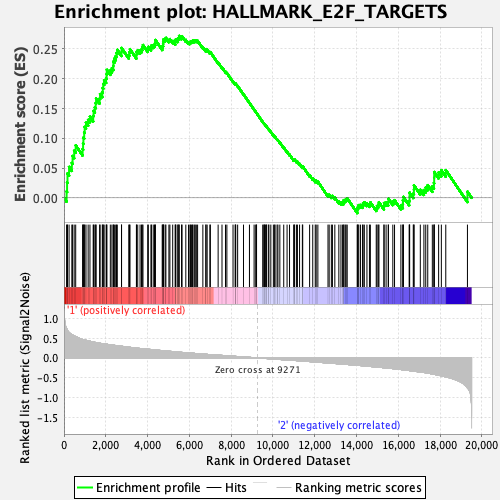
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group2_versus_Group3.MPP_Pheno.cls#Group2_versus_Group3_repos |
| Phenotype | MPP_Pheno.cls#Group2_versus_Group3_repos |
| Upregulated in class | 1 |
| GeneSet | HALLMARK_E2F_TARGETS |
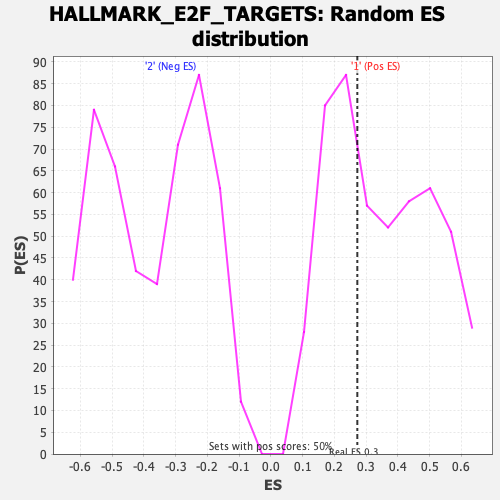
| Enrichment Score (ES) | 0.27197078 |
| Normalized Enrichment Score (NES) | 0.7757351 |
| Nominal p-value | 0.6003976 |
| FDR q-value | 0.98145664 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Prps1 | 125 | 0.743 | 0.0107 | Yes |
| 2 | Ccne1 | 148 | 0.708 | 0.0260 | Yes |
| 3 | Psmc3ip | 167 | 0.687 | 0.0410 | Yes |
| 4 | Rpa3 | 243 | 0.641 | 0.0519 | Yes |
| 5 | Dnmt1 | 376 | 0.587 | 0.0587 | Yes |
| 6 | Dck | 411 | 0.577 | 0.0703 | Yes |
| 7 | Mcm4 | 487 | 0.558 | 0.0793 | Yes |
| 8 | Cse1l | 560 | 0.538 | 0.0880 | Yes |
| 9 | Srsf1 | 891 | 0.466 | 0.0817 | Yes |
| 10 | Mcm3 | 914 | 0.464 | 0.0913 | Yes |
| 11 | Ipo7 | 929 | 0.461 | 0.1013 | Yes |
| 12 | Smc1a | 967 | 0.455 | 0.1099 | Yes |
| 13 | Zw10 | 990 | 0.452 | 0.1192 | Yes |
| 14 | Rbbp7 | 1052 | 0.445 | 0.1263 | Yes |
| 15 | Mcm2 | 1156 | 0.431 | 0.1310 | Yes |
| 16 | Rad51ap1 | 1242 | 0.421 | 0.1363 | Yes |
| 17 | Mcm6 | 1397 | 0.402 | 0.1377 | Yes |
| 18 | Mcm7 | 1416 | 0.399 | 0.1460 | Yes |
| 19 | Mybl2 | 1478 | 0.393 | 0.1519 | Yes |
| 20 | Pola2 | 1516 | 0.390 | 0.1590 | Yes |
| 21 | Msh2 | 1537 | 0.388 | 0.1670 | Yes |
| 22 | Ubr7 | 1706 | 0.371 | 0.1669 | Yes |
| 23 | Prkdc | 1728 | 0.369 | 0.1743 | Yes |
| 24 | Timeless | 1833 | 0.360 | 0.1773 | Yes |
| 25 | Hnrnpd | 1855 | 0.358 | 0.1845 | Yes |
| 26 | Pnn | 1881 | 0.357 | 0.1914 | Yes |
| 27 | Pcna | 1921 | 0.355 | 0.1976 | Yes |
| 28 | Nbn | 2016 | 0.347 | 0.2008 | Yes |
| 29 | Xpo1 | 2039 | 0.345 | 0.2076 | Yes |
| 30 | Lmnb1 | 2052 | 0.344 | 0.2150 | Yes |
| 31 | Smc3 | 2218 | 0.331 | 0.2141 | Yes |
| 32 | Nap1l1 | 2294 | 0.325 | 0.2177 | Yes |
| 33 | Ccp110 | 2361 | 0.320 | 0.2217 | Yes |
| 34 | Lig1 | 2365 | 0.320 | 0.2290 | Yes |
| 35 | Tfrc | 2411 | 0.318 | 0.2340 | Yes |
| 36 | Lyar | 2477 | 0.313 | 0.2379 | Yes |
| 37 | Hells | 2513 | 0.311 | 0.2433 | Yes |
| 38 | Nasp | 2551 | 0.309 | 0.2485 | Yes |
| 39 | Dut | 2753 | 0.295 | 0.2449 | Yes |
| 40 | Eed | 2755 | 0.295 | 0.2517 | Yes |
| 41 | Kif22 | 3101 | 0.272 | 0.2401 | Yes |
| 42 | Rpa1 | 3133 | 0.269 | 0.2447 | Yes |
| 43 | Cdc25a | 3159 | 0.268 | 0.2496 | Yes |
| 44 | Donson | 3467 | 0.248 | 0.2395 | Yes |
| 45 | Melk | 3469 | 0.248 | 0.2452 | Yes |
| 46 | Pold3 | 3529 | 0.244 | 0.2478 | Yes |
| 47 | Lbr | 3641 | 0.238 | 0.2475 | Yes |
| 48 | Cdkn1b | 3711 | 0.234 | 0.2494 | Yes |
| 49 | Eif2s1 | 3747 | 0.232 | 0.2529 | Yes |
| 50 | Rad21 | 3780 | 0.230 | 0.2566 | Yes |
| 51 | Cdkn1a | 4008 | 0.219 | 0.2499 | Yes |
| 52 | Chek2 | 4036 | 0.218 | 0.2536 | Yes |
| 53 | Chek1 | 4164 | 0.212 | 0.2519 | Yes |
| 54 | Cdkn2c | 4183 | 0.211 | 0.2559 | Yes |
| 55 | Nolc1 | 4271 | 0.206 | 0.2561 | Yes |
| 56 | Tacc3 | 4332 | 0.203 | 0.2577 | Yes |
| 57 | Brca1 | 4366 | 0.201 | 0.2607 | Yes |
| 58 | Anp32e | 4372 | 0.201 | 0.2651 | Yes |
| 59 | Ilf3 | 4701 | 0.185 | 0.2524 | Yes |
| 60 | Psip1 | 4726 | 0.184 | 0.2554 | Yes |
| 61 | Mre11a | 4746 | 0.183 | 0.2586 | Yes |
| 62 | Nup205 | 4750 | 0.183 | 0.2627 | Yes |
| 63 | Pole | 4765 | 0.182 | 0.2662 | Yes |
| 64 | Aurka | 4841 | 0.177 | 0.2664 | Yes |
| 65 | Ddx39a | 4877 | 0.175 | 0.2686 | Yes |
| 66 | H2az1 | 5022 | 0.169 | 0.2651 | Yes |
| 67 | Ncapd2 | 5069 | 0.167 | 0.2666 | Yes |
| 68 | Mlh1 | 5203 | 0.160 | 0.2634 | Yes |
| 69 | Syncrip | 5326 | 0.154 | 0.2607 | Yes |
| 70 | Nup107 | 5329 | 0.153 | 0.2641 | Yes |
| 71 | Pa2g4 | 5356 | 0.152 | 0.2663 | Yes |
| 72 | Ssrp1 | 5450 | 0.148 | 0.2649 | Yes |
| 73 | Rad51c | 5458 | 0.147 | 0.2679 | Yes |
| 74 | Atad2 | 5502 | 0.145 | 0.2691 | Yes |
| 75 | Hus1 | 5512 | 0.145 | 0.2720 | Yes |
| 76 | Tra2b | 5626 | 0.140 | 0.2694 | No |
| 77 | Lsm8 | 5651 | 0.139 | 0.2713 | No |
| 78 | Tipin | 5831 | 0.132 | 0.2651 | No |
| 79 | Ezh2 | 5961 | 0.126 | 0.2613 | No |
| 80 | Ppp1r8 | 6027 | 0.123 | 0.2608 | No |
| 81 | Stag1 | 6052 | 0.122 | 0.2624 | No |
| 82 | Mms22l | 6099 | 0.121 | 0.2628 | No |
| 83 | Ran | 6148 | 0.119 | 0.2631 | No |
| 84 | Shmt1 | 6174 | 0.118 | 0.2645 | No |
| 85 | Usp1 | 6238 | 0.115 | 0.2639 | No |
| 86 | Orc2 | 6282 | 0.113 | 0.2643 | No |
| 87 | Gins4 | 6334 | 0.110 | 0.2642 | No |
| 88 | Asf1b | 6395 | 0.108 | 0.2636 | No |
| 89 | Brca2 | 6647 | 0.098 | 0.2529 | No |
| 90 | Rrm2 | 6792 | 0.091 | 0.2475 | No |
| 91 | Paics | 6793 | 0.091 | 0.2497 | No |
| 92 | Rfc3 | 6852 | 0.089 | 0.2487 | No |
| 93 | Wdr90 | 6983 | 0.084 | 0.2439 | No |
| 94 | Ak2 | 7006 | 0.083 | 0.2447 | No |
| 95 | Ranbp1 | 7378 | 0.070 | 0.2271 | No |
| 96 | Mcm5 | 7563 | 0.064 | 0.2191 | No |
| 97 | Slbp | 7730 | 0.057 | 0.2118 | No |
| 98 | Rfc1 | 7796 | 0.055 | 0.2097 | No |
| 99 | Pold2 | 8091 | 0.044 | 0.1955 | No |
| 100 | Brms1l | 8199 | 0.039 | 0.1908 | No |
| 101 | Bub1b | 8200 | 0.039 | 0.1918 | No |
| 102 | Smc6 | 8221 | 0.039 | 0.1916 | No |
| 103 | Spc25 | 8312 | 0.035 | 0.1878 | No |
| 104 | Gspt1 | 8592 | 0.024 | 0.1739 | No |
| 105 | Cdk4 | 8877 | 0.015 | 0.1595 | No |
| 106 | Nme1 | 9100 | 0.007 | 0.1482 | No |
| 107 | Tmpo | 9179 | 0.004 | 0.1442 | No |
| 108 | Cbx5 | 9205 | 0.003 | 0.1430 | No |
| 109 | Pds5b | 9217 | 0.002 | 0.1425 | No |
| 110 | Rad50 | 9512 | -0.008 | 0.1274 | No |
| 111 | Nup153 | 9565 | -0.009 | 0.1249 | No |
| 112 | Snrpb | 9586 | -0.010 | 0.1241 | No |
| 113 | Top2a | 9593 | -0.011 | 0.1241 | No |
| 114 | Bard1 | 9652 | -0.012 | 0.1214 | No |
| 115 | Cdca8 | 9683 | -0.014 | 0.1201 | No |
| 116 | Suv39h1 | 9685 | -0.014 | 0.1204 | No |
| 117 | Tbrg4 | 9779 | -0.017 | 0.1160 | No |
| 118 | Luc7l3 | 9808 | -0.018 | 0.1149 | No |
| 119 | Kif4 | 9900 | -0.021 | 0.1107 | No |
| 120 | Dscc1 | 10022 | -0.026 | 0.1050 | No |
| 121 | Rpa2 | 10084 | -0.028 | 0.1025 | No |
| 122 | Tk1 | 10101 | -0.029 | 0.1024 | No |
| 123 | Prim2 | 10198 | -0.032 | 0.0981 | No |
| 124 | Hmgb3 | 10240 | -0.034 | 0.0968 | No |
| 125 | Orc6 | 10332 | -0.038 | 0.0930 | No |
| 126 | Hmga1b | 10518 | -0.044 | 0.0844 | No |
| 127 | Ctcf | 10678 | -0.050 | 0.0773 | No |
| 128 | Gins3 | 10796 | -0.054 | 0.0725 | No |
| 129 | Rnaseh2a | 10996 | -0.061 | 0.0636 | No |
| 130 | Nop56 | 11019 | -0.062 | 0.0639 | No |
| 131 | Trp53 | 11041 | -0.063 | 0.0643 | No |
| 132 | Tubb5 | 11133 | -0.066 | 0.0611 | No |
| 133 | Ing3 | 11173 | -0.068 | 0.0606 | No |
| 134 | Cdkn2a | 11276 | -0.072 | 0.0570 | No |
| 135 | Nudt21 | 11410 | -0.077 | 0.0519 | No |
| 136 | Xrcc6 | 11427 | -0.077 | 0.0529 | No |
| 137 | Pold1 | 11757 | -0.091 | 0.0379 | No |
| 138 | Dek | 11907 | -0.096 | 0.0324 | No |
| 139 | Pole4 | 12026 | -0.101 | 0.0287 | No |
| 140 | Ung | 12071 | -0.103 | 0.0288 | No |
| 141 | Hmgb2 | 12152 | -0.106 | 0.0271 | No |
| 142 | Cenpe | 12637 | -0.126 | 0.0049 | No |
| 143 | Spag5 | 12693 | -0.128 | 0.0051 | No |
| 144 | Tubg1 | 12798 | -0.132 | 0.0027 | No |
| 145 | Ube2s | 12849 | -0.134 | 0.0033 | No |
| 146 | Smc4 | 12964 | -0.139 | 0.0006 | No |
| 147 | Cnot9 | 13156 | -0.147 | -0.0059 | No |
| 148 | Phf5a | 13257 | -0.150 | -0.0076 | No |
| 149 | Espl1 | 13335 | -0.154 | -0.0080 | No |
| 150 | Ppm1d | 13391 | -0.156 | -0.0073 | No |
| 151 | Ube2t | 13395 | -0.156 | -0.0038 | No |
| 152 | Racgap1 | 13464 | -0.159 | -0.0037 | No |
| 153 | Rad1 | 13505 | -0.161 | -0.0020 | No |
| 154 | Cit | 13556 | -0.164 | -0.0008 | No |
| 155 | Mki67 | 14039 | -0.185 | -0.0215 | No |
| 156 | Plk4 | 14043 | -0.185 | -0.0173 | No |
| 157 | Cks1b | 14076 | -0.186 | -0.0147 | No |
| 158 | Birc5 | 14111 | -0.188 | -0.0121 | No |
| 159 | Mxd3 | 14195 | -0.191 | -0.0120 | No |
| 160 | Cenpm | 14291 | -0.196 | -0.0124 | No |
| 161 | Dlgap5 | 14317 | -0.197 | -0.0091 | No |
| 162 | Cdc20 | 14376 | -0.199 | -0.0075 | No |
| 163 | Diaph3 | 14501 | -0.206 | -0.0091 | No |
| 164 | Cdk1 | 14621 | -0.212 | -0.0104 | No |
| 165 | Kif2c | 14665 | -0.214 | -0.0077 | No |
| 166 | Srsf2 | 14946 | -0.227 | -0.0169 | No |
| 167 | Pop7 | 14985 | -0.229 | -0.0136 | No |
| 168 | Hmmr | 15044 | -0.232 | -0.0112 | No |
| 169 | Wee1 | 15076 | -0.234 | -0.0074 | No |
| 170 | Spc24 | 15314 | -0.247 | -0.0140 | No |
| 171 | Gins1 | 15322 | -0.247 | -0.0086 | No |
| 172 | Pan2 | 15418 | -0.251 | -0.0077 | No |
| 173 | E2f8 | 15521 | -0.257 | -0.0070 | No |
| 174 | Prdx4 | 15530 | -0.258 | -0.0015 | No |
| 175 | Mad2l1 | 15740 | -0.269 | -0.0061 | No |
| 176 | Cdca3 | 15822 | -0.273 | -0.0040 | No |
| 177 | Aurkb | 16126 | -0.292 | -0.0129 | No |
| 178 | Trip13 | 16220 | -0.297 | -0.0109 | No |
| 179 | Ccnb2 | 16224 | -0.297 | -0.0041 | No |
| 180 | Stmn1 | 16248 | -0.299 | 0.0016 | No |
| 181 | Dctpp1 | 16522 | -0.318 | -0.0052 | No |
| 182 | Mthfd2 | 16542 | -0.319 | 0.0012 | No |
| 183 | Asf1a | 16543 | -0.319 | 0.0086 | No |
| 184 | H2ax | 16719 | -0.331 | 0.0072 | No |
| 185 | Myc | 16749 | -0.334 | 0.0134 | No |
| 186 | Rfc2 | 16753 | -0.334 | 0.0210 | No |
| 187 | Jpt1 | 17056 | -0.355 | 0.0136 | No |
| 188 | Pttg1 | 17228 | -0.366 | 0.0132 | No |
| 189 | Pms2 | 17316 | -0.373 | 0.0173 | No |
| 190 | Tcf19 | 17412 | -0.381 | 0.0213 | No |
| 191 | Depdc1a | 17631 | -0.404 | 0.0193 | No |
| 192 | Kif18b | 17708 | -0.412 | 0.0249 | No |
| 193 | Exosc8 | 17720 | -0.412 | 0.0339 | No |
| 194 | Cks2 | 17722 | -0.413 | 0.0434 | No |
| 195 | Dclre1b | 17933 | -0.436 | 0.0426 | No |
| 196 | Plk1 | 18063 | -0.451 | 0.0464 | No |
| 197 | Cdc25b | 18275 | -0.477 | 0.0465 | No |
| 198 | Cdkn3 | 19312 | -0.756 | 0.0104 | No |