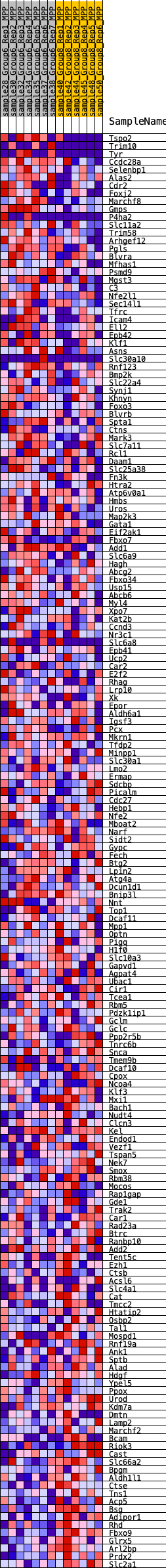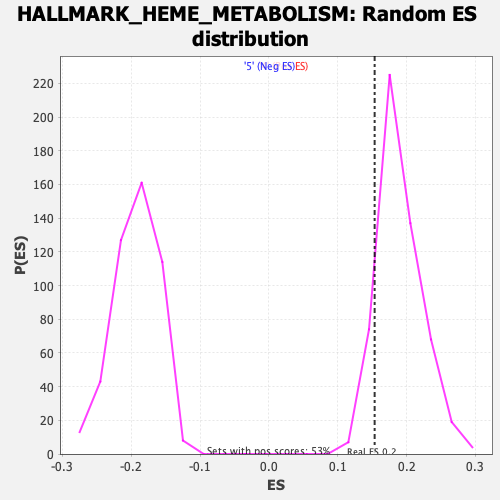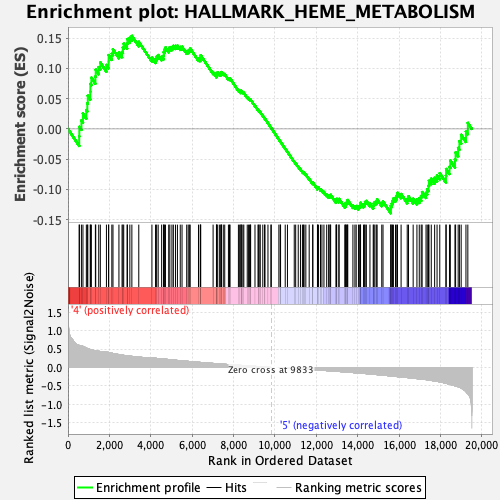
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group6_versus_Group8.MPP_Pheno.cls#Group6_versus_Group8_repos |
| Phenotype | MPP_Pheno.cls#Group6_versus_Group8_repos |
| Upregulated in class | 4 |
| GeneSet | HALLMARK_HEME_METABOLISM |
| Enrichment Score (ES) | 0.15364711 |
| Normalized Enrichment Score (NES) | 0.80816865 |
| Nominal p-value | 0.90636706 |
| FDR q-value | 0.92215514 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Tspo2 | 542 | 0.598 | -0.0121 | Yes |
| 2 | Trim10 | 548 | 0.597 | 0.0036 | Yes |
| 3 | Tyr | 648 | 0.584 | 0.0141 | Yes |
| 4 | Ccdc28a | 722 | 0.570 | 0.0255 | Yes |
| 5 | Selenbp1 | 886 | 0.527 | 0.0311 | Yes |
| 6 | Alas2 | 934 | 0.513 | 0.0424 | Yes |
| 7 | Cdr2 | 957 | 0.509 | 0.0548 | Yes |
| 8 | Foxj2 | 1078 | 0.485 | 0.0616 | Yes |
| 9 | Marchf8 | 1082 | 0.483 | 0.0743 | Yes |
| 10 | Gmps | 1129 | 0.477 | 0.0847 | Yes |
| 11 | P4ha2 | 1323 | 0.451 | 0.0867 | Yes |
| 12 | Slc11a2 | 1340 | 0.449 | 0.0979 | Yes |
| 13 | Trim58 | 1485 | 0.437 | 0.1021 | Yes |
| 14 | Arhgef12 | 1562 | 0.434 | 0.1097 | Yes |
| 15 | Pgls | 1857 | 0.414 | 0.1056 | Yes |
| 16 | Blvra | 1963 | 0.407 | 0.1110 | Yes |
| 17 | Mfhas1 | 1964 | 0.407 | 0.1219 | Yes |
| 18 | Psmd9 | 2118 | 0.386 | 0.1242 | Yes |
| 19 | Mgst3 | 2173 | 0.379 | 0.1316 | Yes |
| 20 | C3 | 2461 | 0.350 | 0.1261 | Yes |
| 21 | Nfe2l1 | 2612 | 0.335 | 0.1273 | Yes |
| 22 | Sec14l1 | 2651 | 0.331 | 0.1341 | Yes |
| 23 | Tfrc | 2692 | 0.328 | 0.1408 | Yes |
| 24 | Icam4 | 2855 | 0.316 | 0.1409 | Yes |
| 25 | Ell2 | 2872 | 0.314 | 0.1484 | Yes |
| 26 | Epb42 | 2988 | 0.306 | 0.1506 | Yes |
| 27 | Klf1 | 3086 | 0.300 | 0.1536 | Yes |
| 28 | Asns | 3420 | 0.279 | 0.1439 | No |
| 29 | Slc30a10 | 4055 | 0.251 | 0.1178 | No |
| 30 | Rnf123 | 4230 | 0.248 | 0.1154 | No |
| 31 | Bmp2k | 4282 | 0.245 | 0.1193 | No |
| 32 | Slc22a4 | 4357 | 0.241 | 0.1219 | No |
| 33 | Synj1 | 4525 | 0.235 | 0.1195 | No |
| 34 | Khnyn | 4620 | 0.229 | 0.1208 | No |
| 35 | Foxo3 | 4625 | 0.229 | 0.1267 | No |
| 36 | Blvrb | 4658 | 0.227 | 0.1311 | No |
| 37 | Spta1 | 4709 | 0.224 | 0.1345 | No |
| 38 | Ctns | 4867 | 0.215 | 0.1321 | No |
| 39 | Mark3 | 4921 | 0.212 | 0.1350 | No |
| 40 | Slc7a11 | 5022 | 0.206 | 0.1353 | No |
| 41 | Rcl1 | 5090 | 0.202 | 0.1373 | No |
| 42 | Daam1 | 5193 | 0.196 | 0.1372 | No |
| 43 | Slc25a38 | 5287 | 0.190 | 0.1375 | No |
| 44 | Fn3k | 5424 | 0.183 | 0.1353 | No |
| 45 | Htra2 | 5510 | 0.179 | 0.1357 | No |
| 46 | Atp6v0a1 | 5736 | 0.168 | 0.1286 | No |
| 47 | Hmbs | 5808 | 0.164 | 0.1293 | No |
| 48 | Uros | 5863 | 0.162 | 0.1308 | No |
| 49 | Map2k3 | 5904 | 0.160 | 0.1330 | No |
| 50 | Gata1 | 6305 | 0.141 | 0.1161 | No |
| 51 | Eif2ak1 | 6388 | 0.137 | 0.1155 | No |
| 52 | Fbxo7 | 6404 | 0.136 | 0.1184 | No |
| 53 | Add1 | 6412 | 0.136 | 0.1216 | No |
| 54 | Slc6a9 | 7011 | 0.109 | 0.0936 | No |
| 55 | Hagh | 7173 | 0.102 | 0.0880 | No |
| 56 | Abcg2 | 7182 | 0.102 | 0.0903 | No |
| 57 | Fbxo34 | 7190 | 0.101 | 0.0926 | No |
| 58 | Usp15 | 7235 | 0.099 | 0.0930 | No |
| 59 | Abcb6 | 7328 | 0.095 | 0.0908 | No |
| 60 | Myl4 | 7380 | 0.093 | 0.0906 | No |
| 61 | Xpo7 | 7410 | 0.092 | 0.0916 | No |
| 62 | Kat2b | 7414 | 0.091 | 0.0938 | No |
| 63 | Ccnd3 | 7503 | 0.088 | 0.0916 | No |
| 64 | Nr3c1 | 7573 | 0.085 | 0.0904 | No |
| 65 | Slc6a8 | 7752 | 0.079 | 0.0833 | No |
| 66 | Epb41 | 7803 | 0.077 | 0.0827 | No |
| 67 | Ucp2 | 7830 | 0.076 | 0.0834 | No |
| 68 | Car2 | 8234 | 0.060 | 0.0641 | No |
| 69 | E2f2 | 8295 | 0.057 | 0.0626 | No |
| 70 | Rhag | 8360 | 0.055 | 0.0607 | No |
| 71 | Lrp10 | 8364 | 0.055 | 0.0620 | No |
| 72 | Xk | 8367 | 0.055 | 0.0634 | No |
| 73 | Epor | 8449 | 0.053 | 0.0606 | No |
| 74 | Aldh6a1 | 8495 | 0.051 | 0.0597 | No |
| 75 | Igsf3 | 8661 | 0.044 | 0.0523 | No |
| 76 | Pcx | 8726 | 0.042 | 0.0501 | No |
| 77 | Mkrn1 | 8774 | 0.040 | 0.0487 | No |
| 78 | Tfdp2 | 8817 | 0.038 | 0.0476 | No |
| 79 | Minpp1 | 8825 | 0.038 | 0.0482 | No |
| 80 | Slc30a1 | 9035 | 0.030 | 0.0382 | No |
| 81 | Lmo2 | 9185 | 0.024 | 0.0311 | No |
| 82 | Ermap | 9198 | 0.024 | 0.0312 | No |
| 83 | Sdcbp | 9277 | 0.020 | 0.0277 | No |
| 84 | Picalm | 9279 | 0.020 | 0.0282 | No |
| 85 | Cdc27 | 9412 | 0.016 | 0.0217 | No |
| 86 | Hebp1 | 9498 | 0.013 | 0.0177 | No |
| 87 | Nfe2 | 9517 | 0.012 | 0.0171 | No |
| 88 | Mboat2 | 9662 | 0.006 | 0.0098 | No |
| 89 | Narf | 9805 | 0.001 | 0.0024 | No |
| 90 | Sidt2 | 9822 | 0.000 | 0.0016 | No |
| 91 | Gypc | 10190 | -0.000 | -0.0174 | No |
| 92 | Fech | 10256 | -0.003 | -0.0206 | No |
| 93 | Btg2 | 10271 | -0.003 | -0.0213 | No |
| 94 | Lpin2 | 10496 | -0.010 | -0.0326 | No |
| 95 | Atg4a | 10604 | -0.013 | -0.0378 | No |
| 96 | Dcun1d1 | 10932 | -0.026 | -0.0540 | No |
| 97 | Bnip3l | 11001 | -0.028 | -0.0568 | No |
| 98 | Nnt | 11125 | -0.033 | -0.0623 | No |
| 99 | Top1 | 11232 | -0.037 | -0.0668 | No |
| 100 | Dcaf11 | 11327 | -0.040 | -0.0706 | No |
| 101 | Mpp1 | 11376 | -0.042 | -0.0719 | No |
| 102 | Optn | 11408 | -0.044 | -0.0724 | No |
| 103 | Pigq | 11492 | -0.046 | -0.0754 | No |
| 104 | H1f0 | 11654 | -0.052 | -0.0824 | No |
| 105 | Slc10a3 | 11823 | -0.058 | -0.0895 | No |
| 106 | Gapvd1 | 11831 | -0.058 | -0.0883 | No |
| 107 | Agpat4 | 12053 | -0.068 | -0.0979 | No |
| 108 | Ubac1 | 12056 | -0.068 | -0.0962 | No |
| 109 | Cir1 | 12106 | -0.070 | -0.0969 | No |
| 110 | Tcea1 | 12201 | -0.074 | -0.0997 | No |
| 111 | Rbm5 | 12270 | -0.076 | -0.1012 | No |
| 112 | Pdzk1ip1 | 12367 | -0.079 | -0.1041 | No |
| 113 | Gclm | 12500 | -0.084 | -0.1087 | No |
| 114 | Gclc | 12597 | -0.088 | -0.1113 | No |
| 115 | Ppp2r5b | 12614 | -0.088 | -0.1098 | No |
| 116 | Tnrc6b | 12674 | -0.091 | -0.1104 | No |
| 117 | Snca | 12691 | -0.091 | -0.1088 | No |
| 118 | Tmem9b | 12947 | -0.103 | -0.1193 | No |
| 119 | Dcaf10 | 12973 | -0.103 | -0.1178 | No |
| 120 | Cpox | 12983 | -0.104 | -0.1155 | No |
| 121 | Ncoa4 | 13090 | -0.107 | -0.1181 | No |
| 122 | Klf3 | 13096 | -0.108 | -0.1155 | No |
| 123 | Mxi1 | 13374 | -0.118 | -0.1267 | No |
| 124 | Bach1 | 13416 | -0.120 | -0.1256 | No |
| 125 | Nudt4 | 13419 | -0.120 | -0.1225 | No |
| 126 | Clcn3 | 13462 | -0.122 | -0.1214 | No |
| 127 | Kel | 13511 | -0.124 | -0.1206 | No |
| 128 | Endod1 | 13514 | -0.124 | -0.1174 | No |
| 129 | Vezf1 | 13767 | -0.136 | -0.1268 | No |
| 130 | Tspan5 | 13864 | -0.140 | -0.1280 | No |
| 131 | Nek7 | 13925 | -0.142 | -0.1273 | No |
| 132 | Smox | 14035 | -0.147 | -0.1290 | No |
| 133 | Rbm38 | 14077 | -0.149 | -0.1272 | No |
| 134 | Mocos | 14128 | -0.152 | -0.1257 | No |
| 135 | Rap1gap | 14136 | -0.152 | -0.1220 | No |
| 136 | Gde1 | 14280 | -0.159 | -0.1252 | No |
| 137 | Trak2 | 14334 | -0.162 | -0.1236 | No |
| 138 | Car1 | 14356 | -0.163 | -0.1203 | No |
| 139 | Rad23a | 14416 | -0.165 | -0.1190 | No |
| 140 | Btrc | 14590 | -0.174 | -0.1233 | No |
| 141 | Ranbp10 | 14748 | -0.181 | -0.1266 | No |
| 142 | Add2 | 14774 | -0.182 | -0.1230 | No |
| 143 | Tent5c | 14826 | -0.185 | -0.1207 | No |
| 144 | Ezh1 | 14897 | -0.190 | -0.1193 | No |
| 145 | Ctsb | 14937 | -0.192 | -0.1162 | No |
| 146 | Acsl6 | 15161 | -0.204 | -0.1223 | No |
| 147 | Slc4a1 | 15221 | -0.207 | -0.1198 | No |
| 148 | Cat | 15595 | -0.227 | -0.1330 | No |
| 149 | Tmcc2 | 15598 | -0.227 | -0.1271 | No |
| 150 | Htatip2 | 15644 | -0.229 | -0.1233 | No |
| 151 | Osbp2 | 15678 | -0.232 | -0.1188 | No |
| 152 | Tal1 | 15724 | -0.234 | -0.1149 | No |
| 153 | Mospd1 | 15824 | -0.240 | -0.1136 | No |
| 154 | Rnf19a | 15886 | -0.244 | -0.1102 | No |
| 155 | Ank1 | 15922 | -0.246 | -0.1054 | No |
| 156 | Sptb | 16095 | -0.257 | -0.1075 | No |
| 157 | Alad | 16391 | -0.271 | -0.1155 | No |
| 158 | Hdgf | 16452 | -0.275 | -0.1113 | No |
| 159 | Ypel5 | 16678 | -0.290 | -0.1152 | No |
| 160 | Ppox | 16860 | -0.300 | -0.1165 | No |
| 161 | Urod | 16973 | -0.309 | -0.1141 | No |
| 162 | Kdm7a | 17066 | -0.312 | -0.1105 | No |
| 163 | Dmtn | 17111 | -0.316 | -0.1043 | No |
| 164 | Lamp2 | 17300 | -0.327 | -0.1053 | No |
| 165 | Marchf2 | 17360 | -0.331 | -0.0996 | No |
| 166 | Bcam | 17421 | -0.337 | -0.0937 | No |
| 167 | Riok3 | 17441 | -0.339 | -0.0856 | No |
| 168 | Cast | 17553 | -0.351 | -0.0820 | No |
| 169 | Slc66a2 | 17716 | -0.362 | -0.0807 | No |
| 170 | Bpgm | 17833 | -0.372 | -0.0768 | No |
| 171 | Aldh1l1 | 17964 | -0.388 | -0.0732 | No |
| 172 | Ctse | 18262 | -0.426 | -0.0772 | No |
| 173 | Tns1 | 18272 | -0.427 | -0.0662 | No |
| 174 | Acp5 | 18433 | -0.452 | -0.0624 | No |
| 175 | Bsg | 18474 | -0.460 | -0.0522 | No |
| 176 | Adipor1 | 18698 | -0.493 | -0.0506 | No |
| 177 | Rhd | 18729 | -0.500 | -0.0388 | No |
| 178 | Fbxo9 | 18853 | -0.521 | -0.0313 | No |
| 179 | Glrx5 | 18913 | -0.534 | -0.0201 | No |
| 180 | Arl2bp | 18998 | -0.553 | -0.0097 | No |
| 181 | Prdx2 | 19229 | -0.650 | -0.0042 | No |
| 182 | Slc2a1 | 19320 | -0.705 | 0.0099 | No |