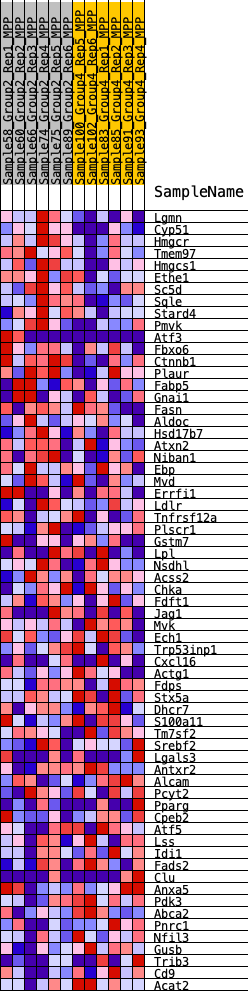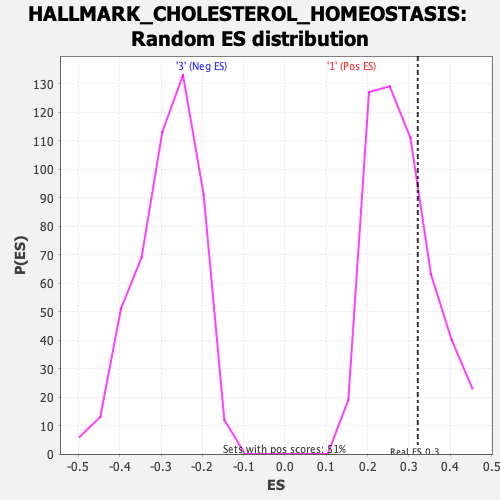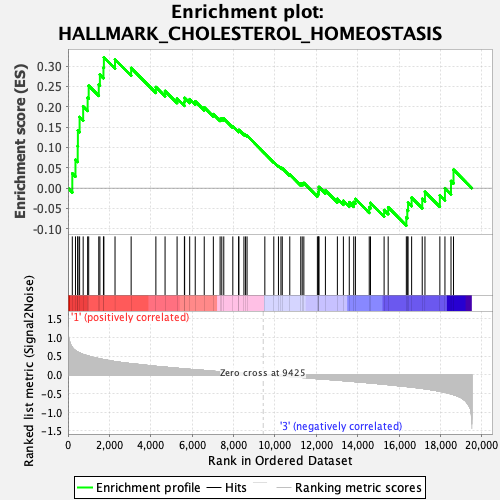
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group2_versus_Group4.MPP_Pheno.cls#Group2_versus_Group4_repos |
| Phenotype | MPP_Pheno.cls#Group2_versus_Group4_repos |
| Upregulated in class | 1 |
| GeneSet | HALLMARK_CHOLESTEROL_HOMEOSTASIS |
| Enrichment Score (ES) | 0.3214031 |
| Normalized Enrichment Score (NES) | 1.1371586 |
| Nominal p-value | 0.25 |
| FDR q-value | 0.90034837 |
| FWER p-Value | 0.97 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Lgmn | 208 | 0.726 | 0.0361 | Yes |
| 2 | Cyp51 | 361 | 0.644 | 0.0698 | Yes |
| 3 | Hmgcr | 469 | 0.605 | 0.1033 | Yes |
| 4 | Tmem97 | 478 | 0.603 | 0.1417 | Yes |
| 5 | Hmgcs1 | 561 | 0.580 | 0.1749 | Yes |
| 6 | Ethe1 | 733 | 0.540 | 0.2009 | Yes |
| 7 | Sc5d | 949 | 0.504 | 0.2224 | Yes |
| 8 | Sqle | 1000 | 0.496 | 0.2518 | Yes |
| 9 | Stard4 | 1487 | 0.430 | 0.2545 | Yes |
| 10 | Pmvk | 1541 | 0.424 | 0.2791 | Yes |
| 11 | Atf3 | 1713 | 0.405 | 0.2964 | Yes |
| 12 | Fbxo6 | 1734 | 0.404 | 0.3214 | Yes |
| 13 | Ctnnb1 | 2272 | 0.350 | 0.3164 | No |
| 14 | Plaur | 3051 | 0.297 | 0.2955 | No |
| 15 | Fabp5 | 4246 | 0.225 | 0.2486 | No |
| 16 | Gnai1 | 4690 | 0.204 | 0.2389 | No |
| 17 | Fasn | 5271 | 0.174 | 0.2203 | No |
| 18 | Aldoc | 5624 | 0.157 | 0.2123 | No |
| 19 | Hsd17b7 | 5636 | 0.156 | 0.2218 | No |
| 20 | Atxn2 | 5880 | 0.145 | 0.2187 | No |
| 21 | Niban1 | 6148 | 0.132 | 0.2134 | No |
| 22 | Ebp | 6583 | 0.114 | 0.1985 | No |
| 23 | Mvd | 7023 | 0.093 | 0.1819 | No |
| 24 | Errfi1 | 7350 | 0.080 | 0.1703 | No |
| 25 | Ldlr | 7431 | 0.077 | 0.1712 | No |
| 26 | Tnfrsf12a | 7527 | 0.074 | 0.1710 | No |
| 27 | Plscr1 | 7964 | 0.056 | 0.1522 | No |
| 28 | Gstm7 | 8244 | 0.045 | 0.1408 | No |
| 29 | Lpl | 8256 | 0.045 | 0.1431 | No |
| 30 | Nsdhl | 8501 | 0.035 | 0.1328 | No |
| 31 | Acss2 | 8580 | 0.031 | 0.1308 | No |
| 32 | Chka | 8652 | 0.028 | 0.1290 | No |
| 33 | Fdft1 | 9511 | -0.001 | 0.0849 | No |
| 34 | Jag1 | 9944 | -0.019 | 0.0639 | No |
| 35 | Mvk | 10169 | -0.027 | 0.0542 | No |
| 36 | Ech1 | 10290 | -0.032 | 0.0500 | No |
| 37 | Trp53inp1 | 10353 | -0.035 | 0.0491 | No |
| 38 | Cxcl16 | 10716 | -0.048 | 0.0336 | No |
| 39 | Actg1 | 11247 | -0.069 | 0.0108 | No |
| 40 | Fdps | 11317 | -0.072 | 0.0119 | No |
| 41 | Stx5a | 11396 | -0.075 | 0.0127 | No |
| 42 | Dhcr7 | 12050 | -0.104 | -0.0142 | No |
| 43 | S100a11 | 12095 | -0.106 | -0.0096 | No |
| 44 | Tm7sf2 | 12117 | -0.107 | -0.0038 | No |
| 45 | Srebf2 | 12122 | -0.107 | 0.0029 | No |
| 46 | Lgals3 | 12438 | -0.120 | -0.0056 | No |
| 47 | Antxr2 | 13016 | -0.142 | -0.0261 | No |
| 48 | Alcam | 13307 | -0.155 | -0.0310 | No |
| 49 | Pcyt2 | 13591 | -0.169 | -0.0347 | No |
| 50 | Pparg | 13806 | -0.178 | -0.0342 | No |
| 51 | Cpeb2 | 13888 | -0.181 | -0.0267 | No |
| 52 | Atf5 | 14556 | -0.214 | -0.0472 | No |
| 53 | Lss | 14616 | -0.217 | -0.0362 | No |
| 54 | Idi1 | 15274 | -0.251 | -0.0538 | No |
| 55 | Fads2 | 15471 | -0.261 | -0.0471 | No |
| 56 | Clu | 16348 | -0.311 | -0.0720 | No |
| 57 | Anxa5 | 16395 | -0.314 | -0.0541 | No |
| 58 | Pdk3 | 16432 | -0.316 | -0.0356 | No |
| 59 | Abca2 | 16604 | -0.327 | -0.0233 | No |
| 60 | Pnrc1 | 17116 | -0.363 | -0.0262 | No |
| 61 | Nfil3 | 17248 | -0.372 | -0.0090 | No |
| 62 | Gusb | 17968 | -0.439 | -0.0177 | No |
| 63 | Trib3 | 18214 | -0.466 | -0.0002 | No |
| 64 | Cd9 | 18507 | -0.509 | 0.0176 | No |
| 65 | Acat2 | 18628 | -0.528 | 0.0455 | No |