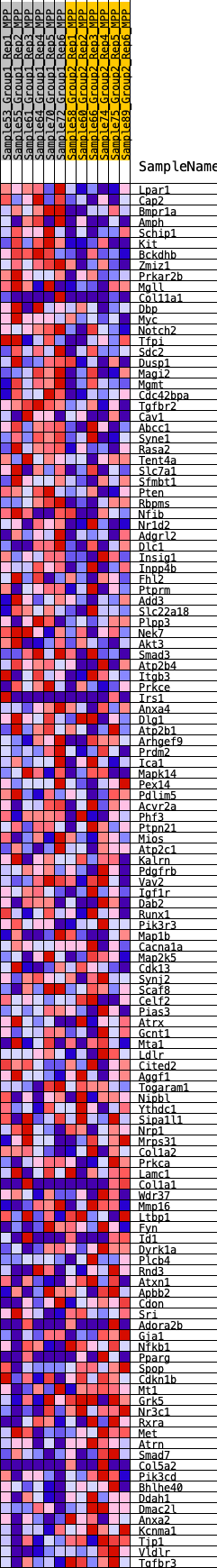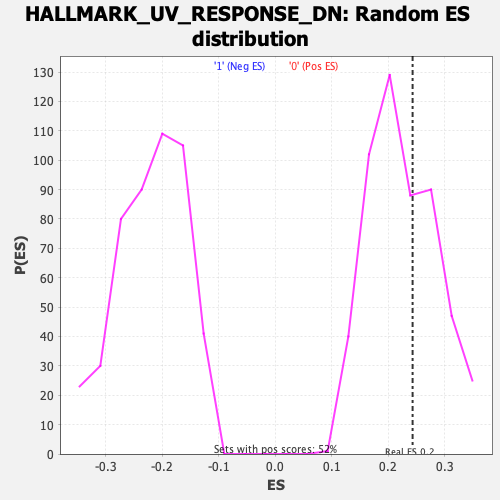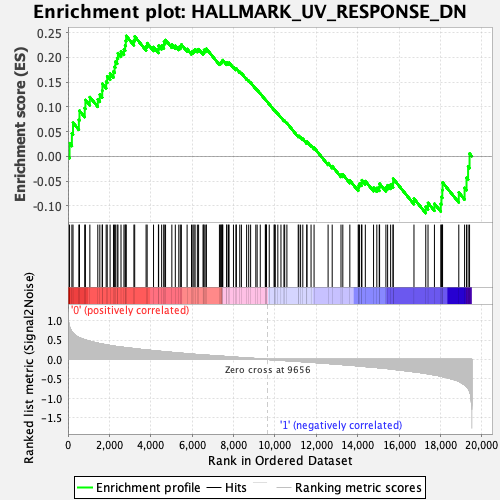
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MPP.MPP_Pheno.cls#Group1_versus_Group2.MPP_Pheno.cls#Group1_versus_Group2_repos |
| Phenotype | MPP_Pheno.cls#Group1_versus_Group2_repos |
| Upregulated in class | 0 |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | 0.24334791 |
| Normalized Enrichment Score (NES) | 1.0810009 |
| Nominal p-value | 0.36206895 |
| FDR q-value | 0.9446022 |
| FWER p-Value | 0.978 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Lpar1 | 75 | 0.841 | 0.0265 | Yes |
| 2 | Cap2 | 188 | 0.709 | 0.0463 | Yes |
| 3 | Bmpr1a | 242 | 0.678 | 0.0680 | Yes |
| 4 | Amph | 525 | 0.561 | 0.0737 | Yes |
| 5 | Schip1 | 555 | 0.554 | 0.0922 | Yes |
| 6 | Kit | 808 | 0.506 | 0.0974 | Yes |
| 7 | Bckdhb | 841 | 0.501 | 0.1139 | Yes |
| 8 | Zmiz1 | 1056 | 0.468 | 0.1197 | Yes |
| 9 | Prkar2b | 1440 | 0.417 | 0.1150 | Yes |
| 10 | Mgll | 1538 | 0.407 | 0.1247 | Yes |
| 11 | Col11a1 | 1653 | 0.394 | 0.1330 | Yes |
| 12 | Dbp | 1663 | 0.393 | 0.1467 | Yes |
| 13 | Myc | 1844 | 0.374 | 0.1509 | Yes |
| 14 | Notch2 | 1895 | 0.369 | 0.1617 | Yes |
| 15 | Tfpi | 2034 | 0.358 | 0.1674 | Yes |
| 16 | Sdc2 | 2197 | 0.344 | 0.1715 | Yes |
| 17 | Dusp1 | 2252 | 0.338 | 0.1809 | Yes |
| 18 | Magi2 | 2287 | 0.335 | 0.1912 | Yes |
| 19 | Mgmt | 2374 | 0.329 | 0.1987 | Yes |
| 20 | Cdc42bpa | 2413 | 0.326 | 0.2084 | Yes |
| 21 | Tgfbr2 | 2561 | 0.316 | 0.2123 | Yes |
| 22 | Cav1 | 2704 | 0.307 | 0.2160 | Yes |
| 23 | Abcc1 | 2760 | 0.305 | 0.2242 | Yes |
| 24 | Syne1 | 2791 | 0.302 | 0.2335 | Yes |
| 25 | Rasa2 | 2812 | 0.301 | 0.2433 | Yes |
| 26 | Tent4a | 3185 | 0.277 | 0.2342 | No |
| 27 | Slc7a1 | 3223 | 0.276 | 0.2422 | No |
| 28 | Sfmbt1 | 3773 | 0.242 | 0.2226 | No |
| 29 | Pten | 3824 | 0.240 | 0.2287 | No |
| 30 | Rbpms | 4133 | 0.223 | 0.2208 | No |
| 31 | Nfib | 4369 | 0.212 | 0.2164 | No |
| 32 | Nr1d2 | 4380 | 0.212 | 0.2235 | No |
| 33 | Adgrl2 | 4525 | 0.203 | 0.2234 | No |
| 34 | Dlc1 | 4633 | 0.198 | 0.2250 | No |
| 35 | Insig1 | 4641 | 0.197 | 0.2318 | No |
| 36 | Inpp4b | 4714 | 0.192 | 0.2350 | No |
| 37 | Fhl2 | 5018 | 0.179 | 0.2258 | No |
| 38 | Ptprm | 5185 | 0.172 | 0.2235 | No |
| 39 | Add3 | 5354 | 0.164 | 0.2207 | No |
| 40 | Slc22a18 | 5427 | 0.160 | 0.2228 | No |
| 41 | Plpp3 | 5478 | 0.158 | 0.2259 | No |
| 42 | Nek7 | 5755 | 0.145 | 0.2169 | No |
| 43 | Akt3 | 5967 | 0.136 | 0.2109 | No |
| 44 | Smad3 | 6033 | 0.133 | 0.2124 | No |
| 45 | Atp2b4 | 6087 | 0.130 | 0.2143 | No |
| 46 | Itgb3 | 6141 | 0.128 | 0.2162 | No |
| 47 | Prkce | 6251 | 0.123 | 0.2151 | No |
| 48 | Irs1 | 6310 | 0.121 | 0.2164 | No |
| 49 | Anxa4 | 6525 | 0.115 | 0.2095 | No |
| 50 | Dlg1 | 6562 | 0.113 | 0.2117 | No |
| 51 | Atp2b1 | 6576 | 0.113 | 0.2151 | No |
| 52 | Arhgef9 | 6663 | 0.109 | 0.2146 | No |
| 53 | Prdm2 | 6686 | 0.108 | 0.2174 | No |
| 54 | Ica1 | 7317 | 0.085 | 0.1880 | No |
| 55 | Mapk14 | 7362 | 0.083 | 0.1887 | No |
| 56 | Pex14 | 7399 | 0.082 | 0.1898 | No |
| 57 | Pdlim5 | 7426 | 0.081 | 0.1914 | No |
| 58 | Acvr2a | 7465 | 0.080 | 0.1923 | No |
| 59 | Phf3 | 7480 | 0.080 | 0.1945 | No |
| 60 | Ptpn21 | 7668 | 0.072 | 0.1874 | No |
| 61 | Mios | 7673 | 0.072 | 0.1898 | No |
| 62 | Atp2c1 | 7755 | 0.069 | 0.1881 | No |
| 63 | Kalrn | 7782 | 0.068 | 0.1892 | No |
| 64 | Pdgfrb | 7995 | 0.060 | 0.1804 | No |
| 65 | Vav2 | 8114 | 0.055 | 0.1763 | No |
| 66 | Igf1r | 8136 | 0.054 | 0.1772 | No |
| 67 | Dab2 | 8300 | 0.047 | 0.1705 | No |
| 68 | Runx1 | 8379 | 0.045 | 0.1681 | No |
| 69 | Pik3r3 | 8631 | 0.036 | 0.1565 | No |
| 70 | Map1b | 8731 | 0.033 | 0.1525 | No |
| 71 | Cacna1a | 8824 | 0.030 | 0.1489 | No |
| 72 | Map2k5 | 9075 | 0.021 | 0.1368 | No |
| 73 | Cdk13 | 9139 | 0.019 | 0.1342 | No |
| 74 | Synj2 | 9290 | 0.013 | 0.1269 | No |
| 75 | Scaf8 | 9538 | 0.004 | 0.1143 | No |
| 76 | Celf2 | 9539 | 0.004 | 0.1145 | No |
| 77 | Pias3 | 9576 | 0.002 | 0.1127 | No |
| 78 | Atrx | 9592 | 0.002 | 0.1120 | No |
| 79 | Gcnt1 | 9724 | -0.002 | 0.1053 | No |
| 80 | Mta1 | 9961 | -0.010 | 0.0935 | No |
| 81 | Ldlr | 9971 | -0.010 | 0.0934 | No |
| 82 | Cited2 | 10017 | -0.012 | 0.0915 | No |
| 83 | Aggf1 | 10139 | -0.016 | 0.0859 | No |
| 84 | Togaram1 | 10290 | -0.020 | 0.0789 | No |
| 85 | Nipbl | 10435 | -0.026 | 0.0724 | No |
| 86 | Ythdc1 | 10470 | -0.027 | 0.0716 | No |
| 87 | Sipa1l1 | 10577 | -0.031 | 0.0672 | No |
| 88 | Nrp1 | 11124 | -0.049 | 0.0409 | No |
| 89 | Mrps31 | 11154 | -0.051 | 0.0412 | No |
| 90 | Col1a2 | 11239 | -0.053 | 0.0388 | No |
| 91 | Prkca | 11346 | -0.057 | 0.0354 | No |
| 92 | Lamc1 | 11537 | -0.064 | 0.0279 | No |
| 93 | Col1a1 | 11546 | -0.064 | 0.0297 | No |
| 94 | Wdr37 | 11744 | -0.072 | 0.0222 | No |
| 95 | Mmp16 | 11890 | -0.077 | 0.0175 | No |
| 96 | Ltbp1 | 12568 | -0.104 | -0.0137 | No |
| 97 | Fyn | 12766 | -0.111 | -0.0199 | No |
| 98 | Id1 | 13191 | -0.129 | -0.0371 | No |
| 99 | Dyrk1a | 13279 | -0.132 | -0.0368 | No |
| 100 | Plcb4 | 13618 | -0.148 | -0.0489 | No |
| 101 | Rnd3 | 14022 | -0.165 | -0.0637 | No |
| 102 | Atxn1 | 14046 | -0.167 | -0.0589 | No |
| 103 | Apbb2 | 14088 | -0.169 | -0.0549 | No |
| 104 | Cdon | 14180 | -0.174 | -0.0534 | No |
| 105 | Sri | 14202 | -0.174 | -0.0481 | No |
| 106 | Adora2b | 14367 | -0.182 | -0.0500 | No |
| 107 | Gja1 | 14764 | -0.199 | -0.0633 | No |
| 108 | Nfkb1 | 14922 | -0.208 | -0.0639 | No |
| 109 | Pparg | 15039 | -0.214 | -0.0621 | No |
| 110 | Spop | 15056 | -0.215 | -0.0552 | No |
| 111 | Cdkn1b | 15361 | -0.230 | -0.0626 | No |
| 112 | Mt1 | 15444 | -0.234 | -0.0584 | No |
| 113 | Grk5 | 15589 | -0.244 | -0.0570 | No |
| 114 | Nr3c1 | 15708 | -0.252 | -0.0540 | No |
| 115 | Rxra | 15711 | -0.252 | -0.0450 | No |
| 116 | Met | 16717 | -0.316 | -0.0854 | No |
| 117 | Atrn | 17287 | -0.359 | -0.1018 | No |
| 118 | Smad7 | 17396 | -0.369 | -0.0941 | No |
| 119 | Col5a2 | 17706 | -0.396 | -0.0958 | No |
| 120 | Pik3cd | 18013 | -0.431 | -0.0960 | No |
| 121 | Bhlhe40 | 18045 | -0.434 | -0.0819 | No |
| 122 | Ddah1 | 18086 | -0.439 | -0.0682 | No |
| 123 | Dmac2l | 18099 | -0.441 | -0.0529 | No |
| 124 | Anxa2 | 18883 | -0.563 | -0.0729 | No |
| 125 | Kcnma1 | 19163 | -0.653 | -0.0638 | No |
| 126 | Tjp1 | 19257 | -0.698 | -0.0434 | No |
| 127 | Vldlr | 19333 | -0.745 | -0.0204 | No |
| 128 | Tgfbr3 | 19403 | -0.819 | 0.0056 | No |