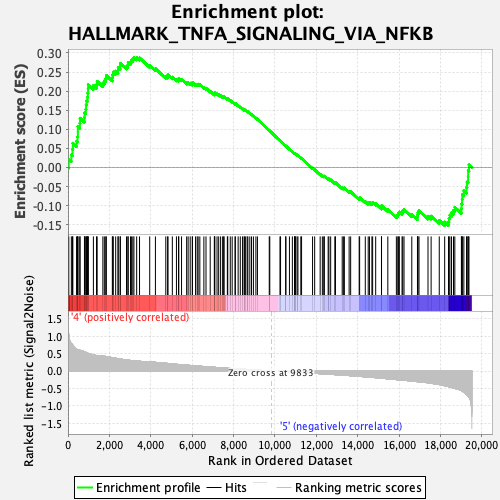
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group6_versus_Group8.MPP_Pheno.cls#Group6_versus_Group8_repos |
| Phenotype | MPP_Pheno.cls#Group6_versus_Group8_repos |
| Upregulated in class | 4 |
| GeneSet | HALLMARK_TNFA_SIGNALING_VIA_NFKB |
| Enrichment Score (ES) | 0.28832778 |
| Normalized Enrichment Score (NES) | 1.2546749 |
| Nominal p-value | 0.12840466 |
| FDR q-value | 0.54804367 |
| FWER p-Value | 0.867 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Klf4 | 38 | 1.039 | 0.0218 | Yes |
| 2 | Plau | 162 | 0.784 | 0.0333 | Yes |
| 3 | Dusp4 | 218 | 0.747 | 0.0475 | Yes |
| 4 | Ccnd1 | 234 | 0.740 | 0.0637 | Yes |
| 5 | Per1 | 413 | 0.625 | 0.0688 | Yes |
| 6 | Olr1 | 460 | 0.611 | 0.0803 | Yes |
| 7 | Btg3 | 476 | 0.609 | 0.0935 | Yes |
| 8 | Plpp3 | 477 | 0.609 | 0.1074 | Yes |
| 9 | Vegfa | 572 | 0.594 | 0.1161 | Yes |
| 10 | Trib1 | 581 | 0.594 | 0.1292 | Yes |
| 11 | Trip10 | 795 | 0.555 | 0.1309 | Yes |
| 12 | Mcl1 | 800 | 0.553 | 0.1433 | Yes |
| 13 | Gem | 860 | 0.535 | 0.1525 | Yes |
| 14 | Fos | 874 | 0.530 | 0.1639 | Yes |
| 15 | Ninj1 | 896 | 0.524 | 0.1748 | Yes |
| 16 | Ets2 | 940 | 0.512 | 0.1842 | Yes |
| 17 | Fosl2 | 956 | 0.509 | 0.1951 | Yes |
| 18 | Zbtb10 | 970 | 0.505 | 0.2060 | Yes |
| 19 | Fosb | 975 | 0.504 | 0.2173 | Yes |
| 20 | Cxcl10 | 1227 | 0.464 | 0.2149 | Yes |
| 21 | Gpr183 | 1378 | 0.443 | 0.2173 | Yes |
| 22 | Phlda1 | 1408 | 0.440 | 0.2258 | Yes |
| 23 | Serpinb2 | 1685 | 0.423 | 0.2212 | Yes |
| 24 | Il7r | 1756 | 0.421 | 0.2272 | Yes |
| 25 | Bcl6 | 1823 | 0.416 | 0.2333 | Yes |
| 26 | Dusp1 | 1853 | 0.414 | 0.2412 | Yes |
| 27 | Nr4a1 | 2142 | 0.384 | 0.2351 | Yes |
| 28 | Birc2 | 2160 | 0.381 | 0.2429 | Yes |
| 29 | Jag1 | 2187 | 0.379 | 0.2503 | Yes |
| 30 | Tubb2a | 2304 | 0.365 | 0.2526 | Yes |
| 31 | Cd83 | 2418 | 0.354 | 0.2548 | Yes |
| 32 | Serpinb8 | 2425 | 0.353 | 0.2626 | Yes |
| 33 | Sqstm1 | 2521 | 0.345 | 0.2656 | Yes |
| 34 | Map3k8 | 2528 | 0.345 | 0.2731 | Yes |
| 35 | Klf6 | 2827 | 0.318 | 0.2650 | Yes |
| 36 | Rela | 2894 | 0.312 | 0.2687 | Yes |
| 37 | Tnf | 2908 | 0.312 | 0.2751 | Yes |
| 38 | Bhlhe40 | 3020 | 0.303 | 0.2763 | Yes |
| 39 | Tnfrsf9 | 3063 | 0.301 | 0.2810 | Yes |
| 40 | Cd80 | 3131 | 0.297 | 0.2844 | Yes |
| 41 | Relb | 3185 | 0.294 | 0.2883 | Yes |
| 42 | Plaur | 3318 | 0.288 | 0.2881 | No |
| 43 | Lif | 3456 | 0.277 | 0.2873 | No |
| 44 | G0s2 | 3949 | 0.252 | 0.2677 | No |
| 45 | Nfat5 | 4224 | 0.249 | 0.2592 | No |
| 46 | Dusp5 | 4725 | 0.223 | 0.2384 | No |
| 47 | Fjx1 | 4815 | 0.218 | 0.2388 | No |
| 48 | Tsc22d1 | 4830 | 0.217 | 0.2430 | No |
| 49 | Pde4b | 5041 | 0.204 | 0.2368 | No |
| 50 | Csf1 | 5240 | 0.193 | 0.2310 | No |
| 51 | Lamb3 | 5340 | 0.187 | 0.2302 | No |
| 52 | Panx1 | 5354 | 0.186 | 0.2338 | No |
| 53 | Id2 | 5486 | 0.180 | 0.2311 | No |
| 54 | Fut4 | 5733 | 0.169 | 0.2222 | No |
| 55 | Jun | 5804 | 0.164 | 0.2224 | No |
| 56 | Map2k3 | 5904 | 0.160 | 0.2209 | No |
| 57 | Atp2b1 | 6008 | 0.156 | 0.2192 | No |
| 58 | Hes1 | 6010 | 0.156 | 0.2227 | No |
| 59 | Birc3 | 6164 | 0.147 | 0.2181 | No |
| 60 | Kdm6b | 6242 | 0.144 | 0.2174 | No |
| 61 | Ier2 | 6287 | 0.142 | 0.2184 | No |
| 62 | Smad3 | 6370 | 0.138 | 0.2173 | No |
| 63 | Abca1 | 6565 | 0.129 | 0.2102 | No |
| 64 | Tlr2 | 6668 | 0.123 | 0.2077 | No |
| 65 | Il18 | 6871 | 0.115 | 0.1999 | No |
| 66 | Slc16a6 | 7070 | 0.107 | 0.1921 | No |
| 67 | Plk2 | 7080 | 0.106 | 0.1941 | No |
| 68 | Tnfaip8 | 7096 | 0.105 | 0.1957 | No |
| 69 | Ppp1r15a | 7201 | 0.101 | 0.1926 | No |
| 70 | Dnajb4 | 7281 | 0.097 | 0.1908 | No |
| 71 | Il6st | 7374 | 0.093 | 0.1882 | No |
| 72 | Ptgs2 | 7473 | 0.089 | 0.1851 | No |
| 73 | Bcl3 | 7506 | 0.088 | 0.1855 | No |
| 74 | Dennd5a | 7545 | 0.086 | 0.1855 | No |
| 75 | B4galt1 | 7700 | 0.081 | 0.1794 | No |
| 76 | Ifit2 | 7719 | 0.080 | 0.1803 | No |
| 77 | Il15ra | 7841 | 0.075 | 0.1757 | No |
| 78 | Clcf1 | 7933 | 0.072 | 0.1727 | No |
| 79 | Phlda2 | 8073 | 0.066 | 0.1670 | No |
| 80 | Ier5 | 8084 | 0.065 | 0.1680 | No |
| 81 | Yrdc | 8221 | 0.060 | 0.1623 | No |
| 82 | Kynu | 8325 | 0.057 | 0.1583 | No |
| 83 | Ehd1 | 8434 | 0.053 | 0.1539 | No |
| 84 | Nfe2l2 | 8513 | 0.050 | 0.1510 | No |
| 85 | Traf1 | 8521 | 0.050 | 0.1518 | No |
| 86 | Pfkfb3 | 8583 | 0.047 | 0.1497 | No |
| 87 | Klf10 | 8659 | 0.044 | 0.1468 | No |
| 88 | Nfil3 | 8691 | 0.043 | 0.1462 | No |
| 89 | Sgk1 | 8785 | 0.040 | 0.1423 | No |
| 90 | Ldlr | 8868 | 0.036 | 0.1389 | No |
| 91 | Snn | 8959 | 0.033 | 0.1350 | No |
| 92 | Spsb1 | 9079 | 0.029 | 0.1295 | No |
| 93 | Sod2 | 9153 | 0.026 | 0.1263 | No |
| 94 | Rel | 9724 | 0.004 | 0.0969 | No |
| 95 | Tank | 9750 | 0.003 | 0.0957 | No |
| 96 | Atf3 | 10243 | -0.003 | 0.0703 | No |
| 97 | Btg2 | 10271 | -0.003 | 0.0690 | No |
| 98 | Icosl | 10516 | -0.011 | 0.0566 | No |
| 99 | Egr1 | 10530 | -0.011 | 0.0562 | No |
| 100 | Sat1 | 10700 | -0.016 | 0.0478 | No |
| 101 | Nampt | 10841 | -0.021 | 0.0411 | No |
| 102 | Zc3h12a | 10946 | -0.026 | 0.0363 | No |
| 103 | Cdkn1a | 10951 | -0.026 | 0.0367 | No |
| 104 | Cd44 | 10970 | -0.027 | 0.0364 | No |
| 105 | Slc2a3 | 10997 | -0.028 | 0.0357 | No |
| 106 | Litaf | 11068 | -0.031 | 0.0328 | No |
| 107 | Gfpt2 | 11110 | -0.032 | 0.0314 | No |
| 108 | Tnip1 | 11250 | -0.037 | 0.0251 | No |
| 109 | Dusp2 | 11286 | -0.039 | 0.0241 | No |
| 110 | Fosl1 | 11817 | -0.058 | -0.0019 | No |
| 111 | Ptpre | 11919 | -0.062 | -0.0057 | No |
| 112 | Ifih1 | 12184 | -0.073 | -0.0177 | No |
| 113 | Irs2 | 12290 | -0.077 | -0.0214 | No |
| 114 | Rigi | 12358 | -0.078 | -0.0231 | No |
| 115 | Icam1 | 12387 | -0.079 | -0.0227 | No |
| 116 | Nr4a2 | 12571 | -0.087 | -0.0302 | No |
| 117 | B4galt5 | 12607 | -0.088 | -0.0300 | No |
| 118 | Tnfaip2 | 12692 | -0.091 | -0.0323 | No |
| 119 | Sphk1 | 12894 | -0.100 | -0.0404 | No |
| 120 | Klf2 | 12924 | -0.101 | -0.0395 | No |
| 121 | Nfkb1 | 13249 | -0.113 | -0.0537 | No |
| 122 | Irf1 | 13306 | -0.116 | -0.0540 | No |
| 123 | Btg1 | 13354 | -0.117 | -0.0537 | No |
| 124 | Gch1 | 13584 | -0.128 | -0.0626 | No |
| 125 | Cd69 | 13658 | -0.131 | -0.0634 | No |
| 126 | Pmepa1 | 14069 | -0.148 | -0.0812 | No |
| 127 | Tgif1 | 14092 | -0.150 | -0.0789 | No |
| 128 | Dram1 | 14365 | -0.163 | -0.0893 | No |
| 129 | Klf9 | 14506 | -0.170 | -0.0926 | No |
| 130 | Myc | 14566 | -0.173 | -0.0917 | No |
| 131 | Hbegf | 14685 | -0.178 | -0.0938 | No |
| 132 | Stat5a | 14720 | -0.179 | -0.0914 | No |
| 133 | Zfp36 | 14867 | -0.188 | -0.0947 | No |
| 134 | Junb | 15147 | -0.203 | -0.1045 | No |
| 135 | Ccnl1 | 15150 | -0.204 | -0.0999 | No |
| 136 | Slc2a6 | 15454 | -0.219 | -0.1106 | No |
| 137 | Marcks | 15867 | -0.243 | -0.1263 | No |
| 138 | Efna1 | 15936 | -0.247 | -0.1242 | No |
| 139 | Ier3 | 15965 | -0.249 | -0.1200 | No |
| 140 | Cflar | 16011 | -0.251 | -0.1165 | No |
| 141 | Plek | 16141 | -0.260 | -0.1173 | No |
| 142 | Rcan1 | 16175 | -0.262 | -0.1130 | No |
| 143 | Ripk2 | 16236 | -0.266 | -0.1100 | No |
| 144 | Pnrc1 | 16612 | -0.287 | -0.1229 | No |
| 145 | Rhob | 16886 | -0.302 | -0.1301 | No |
| 146 | Ccl5 | 16891 | -0.303 | -0.1234 | No |
| 147 | Maff | 16910 | -0.305 | -0.1173 | No |
| 148 | Eif1 | 16960 | -0.308 | -0.1128 | No |
| 149 | Tnfaip3 | 17395 | -0.334 | -0.1276 | No |
| 150 | Gadd45b | 17546 | -0.350 | -0.1274 | No |
| 151 | Tap1 | 17939 | -0.385 | -0.1388 | No |
| 152 | Cebpd | 18201 | -0.418 | -0.1428 | No |
| 153 | Tnip2 | 18385 | -0.445 | -0.1421 | No |
| 154 | Tiparp | 18409 | -0.448 | -0.1330 | No |
| 155 | Gadd45a | 18452 | -0.455 | -0.1248 | No |
| 156 | Pdlim5 | 18532 | -0.468 | -0.1182 | No |
| 157 | Nfkb2 | 18628 | -0.480 | -0.1122 | No |
| 158 | Rnf19b | 18691 | -0.492 | -0.1041 | No |
| 159 | Cebpb | 19008 | -0.557 | -0.1077 | No |
| 160 | Ptger4 | 19017 | -0.561 | -0.0953 | No |
| 161 | Tnfsf9 | 19044 | -0.569 | -0.0837 | No |
| 162 | Nfkbie | 19053 | -0.575 | -0.0710 | No |
| 163 | Nfkbia | 19128 | -0.601 | -0.0610 | No |
| 164 | Ccrl2 | 19255 | -0.670 | -0.0523 | No |
| 165 | Socs3 | 19281 | -0.685 | -0.0379 | No |
| 166 | Ifngr2 | 19334 | -0.715 | -0.0243 | No |
| 167 | Mxd1 | 19337 | -0.715 | -0.0081 | No |
| 168 | Sdc4 | 19375 | -0.746 | 0.0071 | No |