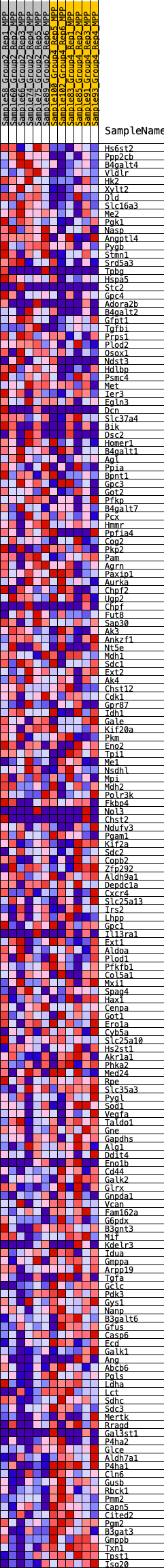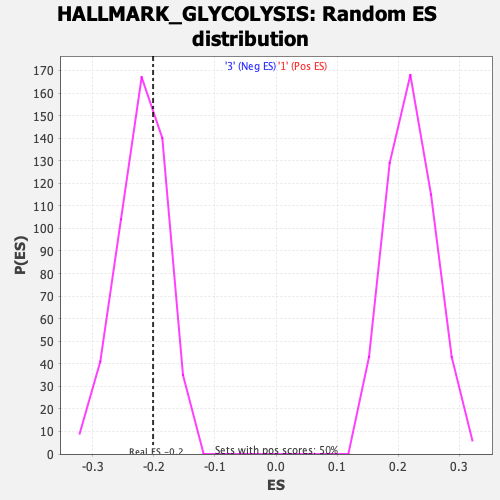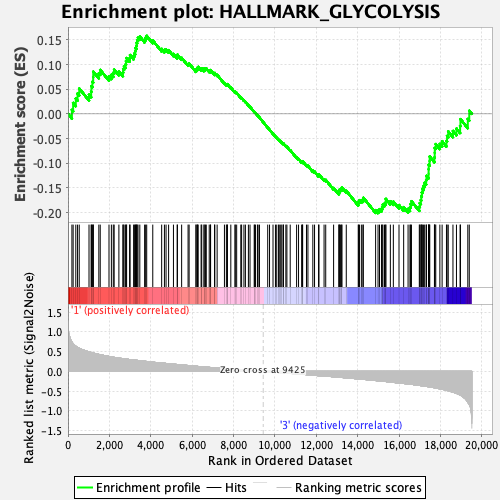
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group2_versus_Group4.MPP_Pheno.cls#Group2_versus_Group4_repos |
| Phenotype | MPP_Pheno.cls#Group2_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_GLYCOLYSIS |
| Enrichment Score (ES) | -0.20118442 |
| Normalized Enrichment Score (NES) | -0.9141648 |
| Nominal p-value | 0.671371 |
| FDR q-value | 0.92950076 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Hs6st2 | 189 | 0.741 | 0.0082 | No |
| 2 | Ppp2cb | 248 | 0.697 | 0.0221 | No |
| 3 | B4galt4 | 377 | 0.636 | 0.0309 | No |
| 4 | Vldlr | 462 | 0.607 | 0.0413 | No |
| 5 | Hk2 | 543 | 0.583 | 0.0514 | No |
| 6 | Xylt2 | 1015 | 0.493 | 0.0390 | No |
| 7 | Dld | 1119 | 0.479 | 0.0453 | No |
| 8 | Slc16a3 | 1132 | 0.476 | 0.0562 | No |
| 9 | Me2 | 1179 | 0.469 | 0.0652 | No |
| 10 | Pgk1 | 1218 | 0.464 | 0.0745 | No |
| 11 | Nasp | 1221 | 0.463 | 0.0856 | No |
| 12 | Angptl4 | 1490 | 0.429 | 0.0822 | No |
| 13 | Pygb | 1557 | 0.422 | 0.0890 | No |
| 14 | Stmn1 | 1981 | 0.379 | 0.0763 | No |
| 15 | Srd5a3 | 2096 | 0.367 | 0.0793 | No |
| 16 | Tpbg | 2187 | 0.358 | 0.0834 | No |
| 17 | Hspa5 | 2230 | 0.355 | 0.0898 | No |
| 18 | Stc2 | 2459 | 0.336 | 0.0862 | No |
| 19 | Gpc4 | 2654 | 0.322 | 0.0840 | No |
| 20 | Adora2b | 2667 | 0.321 | 0.0911 | No |
| 21 | B4galt2 | 2709 | 0.318 | 0.0967 | No |
| 22 | Gfpt1 | 2785 | 0.312 | 0.1004 | No |
| 23 | Tgfbi | 2804 | 0.311 | 0.1070 | No |
| 24 | Prps1 | 2832 | 0.309 | 0.1131 | No |
| 25 | Plod2 | 2978 | 0.301 | 0.1129 | No |
| 26 | Qsox1 | 2999 | 0.300 | 0.1192 | No |
| 27 | Ndst3 | 3171 | 0.289 | 0.1173 | No |
| 28 | Hdlbp | 3207 | 0.286 | 0.1225 | No |
| 29 | Psmc4 | 3249 | 0.284 | 0.1272 | No |
| 30 | Met | 3263 | 0.283 | 0.1334 | No |
| 31 | Ier3 | 3305 | 0.281 | 0.1381 | No |
| 32 | Egln3 | 3320 | 0.279 | 0.1442 | No |
| 33 | Dcn | 3341 | 0.278 | 0.1499 | No |
| 34 | Slc37a4 | 3382 | 0.277 | 0.1545 | No |
| 35 | Bik | 3470 | 0.272 | 0.1566 | No |
| 36 | Dsc2 | 3702 | 0.257 | 0.1509 | No |
| 37 | Homer1 | 3752 | 0.254 | 0.1546 | No |
| 38 | B4galt1 | 3803 | 0.251 | 0.1581 | No |
| 39 | Agl | 4096 | 0.233 | 0.1486 | No |
| 40 | Ppia | 4526 | 0.212 | 0.1316 | No |
| 41 | Bpnt1 | 4659 | 0.205 | 0.1297 | No |
| 42 | Gpc3 | 4738 | 0.201 | 0.1306 | No |
| 43 | Got2 | 4861 | 0.194 | 0.1290 | No |
| 44 | Pfkp | 5095 | 0.184 | 0.1214 | No |
| 45 | B4galt7 | 5276 | 0.174 | 0.1163 | No |
| 46 | Pcx | 5287 | 0.173 | 0.1200 | No |
| 47 | Hmmr | 5491 | 0.163 | 0.1135 | No |
| 48 | Ppfia4 | 5802 | 0.148 | 0.1010 | No |
| 49 | Cog2 | 5852 | 0.146 | 0.1020 | No |
| 50 | Pkp2 | 6182 | 0.131 | 0.0882 | No |
| 51 | Pam | 6192 | 0.130 | 0.0909 | No |
| 52 | Agrn | 6248 | 0.127 | 0.0911 | No |
| 53 | Paxip1 | 6259 | 0.127 | 0.0937 | No |
| 54 | Aurka | 6290 | 0.125 | 0.0952 | No |
| 55 | Chpf2 | 6431 | 0.121 | 0.0909 | No |
| 56 | Ugp2 | 6462 | 0.119 | 0.0922 | No |
| 57 | Chpf | 6575 | 0.115 | 0.0892 | No |
| 58 | Fut8 | 6576 | 0.115 | 0.0920 | No |
| 59 | Sap30 | 6647 | 0.112 | 0.0911 | No |
| 60 | Ak3 | 6680 | 0.109 | 0.0921 | No |
| 61 | Ankzf1 | 6824 | 0.102 | 0.0872 | No |
| 62 | Nt5e | 6887 | 0.099 | 0.0864 | No |
| 63 | Mdh1 | 6889 | 0.099 | 0.0887 | No |
| 64 | Sdc1 | 7084 | 0.090 | 0.0809 | No |
| 65 | Ext2 | 7090 | 0.090 | 0.0828 | No |
| 66 | Ak4 | 7206 | 0.085 | 0.0789 | No |
| 67 | Chst12 | 7559 | 0.073 | 0.0625 | No |
| 68 | Cdk1 | 7652 | 0.070 | 0.0594 | No |
| 69 | Gpr87 | 7692 | 0.068 | 0.0591 | No |
| 70 | Idh1 | 7702 | 0.067 | 0.0602 | No |
| 71 | Gale | 7867 | 0.061 | 0.0532 | No |
| 72 | Kif20a | 8069 | 0.052 | 0.0441 | No |
| 73 | Pkm | 8081 | 0.052 | 0.0448 | No |
| 74 | Eno2 | 8150 | 0.049 | 0.0425 | No |
| 75 | Tpi1 | 8350 | 0.041 | 0.0332 | No |
| 76 | Me1 | 8389 | 0.040 | 0.0322 | No |
| 77 | Nsdhl | 8501 | 0.035 | 0.0273 | No |
| 78 | Mpi | 8572 | 0.031 | 0.0244 | No |
| 79 | Mdh2 | 8717 | 0.026 | 0.0176 | No |
| 80 | Polr3k | 8795 | 0.022 | 0.0142 | No |
| 81 | Fkbp4 | 8997 | 0.015 | 0.0041 | No |
| 82 | Nol3 | 9017 | 0.014 | 0.0035 | No |
| 83 | Chst2 | 9062 | 0.013 | 0.0015 | No |
| 84 | Ndufv3 | 9154 | 0.010 | -0.0029 | No |
| 85 | Pgam1 | 9185 | 0.009 | -0.0043 | No |
| 86 | Kif2a | 9252 | 0.007 | -0.0075 | No |
| 87 | Sdc2 | 9650 | -0.006 | -0.0279 | No |
| 88 | Copb2 | 9738 | -0.010 | -0.0321 | No |
| 89 | Zfp292 | 9906 | -0.017 | -0.0404 | No |
| 90 | Aldh9a1 | 10024 | -0.022 | -0.0459 | No |
| 91 | Depdc1a | 10086 | -0.023 | -0.0485 | No |
| 92 | Cxcr4 | 10175 | -0.027 | -0.0524 | No |
| 93 | Slc25a13 | 10207 | -0.029 | -0.0533 | No |
| 94 | Irs2 | 10278 | -0.032 | -0.0561 | No |
| 95 | Lhpp | 10331 | -0.034 | -0.0580 | No |
| 96 | Gpc1 | 10397 | -0.037 | -0.0605 | No |
| 97 | Il13ra1 | 10411 | -0.037 | -0.0602 | No |
| 98 | Ext1 | 10527 | -0.042 | -0.0652 | No |
| 99 | Aldoa | 10577 | -0.043 | -0.0666 | No |
| 100 | Plod1 | 10737 | -0.049 | -0.0737 | No |
| 101 | Pfkfb1 | 11046 | -0.062 | -0.0881 | No |
| 102 | Col5a1 | 11140 | -0.066 | -0.0913 | No |
| 103 | Mxi1 | 11289 | -0.071 | -0.0973 | No |
| 104 | Spag4 | 11334 | -0.073 | -0.0978 | No |
| 105 | Hax1 | 11336 | -0.073 | -0.0961 | No |
| 106 | Cenpa | 11523 | -0.080 | -0.1037 | No |
| 107 | Got1 | 11593 | -0.083 | -0.1053 | No |
| 108 | Ero1a | 11830 | -0.095 | -0.1152 | No |
| 109 | Cyb5a | 11913 | -0.098 | -0.1171 | No |
| 110 | Slc25a10 | 12108 | -0.106 | -0.1245 | No |
| 111 | Hs2st1 | 12126 | -0.107 | -0.1228 | No |
| 112 | Akr1a1 | 12373 | -0.117 | -0.1327 | No |
| 113 | Phka2 | 12453 | -0.120 | -0.1338 | No |
| 114 | Med24 | 12834 | -0.134 | -0.1502 | No |
| 115 | Rpe | 13089 | -0.145 | -0.1598 | No |
| 116 | Slc35a3 | 13105 | -0.146 | -0.1571 | No |
| 117 | Pygl | 13109 | -0.146 | -0.1537 | No |
| 118 | Sod1 | 13171 | -0.149 | -0.1532 | No |
| 119 | Vegfa | 13223 | -0.151 | -0.1522 | No |
| 120 | Taldo1 | 13235 | -0.152 | -0.1491 | No |
| 121 | Gne | 13446 | -0.163 | -0.1560 | No |
| 122 | Gapdhs | 14024 | -0.188 | -0.1813 | No |
| 123 | Alg1 | 14041 | -0.189 | -0.1775 | No |
| 124 | Ddit4 | 14083 | -0.191 | -0.1750 | No |
| 125 | Eno1b | 14175 | -0.195 | -0.1750 | No |
| 126 | Cd44 | 14250 | -0.199 | -0.1740 | No |
| 127 | Galk2 | 14269 | -0.200 | -0.1700 | No |
| 128 | Glrx | 14864 | -0.229 | -0.1952 | No |
| 129 | Gnpda1 | 14981 | -0.236 | -0.1955 | Yes |
| 130 | Vcan | 15051 | -0.240 | -0.1932 | Yes |
| 131 | Fam162a | 15147 | -0.244 | -0.1922 | Yes |
| 132 | G6pdx | 15183 | -0.246 | -0.1881 | Yes |
| 133 | B3gnt3 | 15216 | -0.248 | -0.1837 | Yes |
| 134 | Mif | 15293 | -0.252 | -0.1815 | Yes |
| 135 | Kdelr3 | 15354 | -0.255 | -0.1784 | Yes |
| 136 | Idua | 15356 | -0.255 | -0.1723 | Yes |
| 137 | Gmppa | 15576 | -0.268 | -0.1771 | Yes |
| 138 | Arpp19 | 15720 | -0.276 | -0.1778 | Yes |
| 139 | Tgfa | 15994 | -0.292 | -0.1849 | Yes |
| 140 | Gclc | 16219 | -0.303 | -0.1891 | Yes |
| 141 | Pdk3 | 16432 | -0.316 | -0.1924 | Yes |
| 142 | Gys1 | 16524 | -0.321 | -0.1893 | Yes |
| 143 | Nanp | 16554 | -0.323 | -0.1829 | Yes |
| 144 | B3galt6 | 16595 | -0.326 | -0.1771 | Yes |
| 145 | Gfus | 16975 | -0.353 | -0.1881 | Yes |
| 146 | Casp6 | 17000 | -0.355 | -0.1808 | Yes |
| 147 | Ecd | 17050 | -0.358 | -0.1746 | Yes |
| 148 | Galk1 | 17073 | -0.360 | -0.1670 | Yes |
| 149 | Ang | 17096 | -0.361 | -0.1594 | Yes |
| 150 | Abcb6 | 17122 | -0.363 | -0.1519 | Yes |
| 151 | Pgls | 17181 | -0.367 | -0.1459 | Yes |
| 152 | Ldha | 17234 | -0.371 | -0.1396 | Yes |
| 153 | Lct | 17307 | -0.377 | -0.1342 | Yes |
| 154 | Sdhc | 17320 | -0.377 | -0.1257 | Yes |
| 155 | Sdc3 | 17424 | -0.386 | -0.1216 | Yes |
| 156 | Mertk | 17425 | -0.386 | -0.1123 | Yes |
| 157 | Rragd | 17430 | -0.386 | -0.1031 | Yes |
| 158 | Gal3st1 | 17473 | -0.391 | -0.0958 | Yes |
| 159 | P4ha2 | 17476 | -0.391 | -0.0864 | Yes |
| 160 | Glce | 17706 | -0.411 | -0.0883 | Yes |
| 161 | Aldh7a1 | 17720 | -0.413 | -0.0789 | Yes |
| 162 | P4ha1 | 17723 | -0.413 | -0.0690 | Yes |
| 163 | Cln6 | 17770 | -0.417 | -0.0613 | Yes |
| 164 | Gusb | 17968 | -0.439 | -0.0608 | Yes |
| 165 | Rbck1 | 18077 | -0.450 | -0.0555 | Yes |
| 166 | Pmm2 | 18287 | -0.475 | -0.0547 | Yes |
| 167 | Capn5 | 18317 | -0.479 | -0.0446 | Yes |
| 168 | Cited2 | 18373 | -0.488 | -0.0356 | Yes |
| 169 | Pgm2 | 18601 | -0.524 | -0.0346 | Yes |
| 170 | B3gat3 | 18770 | -0.555 | -0.0299 | Yes |
| 171 | Gmppb | 18947 | -0.595 | -0.0245 | Yes |
| 172 | Txn1 | 18963 | -0.599 | -0.0108 | Yes |
| 173 | Tpst1 | 19319 | -0.778 | -0.0102 | Yes |
| 174 | Isg20 | 19389 | -0.831 | 0.0064 | Yes |