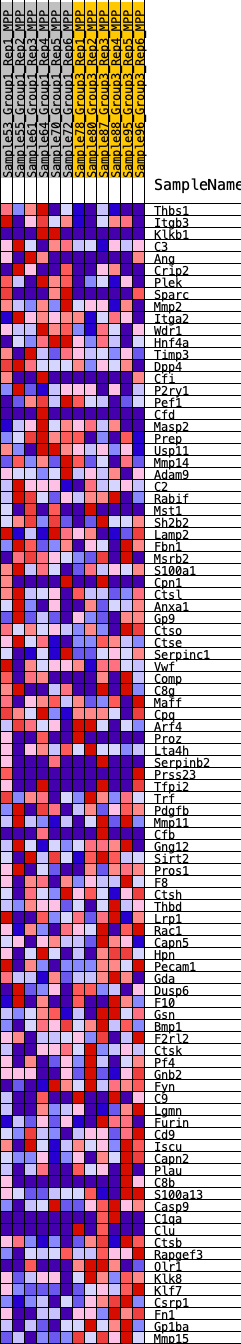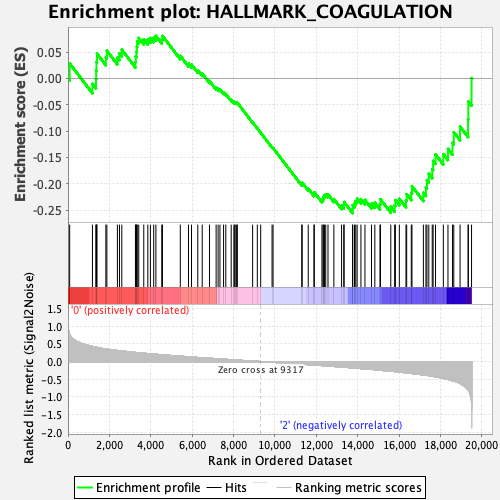
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group1_versus_Group3.MPP_Pheno.cls#Group1_versus_Group3_repos |
| Phenotype | MPP_Pheno.cls#Group1_versus_Group3_repos |
| Upregulated in class | 2 |
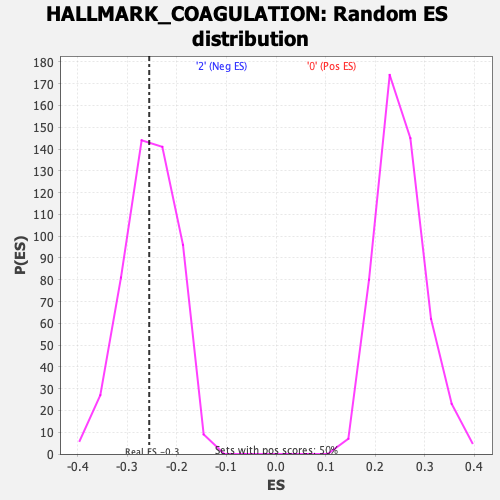
| GeneSet | HALLMARK_COAGULATION |
| Enrichment Score (ES) | -0.25570554 |
| Normalized Enrichment Score (NES) | -1.0045741 |
| Nominal p-value | 0.4702381 |
| FDR q-value | 0.84321594 |
| FWER p-Value | 0.998 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Thbs1 | 80 | 0.775 | 0.0285 | No |
| 2 | Itgb3 | 1181 | 0.430 | -0.0101 | No |
| 3 | Klkb1 | 1351 | 0.408 | -0.0016 | No |
| 4 | C3 | 1360 | 0.407 | 0.0151 | No |
| 5 | Ang | 1375 | 0.405 | 0.0314 | No |
| 6 | Crip2 | 1401 | 0.402 | 0.0470 | No |
| 7 | Plek | 1829 | 0.359 | 0.0402 | No |
| 8 | Sparc | 1878 | 0.356 | 0.0527 | No |
| 9 | Mmp2 | 2387 | 0.315 | 0.0398 | No |
| 10 | Itga2 | 2487 | 0.308 | 0.0476 | No |
| 11 | Wdr1 | 2600 | 0.303 | 0.0546 | No |
| 12 | Hnf4a | 3259 | 0.258 | 0.0316 | No |
| 13 | Timp3 | 3271 | 0.257 | 0.0418 | No |
| 14 | Dpp4 | 3307 | 0.254 | 0.0507 | No |
| 15 | Cfi | 3326 | 0.253 | 0.0604 | No |
| 16 | P2ry1 | 3342 | 0.252 | 0.0703 | No |
| 17 | Pef1 | 3418 | 0.249 | 0.0769 | No |
| 18 | Cfd | 3661 | 0.236 | 0.0744 | No |
| 19 | Masp2 | 3857 | 0.227 | 0.0739 | No |
| 20 | Prep | 3981 | 0.221 | 0.0768 | No |
| 21 | Usp11 | 4141 | 0.212 | 0.0776 | No |
| 22 | Mmp14 | 4247 | 0.207 | 0.0809 | No |
| 23 | Adam9 | 4530 | 0.194 | 0.0745 | No |
| 24 | C2 | 4564 | 0.192 | 0.0809 | No |
| 25 | Rabif | 5426 | 0.155 | 0.0430 | No |
| 26 | Mst1 | 5823 | 0.135 | 0.0283 | No |
| 27 | Sh2b2 | 5968 | 0.128 | 0.0263 | No |
| 28 | Lamp2 | 6271 | 0.117 | 0.0157 | No |
| 29 | Fbn1 | 6485 | 0.110 | 0.0093 | No |
| 30 | Msrb2 | 6836 | 0.095 | -0.0047 | No |
| 31 | S100a1 | 7157 | 0.083 | -0.0177 | No |
| 32 | Cpn1 | 7254 | 0.079 | -0.0193 | No |
| 33 | Ctsl | 7342 | 0.075 | -0.0206 | No |
| 34 | Anxa1 | 7517 | 0.068 | -0.0267 | No |
| 35 | Gp9 | 7625 | 0.064 | -0.0295 | No |
| 36 | Ctso | 7885 | 0.053 | -0.0406 | No |
| 37 | Ctse | 8019 | 0.048 | -0.0455 | No |
| 38 | Serpinc1 | 8036 | 0.048 | -0.0443 | No |
| 39 | Vwf | 8108 | 0.046 | -0.0460 | No |
| 40 | Comp | 8149 | 0.044 | -0.0462 | No |
| 41 | C8g | 8183 | 0.043 | -0.0461 | No |
| 42 | Maff | 8921 | 0.015 | -0.0834 | No |
| 43 | Cpq | 9148 | 0.007 | -0.0948 | No |
| 44 | Arf4 | 9311 | 0.000 | -0.1031 | No |
| 45 | Proz | 9860 | -0.019 | -0.1305 | No |
| 46 | Lta4h | 9909 | -0.021 | -0.1321 | No |
| 47 | Serpinb2 | 11295 | -0.076 | -0.2002 | No |
| 48 | Prss23 | 11315 | -0.076 | -0.1980 | No |
| 49 | Tfpi2 | 11608 | -0.087 | -0.2094 | No |
| 50 | Trf | 11883 | -0.097 | -0.2194 | No |
| 51 | Pdgfb | 11903 | -0.098 | -0.2163 | No |
| 52 | Mmp11 | 12270 | -0.113 | -0.2304 | No |
| 53 | Cfb | 12323 | -0.115 | -0.2282 | No |
| 54 | Gng12 | 12351 | -0.116 | -0.2247 | No |
| 55 | Sirt2 | 12392 | -0.118 | -0.2218 | No |
| 56 | Pros1 | 12454 | -0.121 | -0.2199 | No |
| 57 | F8 | 12563 | -0.124 | -0.2202 | No |
| 58 | Ctsh | 12847 | -0.138 | -0.2290 | No |
| 59 | Thbd | 13219 | -0.153 | -0.2416 | No |
| 60 | Lrp1 | 13327 | -0.158 | -0.2405 | No |
| 61 | Rac1 | 13341 | -0.158 | -0.2345 | No |
| 62 | Capn5 | 13753 | -0.179 | -0.2482 | Yes |
| 63 | Hpn | 13756 | -0.179 | -0.2407 | Yes |
| 64 | Pecam1 | 13842 | -0.183 | -0.2374 | Yes |
| 65 | Gda | 13904 | -0.186 | -0.2327 | Yes |
| 66 | Dusp6 | 13971 | -0.189 | -0.2282 | Yes |
| 67 | F10 | 14154 | -0.198 | -0.2292 | Yes |
| 68 | Gsn | 14349 | -0.207 | -0.2305 | Yes |
| 69 | Bmp1 | 14665 | -0.222 | -0.2374 | Yes |
| 70 | F2rl2 | 14826 | -0.229 | -0.2360 | Yes |
| 71 | Ctsk | 15075 | -0.244 | -0.2385 | Yes |
| 72 | Pf4 | 15098 | -0.246 | -0.2293 | Yes |
| 73 | Gnb2 | 15597 | -0.273 | -0.2434 | Yes |
| 74 | Fyn | 15783 | -0.282 | -0.2411 | Yes |
| 75 | C9 | 15819 | -0.284 | -0.2309 | Yes |
| 76 | Lgmn | 16010 | -0.296 | -0.2283 | Yes |
| 77 | Furin | 16331 | -0.317 | -0.2314 | Yes |
| 78 | Cd9 | 16365 | -0.320 | -0.2197 | Yes |
| 79 | Iscu | 16586 | -0.335 | -0.2169 | Yes |
| 80 | Capn2 | 16621 | -0.338 | -0.2045 | Yes |
| 81 | Plau | 17172 | -0.377 | -0.2169 | Yes |
| 82 | C8b | 17295 | -0.386 | -0.2070 | Yes |
| 83 | S100a13 | 17344 | -0.389 | -0.1931 | Yes |
| 84 | Casp9 | 17434 | -0.398 | -0.1809 | Yes |
| 85 | C1qa | 17599 | -0.415 | -0.1719 | Yes |
| 86 | Clu | 17642 | -0.419 | -0.1564 | Yes |
| 87 | Ctsb | 17758 | -0.432 | -0.1442 | Yes |
| 88 | Rapgef3 | 18136 | -0.477 | -0.1435 | Yes |
| 89 | Olr1 | 18360 | -0.506 | -0.1337 | Yes |
| 90 | Klk8 | 18578 | -0.542 | -0.1221 | Yes |
| 91 | Klf7 | 18641 | -0.553 | -0.1020 | Yes |
| 92 | Csrp1 | 18944 | -0.621 | -0.0914 | Yes |
| 93 | Fn1 | 19332 | -0.803 | -0.0775 | Yes |
| 94 | Gp1ba | 19345 | -0.817 | -0.0438 | Yes |
| 95 | Mmp15 | 19499 | -1.243 | 0.0007 | Yes |