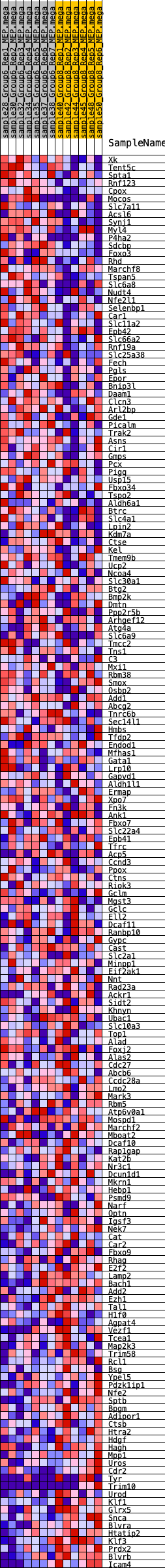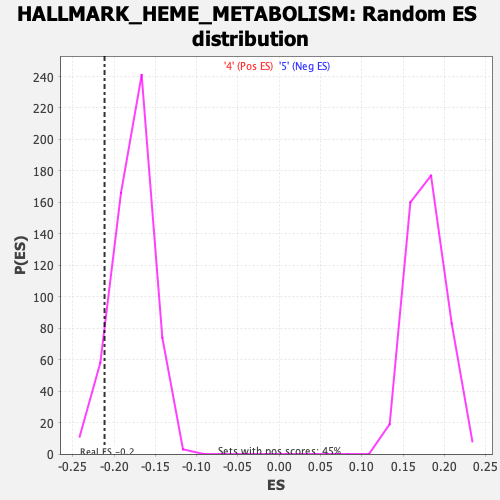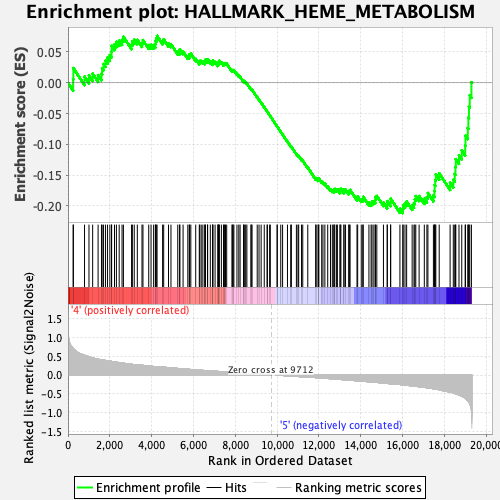
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group6_versus_Group8.MEP.mega_Pheno.cls #Group6_versus_Group8_repos |
| Phenotype | MEP.mega_Pheno.cls#Group6_versus_Group8_repos |
| Upregulated in class | 5 |
| GeneSet | HALLMARK_HEME_METABOLISM |
| Enrichment Score (ES) | -0.21163249 |
| Normalized Enrichment Score (NES) | -1.1977795 |
| Nominal p-value | 0.07594936 |
| FDR q-value | 0.42130303 |
| FWER p-Value | 0.936 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Xk | 246 | 0.703 | 0.0059 | No |
| 2 | Tent5c | 257 | 0.699 | 0.0239 | No |
| 3 | Spta1 | 790 | 0.523 | 0.0101 | No |
| 4 | Rnf123 | 1003 | 0.485 | 0.0119 | No |
| 5 | Cpox | 1179 | 0.453 | 0.0148 | No |
| 6 | Mocos | 1439 | 0.421 | 0.0125 | No |
| 7 | Slc7a11 | 1600 | 0.398 | 0.0147 | No |
| 8 | Acsl6 | 1631 | 0.395 | 0.0237 | No |
| 9 | Synj1 | 1700 | 0.391 | 0.0305 | No |
| 10 | Myl4 | 1795 | 0.386 | 0.0359 | No |
| 11 | P4ha2 | 1891 | 0.380 | 0.0411 | No |
| 12 | Sdcbp | 2001 | 0.368 | 0.0452 | No |
| 13 | Foxo3 | 2081 | 0.360 | 0.0506 | No |
| 14 | Rhd | 2086 | 0.360 | 0.0600 | No |
| 15 | Marchf8 | 2223 | 0.350 | 0.0622 | No |
| 16 | Tspan5 | 2314 | 0.340 | 0.0666 | No |
| 17 | Slc6a8 | 2447 | 0.330 | 0.0685 | No |
| 18 | Nudt4 | 2584 | 0.319 | 0.0698 | No |
| 19 | Nfe2l1 | 2650 | 0.312 | 0.0748 | No |
| 20 | Selenbp1 | 3039 | 0.284 | 0.0620 | No |
| 21 | Car1 | 3077 | 0.280 | 0.0676 | No |
| 22 | Slc11a2 | 3167 | 0.275 | 0.0702 | No |
| 23 | Epb42 | 3314 | 0.268 | 0.0697 | No |
| 24 | Slc66a2 | 3537 | 0.257 | 0.0650 | No |
| 25 | Rnf19a | 3584 | 0.255 | 0.0693 | No |
| 26 | Slc25a38 | 3860 | 0.239 | 0.0613 | No |
| 27 | Fech | 3971 | 0.232 | 0.0617 | No |
| 28 | Pgls | 4094 | 0.225 | 0.0613 | No |
| 29 | Epor | 4177 | 0.219 | 0.0629 | No |
| 30 | Bnip3l | 4187 | 0.219 | 0.0683 | No |
| 31 | Daam1 | 4222 | 0.217 | 0.0723 | No |
| 32 | Clcn3 | 4258 | 0.215 | 0.0762 | No |
| 33 | Arl2bp | 4523 | 0.207 | 0.0679 | No |
| 34 | Gde1 | 4575 | 0.204 | 0.0706 | No |
| 35 | Picalm | 4805 | 0.195 | 0.0638 | No |
| 36 | Trak2 | 4923 | 0.188 | 0.0627 | No |
| 37 | Asns | 5248 | 0.173 | 0.0504 | No |
| 38 | Cir1 | 5337 | 0.170 | 0.0503 | No |
| 39 | Gmps | 5348 | 0.169 | 0.0543 | No |
| 40 | Pcx | 5502 | 0.162 | 0.0507 | No |
| 41 | Pigq | 5729 | 0.152 | 0.0429 | No |
| 42 | Usp15 | 5797 | 0.149 | 0.0433 | No |
| 43 | Fbxo34 | 5814 | 0.148 | 0.0465 | No |
| 44 | Tspo2 | 5872 | 0.146 | 0.0474 | No |
| 45 | Aldh6a1 | 6106 | 0.137 | 0.0388 | No |
| 46 | Btrc | 6277 | 0.129 | 0.0334 | No |
| 47 | Slc4a1 | 6300 | 0.128 | 0.0356 | No |
| 48 | Lpin2 | 6369 | 0.126 | 0.0354 | No |
| 49 | Kdm7a | 6448 | 0.122 | 0.0346 | No |
| 50 | Ctse | 6539 | 0.118 | 0.0330 | No |
| 51 | Kel | 6548 | 0.118 | 0.0357 | No |
| 52 | Tmem9b | 6572 | 0.116 | 0.0376 | No |
| 53 | Ucp2 | 6672 | 0.113 | 0.0355 | No |
| 54 | Ncoa4 | 6675 | 0.113 | 0.0384 | No |
| 55 | Slc30a1 | 6804 | 0.107 | 0.0346 | No |
| 56 | Btg2 | 6912 | 0.104 | 0.0317 | No |
| 57 | Bmp2k | 6918 | 0.104 | 0.0342 | No |
| 58 | Dmtn | 6929 | 0.103 | 0.0365 | No |
| 59 | Ppp2r5b | 7032 | 0.100 | 0.0338 | No |
| 60 | Arhgef12 | 7162 | 0.095 | 0.0296 | No |
| 61 | Atg4a | 7180 | 0.094 | 0.0312 | No |
| 62 | Slc6a9 | 7202 | 0.093 | 0.0326 | No |
| 63 | Tmcc2 | 7215 | 0.093 | 0.0344 | No |
| 64 | Tns1 | 7235 | 0.092 | 0.0359 | No |
| 65 | C3 | 7333 | 0.089 | 0.0332 | No |
| 66 | Mxi1 | 7437 | 0.085 | 0.0301 | No |
| 67 | Rbm38 | 7460 | 0.084 | 0.0311 | No |
| 68 | Smox | 7491 | 0.083 | 0.0318 | No |
| 69 | Osbp2 | 7531 | 0.081 | 0.0319 | No |
| 70 | Add1 | 7588 | 0.079 | 0.0311 | No |
| 71 | Abcg2 | 7841 | 0.069 | 0.0197 | No |
| 72 | Tnrc6b | 7859 | 0.069 | 0.0207 | No |
| 73 | Sec14l1 | 7895 | 0.067 | 0.0206 | No |
| 74 | Hmbs | 7931 | 0.065 | 0.0206 | No |
| 75 | Tfdp2 | 8078 | 0.059 | 0.0145 | No |
| 76 | Endod1 | 8167 | 0.056 | 0.0114 | No |
| 77 | Mfhas1 | 8228 | 0.053 | 0.0097 | No |
| 78 | Gata1 | 8393 | 0.047 | 0.0024 | No |
| 79 | Lrp10 | 8401 | 0.046 | 0.0032 | No |
| 80 | Gapvd1 | 8449 | 0.045 | 0.0020 | No |
| 81 | Aldh1l1 | 8497 | 0.043 | 0.0006 | No |
| 82 | Ermap | 8552 | 0.041 | -0.0011 | No |
| 83 | Xpo7 | 8747 | 0.034 | -0.0103 | No |
| 84 | Fn3k | 8760 | 0.034 | -0.0101 | No |
| 85 | Ank1 | 8800 | 0.032 | -0.0113 | No |
| 86 | Fbxo7 | 9047 | 0.024 | -0.0235 | No |
| 87 | Slc22a4 | 9133 | 0.020 | -0.0274 | No |
| 88 | Epb41 | 9234 | 0.017 | -0.0322 | No |
| 89 | Tfrc | 9383 | 0.011 | -0.0396 | No |
| 90 | Acp5 | 9518 | 0.007 | -0.0464 | No |
| 91 | Ccnd3 | 9533 | 0.006 | -0.0470 | No |
| 92 | Ppox | 9640 | 0.002 | -0.0525 | No |
| 93 | Ctns | 9672 | 0.001 | -0.0541 | No |
| 94 | Riok3 | 9999 | -0.003 | -0.0710 | No |
| 95 | Gclm | 10004 | -0.003 | -0.0712 | No |
| 96 | Mgst3 | 10172 | -0.009 | -0.0797 | No |
| 97 | Gclc | 10260 | -0.012 | -0.0839 | No |
| 98 | Ell2 | 10493 | -0.021 | -0.0955 | No |
| 99 | Dcaf11 | 10648 | -0.026 | -0.1028 | No |
| 100 | Ranbp10 | 10680 | -0.027 | -0.1037 | No |
| 101 | Gypc | 10920 | -0.036 | -0.1153 | No |
| 102 | Cast | 10994 | -0.038 | -0.1181 | No |
| 103 | Slc2a1 | 11030 | -0.040 | -0.1188 | No |
| 104 | Minpp1 | 11165 | -0.044 | -0.1246 | No |
| 105 | Eif2ak1 | 11224 | -0.046 | -0.1265 | No |
| 106 | Nnt | 11464 | -0.055 | -0.1375 | No |
| 107 | Rad23a | 11841 | -0.068 | -0.1553 | No |
| 108 | Ackr1 | 11865 | -0.069 | -0.1547 | No |
| 109 | Sidt2 | 11958 | -0.072 | -0.1576 | No |
| 110 | Khnyn | 11968 | -0.072 | -0.1561 | No |
| 111 | Ubac1 | 11995 | -0.073 | -0.1555 | No |
| 112 | Slc10a3 | 12130 | -0.079 | -0.1604 | No |
| 113 | Top1 | 12190 | -0.081 | -0.1614 | No |
| 114 | Alad | 12281 | -0.086 | -0.1638 | No |
| 115 | Foxj2 | 12417 | -0.091 | -0.1684 | No |
| 116 | Alas2 | 12550 | -0.097 | -0.1727 | No |
| 117 | Cdc27 | 12638 | -0.101 | -0.1746 | No |
| 118 | Abcb6 | 12708 | -0.104 | -0.1754 | No |
| 119 | Ccdc28a | 12724 | -0.104 | -0.1734 | No |
| 120 | Lmo2 | 12756 | -0.106 | -0.1723 | No |
| 121 | Mark3 | 12842 | -0.109 | -0.1738 | No |
| 122 | Rbm5 | 12880 | -0.111 | -0.1728 | No |
| 123 | Atp6v0a1 | 13007 | -0.116 | -0.1763 | No |
| 124 | Mospd1 | 13010 | -0.116 | -0.1733 | No |
| 125 | Marchf2 | 13048 | -0.118 | -0.1721 | No |
| 126 | Mboat2 | 13193 | -0.124 | -0.1763 | No |
| 127 | Dcaf10 | 13194 | -0.124 | -0.1730 | No |
| 128 | Rap1gap | 13270 | -0.127 | -0.1735 | No |
| 129 | Kat2b | 13419 | -0.134 | -0.1777 | No |
| 130 | Nr3c1 | 13453 | -0.136 | -0.1758 | No |
| 131 | Dcun1d1 | 13487 | -0.137 | -0.1739 | No |
| 132 | Mkrn1 | 13817 | -0.153 | -0.1870 | No |
| 133 | Hebp1 | 13850 | -0.154 | -0.1846 | No |
| 134 | Psmd9 | 14021 | -0.162 | -0.1891 | No |
| 135 | Narf | 14093 | -0.166 | -0.1884 | No |
| 136 | Optn | 14116 | -0.167 | -0.1851 | No |
| 137 | Igsf3 | 14388 | -0.178 | -0.1946 | No |
| 138 | Nek7 | 14492 | -0.183 | -0.1951 | No |
| 139 | Cat | 14537 | -0.185 | -0.1925 | No |
| 140 | Car2 | 14624 | -0.189 | -0.1919 | No |
| 141 | Fbxo9 | 14693 | -0.193 | -0.1904 | No |
| 142 | Rhag | 14698 | -0.193 | -0.1854 | No |
| 143 | E2f2 | 14765 | -0.196 | -0.1837 | No |
| 144 | Lamp2 | 15082 | -0.213 | -0.1945 | No |
| 145 | Bach1 | 15264 | -0.223 | -0.1980 | No |
| 146 | Add2 | 15265 | -0.223 | -0.1921 | No |
| 147 | Ezh1 | 15422 | -0.231 | -0.1941 | No |
| 148 | Tal1 | 15429 | -0.232 | -0.1882 | No |
| 149 | H1f0 | 15870 | -0.252 | -0.2045 | No |
| 150 | Agpat4 | 16007 | -0.261 | -0.2047 | Yes |
| 151 | Vezf1 | 16024 | -0.263 | -0.1985 | Yes |
| 152 | Tcea1 | 16112 | -0.268 | -0.1959 | Yes |
| 153 | Map2k3 | 16191 | -0.273 | -0.1928 | Yes |
| 154 | Trim58 | 16454 | -0.290 | -0.1987 | Yes |
| 155 | Rcl1 | 16542 | -0.296 | -0.1954 | Yes |
| 156 | Bsg | 16581 | -0.298 | -0.1894 | Yes |
| 157 | Ypel5 | 16628 | -0.301 | -0.1838 | Yes |
| 158 | Pdzk1ip1 | 16787 | -0.311 | -0.1838 | Yes |
| 159 | Nfe2 | 17036 | -0.327 | -0.1881 | Yes |
| 160 | Sptb | 17160 | -0.338 | -0.1855 | Yes |
| 161 | Bpgm | 17208 | -0.341 | -0.1789 | Yes |
| 162 | Adipor1 | 17471 | -0.365 | -0.1828 | Yes |
| 163 | Ctsb | 17522 | -0.369 | -0.1756 | Yes |
| 164 | Htra2 | 17530 | -0.370 | -0.1661 | Yes |
| 165 | Hdgf | 17558 | -0.373 | -0.1576 | Yes |
| 166 | Hagh | 17586 | -0.376 | -0.1490 | Yes |
| 167 | Mpp1 | 17748 | -0.392 | -0.1469 | Yes |
| 168 | Uros | 18269 | -0.453 | -0.1621 | Yes |
| 169 | Cdr2 | 18422 | -0.476 | -0.1573 | Yes |
| 170 | Tyr | 18492 | -0.490 | -0.1479 | Yes |
| 171 | Trim10 | 18523 | -0.496 | -0.1362 | Yes |
| 172 | Urod | 18542 | -0.500 | -0.1238 | Yes |
| 173 | Klf1 | 18694 | -0.528 | -0.1177 | Yes |
| 174 | Glrx5 | 18826 | -0.561 | -0.1096 | Yes |
| 175 | Snca | 18990 | -0.615 | -0.1017 | Yes |
| 176 | Blvra | 19003 | -0.621 | -0.0858 | Yes |
| 177 | Htatip2 | 19114 | -0.679 | -0.0735 | Yes |
| 178 | Klf3 | 19143 | -0.695 | -0.0564 | Yes |
| 179 | Prdx2 | 19171 | -0.716 | -0.0388 | Yes |
| 180 | Blvrb | 19208 | -0.758 | -0.0205 | Yes |
| 181 | Icam4 | 19288 | -0.961 | 0.0010 | Yes |