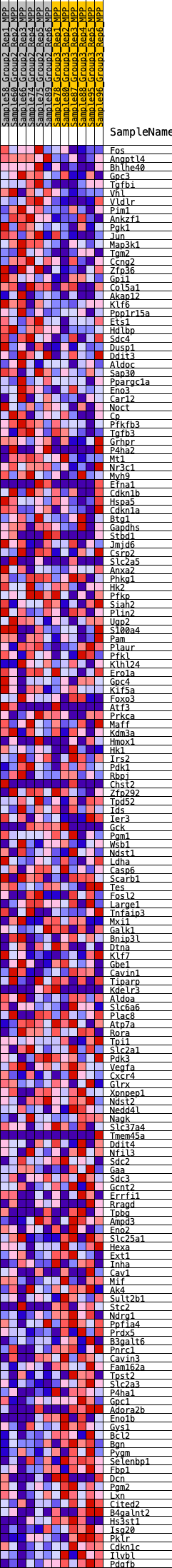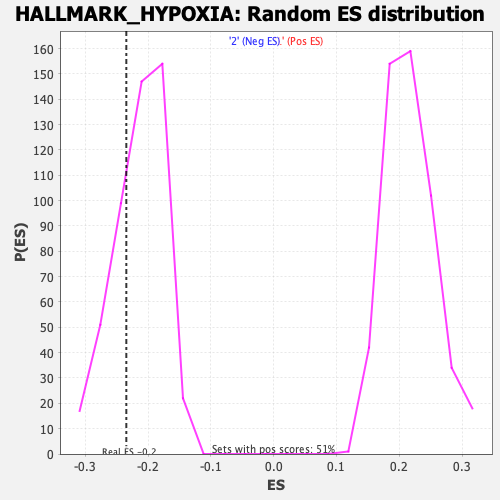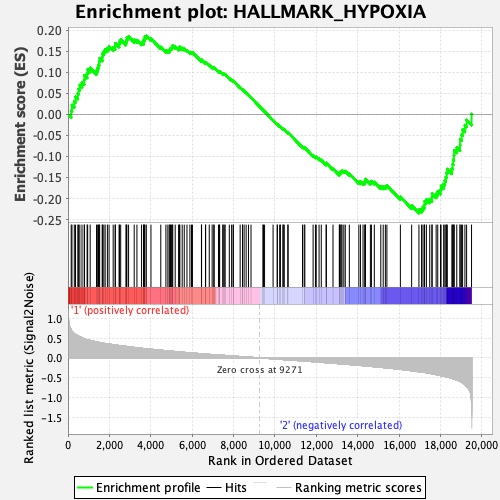
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group2_versus_Group3.MPP_Pheno.cls#Group2_versus_Group3_repos |
| Phenotype | MPP_Pheno.cls#Group2_versus_Group3_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_HYPOXIA |
| Enrichment Score (ES) | -0.23466419 |
| Normalized Enrichment Score (NES) | -1.0983914 |
| Nominal p-value | 0.26122448 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.978 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Fos | 154 | 0.699 | 0.0084 | No |
| 2 | Angptl4 | 191 | 0.671 | 0.0222 | No |
| 3 | Bhlhe40 | 304 | 0.614 | 0.0308 | No |
| 4 | Gpc3 | 364 | 0.592 | 0.0416 | No |
| 5 | Tgfbi | 475 | 0.561 | 0.0490 | No |
| 6 | Vhl | 513 | 0.549 | 0.0600 | No |
| 7 | Vldlr | 572 | 0.535 | 0.0695 | No |
| 8 | Pim1 | 677 | 0.508 | 0.0760 | No |
| 9 | Ankzf1 | 784 | 0.484 | 0.0818 | No |
| 10 | Pgk1 | 794 | 0.481 | 0.0926 | No |
| 11 | Jun | 923 | 0.462 | 0.0968 | No |
| 12 | Map3k1 | 950 | 0.457 | 0.1062 | No |
| 13 | Tgm2 | 1071 | 0.442 | 0.1103 | No |
| 14 | Ccng2 | 1379 | 0.404 | 0.1039 | No |
| 15 | Zfp36 | 1435 | 0.397 | 0.1103 | No |
| 16 | Gpi1 | 1475 | 0.393 | 0.1175 | No |
| 17 | Col5a1 | 1508 | 0.391 | 0.1250 | No |
| 18 | Akap12 | 1529 | 0.389 | 0.1331 | No |
| 19 | Klf6 | 1657 | 0.377 | 0.1353 | No |
| 20 | Ppp1r15a | 1660 | 0.376 | 0.1440 | No |
| 21 | Ets1 | 1730 | 0.369 | 0.1491 | No |
| 22 | Hdlbp | 1795 | 0.363 | 0.1543 | No |
| 23 | Sdc4 | 1904 | 0.355 | 0.1570 | No |
| 24 | Dusp1 | 1985 | 0.349 | 0.1610 | No |
| 25 | Ddit3 | 2183 | 0.335 | 0.1587 | No |
| 26 | Aldoc | 2278 | 0.327 | 0.1614 | No |
| 27 | Sap30 | 2288 | 0.326 | 0.1686 | No |
| 28 | Ppargc1a | 2468 | 0.314 | 0.1667 | No |
| 29 | Eno3 | 2491 | 0.312 | 0.1728 | No |
| 30 | Car12 | 2546 | 0.309 | 0.1773 | No |
| 31 | Noct | 2799 | 0.292 | 0.1711 | No |
| 32 | Cp | 2827 | 0.290 | 0.1765 | No |
| 33 | Pfkfb3 | 2846 | 0.288 | 0.1823 | No |
| 34 | Tgfb3 | 2927 | 0.283 | 0.1848 | No |
| 35 | Grhpr | 3199 | 0.265 | 0.1770 | No |
| 36 | P4ha2 | 3331 | 0.255 | 0.1762 | No |
| 37 | Mt1 | 3555 | 0.243 | 0.1703 | No |
| 38 | Nr3c1 | 3651 | 0.237 | 0.1710 | No |
| 39 | Myh9 | 3658 | 0.237 | 0.1762 | No |
| 40 | Efna1 | 3703 | 0.235 | 0.1794 | No |
| 41 | Cdkn1b | 3711 | 0.234 | 0.1845 | No |
| 42 | Hspa5 | 3784 | 0.230 | 0.1862 | No |
| 43 | Cdkn1a | 4008 | 0.219 | 0.1798 | No |
| 44 | Btg1 | 4483 | 0.195 | 0.1599 | No |
| 45 | Gapdhs | 4731 | 0.183 | 0.1514 | No |
| 46 | Stbd1 | 4826 | 0.178 | 0.1507 | No |
| 47 | Jmjd6 | 4892 | 0.175 | 0.1514 | No |
| 48 | Csrp2 | 4926 | 0.174 | 0.1538 | No |
| 49 | Slc2a5 | 4951 | 0.173 | 0.1566 | No |
| 50 | Anxa2 | 5007 | 0.170 | 0.1577 | No |
| 51 | Phkg1 | 5033 | 0.169 | 0.1604 | No |
| 52 | Hk2 | 5062 | 0.167 | 0.1629 | No |
| 53 | Pfkp | 5180 | 0.162 | 0.1606 | No |
| 54 | Siah2 | 5344 | 0.153 | 0.1557 | No |
| 55 | Plin2 | 5396 | 0.150 | 0.1566 | No |
| 56 | Ugp2 | 5398 | 0.150 | 0.1601 | No |
| 57 | S100a4 | 5517 | 0.145 | 0.1574 | No |
| 58 | Pam | 5617 | 0.140 | 0.1555 | No |
| 59 | Plaur | 5748 | 0.135 | 0.1520 | No |
| 60 | Pfkl | 5890 | 0.129 | 0.1477 | No |
| 61 | Klhl24 | 5968 | 0.125 | 0.1466 | No |
| 62 | Ero1a | 6013 | 0.123 | 0.1473 | No |
| 63 | Gpc4 | 6448 | 0.106 | 0.1273 | No |
| 64 | Kif5a | 6455 | 0.105 | 0.1295 | No |
| 65 | Foxo3 | 6643 | 0.098 | 0.1221 | No |
| 66 | Atf3 | 6651 | 0.098 | 0.1240 | No |
| 67 | Prkca | 6825 | 0.090 | 0.1172 | No |
| 68 | Maff | 6965 | 0.085 | 0.1119 | No |
| 69 | Kdm3a | 7045 | 0.082 | 0.1098 | No |
| 70 | Hmox1 | 7062 | 0.081 | 0.1108 | No |
| 71 | Hk1 | 7271 | 0.074 | 0.1018 | No |
| 72 | Irs2 | 7314 | 0.072 | 0.1013 | No |
| 73 | Pdk1 | 7328 | 0.072 | 0.1023 | No |
| 74 | Rbpj | 7478 | 0.067 | 0.0962 | No |
| 75 | Chst2 | 7497 | 0.066 | 0.0968 | No |
| 76 | Zfp292 | 7549 | 0.064 | 0.0957 | No |
| 77 | Tpd52 | 7594 | 0.062 | 0.0949 | No |
| 78 | Ids | 7794 | 0.055 | 0.0859 | No |
| 79 | Ier3 | 7906 | 0.050 | 0.0813 | No |
| 80 | Gck | 7982 | 0.048 | 0.0785 | No |
| 81 | Pgm1 | 7988 | 0.047 | 0.0794 | No |
| 82 | Wsb1 | 8318 | 0.034 | 0.0632 | No |
| 83 | Ndst1 | 8432 | 0.030 | 0.0581 | No |
| 84 | Ldha | 8434 | 0.030 | 0.0587 | No |
| 85 | Casp6 | 8510 | 0.027 | 0.0555 | No |
| 86 | Scarb1 | 8608 | 0.023 | 0.0510 | No |
| 87 | Tes | 8720 | 0.019 | 0.0457 | No |
| 88 | Fosl2 | 8843 | 0.015 | 0.0398 | No |
| 89 | Large1 | 9417 | -0.004 | 0.0102 | No |
| 90 | Tnfaip3 | 9446 | -0.005 | 0.0089 | No |
| 91 | Mxi1 | 9498 | -0.007 | 0.0064 | No |
| 92 | Galk1 | 9909 | -0.021 | -0.0142 | No |
| 93 | Bnip3l | 10111 | -0.029 | -0.0240 | No |
| 94 | Dtna | 10113 | -0.029 | -0.0233 | No |
| 95 | Klf7 | 10228 | -0.033 | -0.0285 | No |
| 96 | Gbe1 | 10264 | -0.035 | -0.0294 | No |
| 97 | Cavin1 | 10385 | -0.040 | -0.0347 | No |
| 98 | Tiparp | 10400 | -0.041 | -0.0345 | No |
| 99 | Kdelr3 | 10450 | -0.042 | -0.0360 | No |
| 100 | Aldoa | 10628 | -0.048 | -0.0441 | No |
| 101 | Slc6a6 | 10629 | -0.048 | -0.0429 | No |
| 102 | Plac8 | 11331 | -0.074 | -0.0774 | No |
| 103 | Atp7a | 11411 | -0.077 | -0.0797 | No |
| 104 | Rora | 11444 | -0.078 | -0.0796 | No |
| 105 | Tpi1 | 11843 | -0.094 | -0.0979 | No |
| 106 | Slc2a1 | 11956 | -0.098 | -0.1014 | No |
| 107 | Pdk3 | 11994 | -0.100 | -0.1010 | No |
| 108 | Vegfa | 12131 | -0.105 | -0.1056 | No |
| 109 | Cxcr4 | 12238 | -0.109 | -0.1085 | No |
| 110 | Glrx | 12473 | -0.119 | -0.1178 | No |
| 111 | Xpnpep1 | 12482 | -0.120 | -0.1154 | No |
| 112 | Ndst2 | 12805 | -0.133 | -0.1290 | No |
| 113 | Nedd4l | 13112 | -0.145 | -0.1414 | No |
| 114 | Nagk | 13116 | -0.145 | -0.1382 | No |
| 115 | Slc37a4 | 13181 | -0.148 | -0.1380 | No |
| 116 | Tmem45a | 13195 | -0.148 | -0.1352 | No |
| 117 | Ddit4 | 13242 | -0.149 | -0.1341 | No |
| 118 | Nfil3 | 13328 | -0.154 | -0.1349 | No |
| 119 | Sdc2 | 13398 | -0.157 | -0.1348 | No |
| 120 | Gaa | 13594 | -0.165 | -0.1410 | No |
| 121 | Sdc3 | 14053 | -0.185 | -0.1604 | No |
| 122 | Gcnt2 | 14131 | -0.189 | -0.1599 | No |
| 123 | Errfi1 | 14256 | -0.194 | -0.1618 | No |
| 124 | Rragd | 14339 | -0.197 | -0.1614 | No |
| 125 | Tpbg | 14362 | -0.199 | -0.1579 | No |
| 126 | Ampd3 | 14371 | -0.199 | -0.1537 | No |
| 127 | Eno2 | 14611 | -0.211 | -0.1611 | No |
| 128 | Slc25a1 | 14661 | -0.214 | -0.1586 | No |
| 129 | Hexa | 14801 | -0.220 | -0.1607 | No |
| 130 | Ext1 | 15114 | -0.235 | -0.1713 | No |
| 131 | Inha | 15230 | -0.242 | -0.1716 | No |
| 132 | Cav1 | 15335 | -0.248 | -0.1711 | No |
| 133 | Mif | 15409 | -0.251 | -0.1691 | No |
| 134 | Ak4 | 16060 | -0.286 | -0.1959 | No |
| 135 | Sult2b1 | 16605 | -0.323 | -0.2165 | No |
| 136 | Stc2 | 16957 | -0.348 | -0.2265 | Yes |
| 137 | Ndrg1 | 17084 | -0.356 | -0.2247 | Yes |
| 138 | Ppfia4 | 17153 | -0.361 | -0.2198 | Yes |
| 139 | Prdx5 | 17220 | -0.366 | -0.2146 | Yes |
| 140 | B3galt6 | 17231 | -0.366 | -0.2066 | Yes |
| 141 | Pnrc1 | 17318 | -0.373 | -0.2023 | Yes |
| 142 | Cavin3 | 17479 | -0.388 | -0.2015 | Yes |
| 143 | Fam162a | 17586 | -0.401 | -0.1976 | Yes |
| 144 | Tpst2 | 17592 | -0.401 | -0.1885 | Yes |
| 145 | Slc2a3 | 17789 | -0.419 | -0.1888 | Yes |
| 146 | P4ha1 | 17866 | -0.429 | -0.1827 | Yes |
| 147 | Gpc1 | 18000 | -0.444 | -0.1792 | Yes |
| 148 | Adora2b | 18025 | -0.447 | -0.1700 | Yes |
| 149 | Eno1b | 18152 | -0.462 | -0.1657 | Yes |
| 150 | Gys1 | 18210 | -0.469 | -0.1576 | Yes |
| 151 | Bcl2 | 18258 | -0.475 | -0.1489 | Yes |
| 152 | Bgn | 18293 | -0.479 | -0.1395 | Yes |
| 153 | Pygm | 18328 | -0.483 | -0.1300 | Yes |
| 154 | Selenbp1 | 18549 | -0.515 | -0.1293 | Yes |
| 155 | Fbp1 | 18585 | -0.521 | -0.1189 | Yes |
| 156 | Dcn | 18612 | -0.526 | -0.1079 | Yes |
| 157 | Pgm2 | 18646 | -0.533 | -0.0972 | Yes |
| 158 | Lxn | 18660 | -0.535 | -0.0853 | Yes |
| 159 | Cited2 | 18785 | -0.557 | -0.0787 | Yes |
| 160 | B4galnt2 | 18940 | -0.593 | -0.0728 | Yes |
| 161 | Hs3st1 | 18948 | -0.596 | -0.0592 | Yes |
| 162 | Isg20 | 19019 | -0.613 | -0.0485 | Yes |
| 163 | Pklr | 19067 | -0.629 | -0.0362 | Yes |
| 164 | Cdkn1c | 19179 | -0.679 | -0.0261 | Yes |
| 165 | Ilvbl | 19260 | -0.718 | -0.0134 | Yes |
| 166 | Pdgfb | 19499 | -1.129 | 0.0007 | Yes |