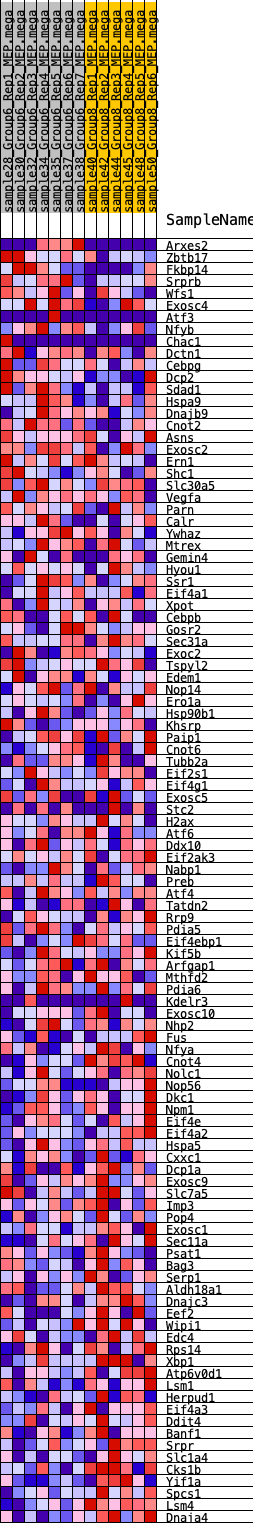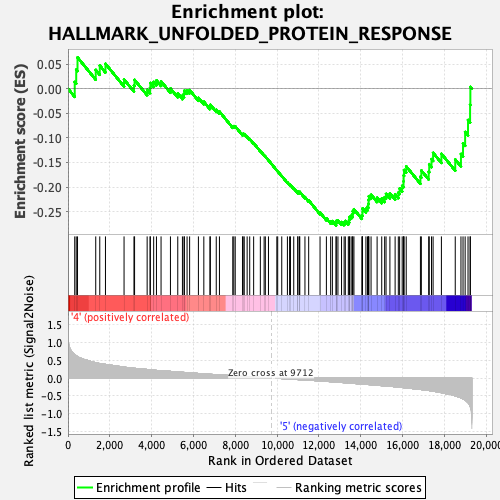
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group6_versus_Group8.MEP.mega_Pheno.cls #Group6_versus_Group8_repos |
| Phenotype | MEP.mega_Pheno.cls#Group6_versus_Group8_repos |
| Upregulated in class | 5 |
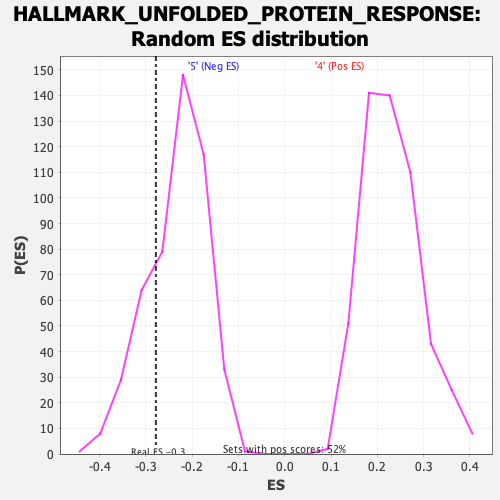
| GeneSet | HALLMARK_UNFOLDED_PROTEIN_RESPONSE |
| Enrichment Score (ES) | -0.27847692 |
| Normalized Enrichment Score (NES) | -1.1925504 |
| Nominal p-value | 0.23541667 |
| FDR q-value | 0.40635684 |
| FWER p-Value | 0.942 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Arxes2 | 322 | 0.663 | 0.0138 | No |
| 2 | Zbtb17 | 393 | 0.625 | 0.0389 | No |
| 3 | Fkbp14 | 455 | 0.604 | 0.0635 | No |
| 4 | Srprb | 1326 | 0.433 | 0.0382 | No |
| 5 | Wfs1 | 1519 | 0.408 | 0.0469 | No |
| 6 | Exosc4 | 1791 | 0.386 | 0.0506 | No |
| 7 | Atf3 | 2680 | 0.310 | 0.0187 | No |
| 8 | Nfyb | 3154 | 0.276 | 0.0067 | No |
| 9 | Chac1 | 3183 | 0.274 | 0.0179 | No |
| 10 | Dctn1 | 3784 | 0.244 | -0.0021 | No |
| 11 | Cebpg | 3923 | 0.234 | 0.0015 | No |
| 12 | Dcp2 | 3937 | 0.234 | 0.0116 | No |
| 13 | Sdad1 | 4096 | 0.224 | 0.0137 | No |
| 14 | Hspa9 | 4229 | 0.217 | 0.0168 | No |
| 15 | Dnajb9 | 4449 | 0.208 | 0.0150 | No |
| 16 | Cnot2 | 4898 | 0.190 | 0.0004 | No |
| 17 | Asns | 5248 | 0.173 | -0.0098 | No |
| 18 | Exosc2 | 5471 | 0.164 | -0.0138 | No |
| 19 | Ern1 | 5547 | 0.160 | -0.0103 | No |
| 20 | Shc1 | 5565 | 0.160 | -0.0039 | No |
| 21 | Slc30a5 | 5693 | 0.153 | -0.0034 | No |
| 22 | Vegfa | 5815 | 0.148 | -0.0029 | No |
| 23 | Parn | 6235 | 0.131 | -0.0187 | No |
| 24 | Calr | 6494 | 0.120 | -0.0266 | No |
| 25 | Ywhaz | 6786 | 0.108 | -0.0367 | No |
| 26 | Mtrex | 6805 | 0.107 | -0.0327 | No |
| 27 | Gemin4 | 7088 | 0.098 | -0.0429 | No |
| 28 | Hyou1 | 7241 | 0.091 | -0.0466 | No |
| 29 | Ssr1 | 7878 | 0.068 | -0.0766 | No |
| 30 | Eif4a1 | 7928 | 0.065 | -0.0762 | No |
| 31 | Xpot | 7995 | 0.063 | -0.0767 | No |
| 32 | Cebpb | 8342 | 0.049 | -0.0925 | No |
| 33 | Gosr2 | 8372 | 0.048 | -0.0918 | No |
| 34 | Sec31a | 8426 | 0.046 | -0.0924 | No |
| 35 | Exoc2 | 8565 | 0.041 | -0.0978 | No |
| 36 | Tspyl2 | 8688 | 0.036 | -0.1025 | No |
| 37 | Edem1 | 8876 | 0.029 | -0.1108 | No |
| 38 | Nop14 | 9196 | 0.018 | -0.1266 | No |
| 39 | Ero1a | 9377 | 0.011 | -0.1355 | No |
| 40 | Hsp90b1 | 9435 | 0.010 | -0.1380 | No |
| 41 | Khsrp | 9585 | 0.004 | -0.1456 | No |
| 42 | Paip1 | 9985 | -0.003 | -0.1662 | No |
| 43 | Cnot6 | 10026 | -0.004 | -0.1681 | No |
| 44 | Tubb2a | 10220 | -0.011 | -0.1777 | No |
| 45 | Eif2s1 | 10490 | -0.021 | -0.1907 | No |
| 46 | Eif4g1 | 10595 | -0.025 | -0.1950 | No |
| 47 | Exosc5 | 10637 | -0.026 | -0.1960 | No |
| 48 | Stc2 | 10793 | -0.031 | -0.2026 | No |
| 49 | H2ax | 10988 | -0.038 | -0.2109 | No |
| 50 | Atf6 | 10989 | -0.038 | -0.2092 | No |
| 51 | Ddx10 | 11074 | -0.042 | -0.2116 | No |
| 52 | Eif2ak3 | 11078 | -0.042 | -0.2099 | No |
| 53 | Nabp1 | 11329 | -0.050 | -0.2206 | No |
| 54 | Preb | 11507 | -0.056 | -0.2272 | No |
| 55 | Atf4 | 12051 | -0.076 | -0.2520 | No |
| 56 | Tatdn2 | 12354 | -0.089 | -0.2636 | No |
| 57 | Rrp9 | 12558 | -0.097 | -0.2698 | No |
| 58 | Pdia5 | 12640 | -0.101 | -0.2693 | No |
| 59 | Eif4ebp1 | 12817 | -0.108 | -0.2735 | Yes |
| 60 | Kif5b | 12824 | -0.108 | -0.2688 | Yes |
| 61 | Arfgap1 | 12906 | -0.112 | -0.2679 | Yes |
| 62 | Mthfd2 | 13078 | -0.120 | -0.2712 | Yes |
| 63 | Pdia6 | 13202 | -0.124 | -0.2719 | Yes |
| 64 | Kdelr3 | 13259 | -0.127 | -0.2690 | Yes |
| 65 | Exosc10 | 13403 | -0.133 | -0.2703 | Yes |
| 66 | Nhp2 | 13446 | -0.135 | -0.2663 | Yes |
| 67 | Fus | 13463 | -0.136 | -0.2608 | Yes |
| 68 | Nfya | 13538 | -0.139 | -0.2583 | Yes |
| 69 | Cnot4 | 13605 | -0.142 | -0.2552 | Yes |
| 70 | Nolc1 | 13617 | -0.143 | -0.2491 | Yes |
| 71 | Nop56 | 13671 | -0.146 | -0.2452 | Yes |
| 72 | Dkc1 | 14050 | -0.164 | -0.2573 | Yes |
| 73 | Npm1 | 14079 | -0.165 | -0.2512 | Yes |
| 74 | Eif4e | 14080 | -0.165 | -0.2436 | Yes |
| 75 | Eif4a2 | 14244 | -0.172 | -0.2441 | Yes |
| 76 | Hspa5 | 14319 | -0.175 | -0.2399 | Yes |
| 77 | Cxxc1 | 14364 | -0.177 | -0.2340 | Yes |
| 78 | Dcp1a | 14372 | -0.178 | -0.2262 | Yes |
| 79 | Exosc9 | 14386 | -0.178 | -0.2187 | Yes |
| 80 | Slc7a5 | 14490 | -0.182 | -0.2157 | Yes |
| 81 | Imp3 | 14779 | -0.197 | -0.2216 | Yes |
| 82 | Pop4 | 15000 | -0.209 | -0.2234 | Yes |
| 83 | Exosc1 | 15140 | -0.216 | -0.2207 | Yes |
| 84 | Sec11a | 15206 | -0.220 | -0.2139 | Yes |
| 85 | Psat1 | 15391 | -0.230 | -0.2129 | Yes |
| 86 | Bag3 | 15640 | -0.240 | -0.2148 | Yes |
| 87 | Serp1 | 15793 | -0.247 | -0.2113 | Yes |
| 88 | Aldh18a1 | 15856 | -0.251 | -0.2030 | Yes |
| 89 | Dnajc3 | 15978 | -0.259 | -0.1973 | Yes |
| 90 | Eef2 | 16042 | -0.264 | -0.1885 | Yes |
| 91 | Wipi1 | 16048 | -0.264 | -0.1765 | Yes |
| 92 | Edc4 | 16068 | -0.265 | -0.1653 | Yes |
| 93 | Rps14 | 16171 | -0.272 | -0.1581 | Yes |
| 94 | Xbp1 | 16850 | -0.316 | -0.1789 | Yes |
| 95 | Atp6v0d1 | 16894 | -0.319 | -0.1664 | Yes |
| 96 | Lsm1 | 17246 | -0.345 | -0.1688 | Yes |
| 97 | Herpud1 | 17271 | -0.347 | -0.1541 | Yes |
| 98 | Eif4a3 | 17380 | -0.357 | -0.1433 | Yes |
| 99 | Ddit4 | 17456 | -0.363 | -0.1305 | Yes |
| 100 | Banf1 | 17853 | -0.406 | -0.1324 | Yes |
| 101 | Srpr | 18515 | -0.495 | -0.1440 | Yes |
| 102 | Slc1a4 | 18788 | -0.552 | -0.1327 | Yes |
| 103 | Cks1b | 18887 | -0.580 | -0.1111 | Yes |
| 104 | Yif1a | 18988 | -0.615 | -0.0880 | Yes |
| 105 | Spcs1 | 19126 | -0.684 | -0.0636 | Yes |
| 106 | Lsm4 | 19225 | -0.780 | -0.0328 | Yes |
| 107 | Dnaja4 | 19238 | -0.804 | 0.0036 | Yes |