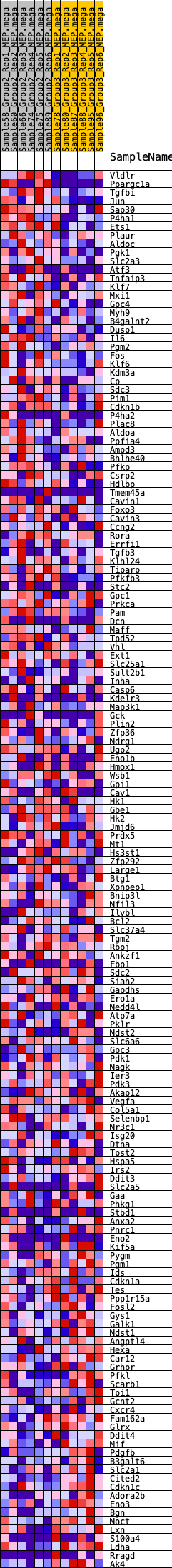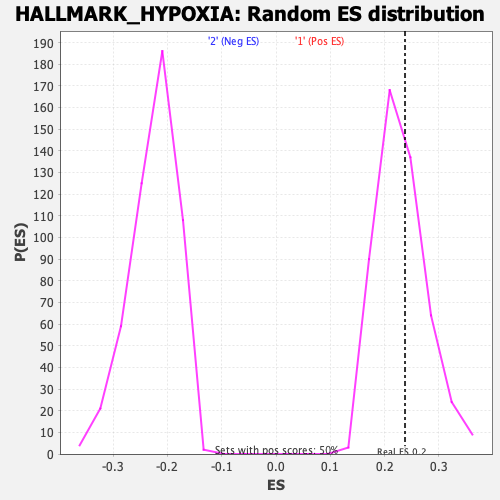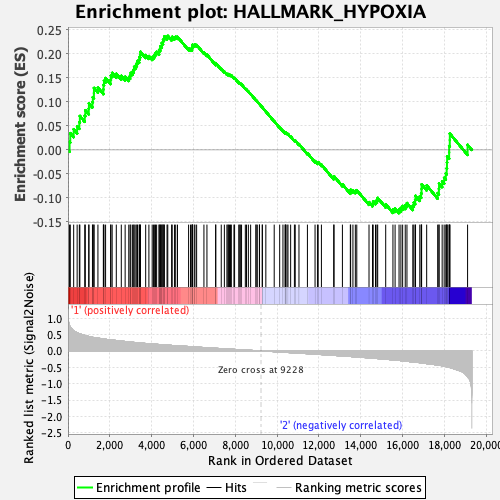
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group2_versus_Group3.MEP.mega_Pheno.cls #Group2_versus_Group3_repos |
| Phenotype | MEP.mega_Pheno.cls#Group2_versus_Group3_repos |
| Upregulated in class | 1 |
| GeneSet | HALLMARK_HYPOXIA |
| Enrichment Score (ES) | 0.23745911 |
| Normalized Enrichment Score (NES) | 1.0241809 |
| Nominal p-value | 0.4040404 |
| FDR q-value | 0.8800271 |
| FWER p-Value | 0.993 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Vldlr | 75 | 0.776 | 0.0169 | Yes |
| 2 | Ppargc1a | 109 | 0.733 | 0.0348 | Yes |
| 3 | Tgfbi | 271 | 0.617 | 0.0430 | Yes |
| 4 | Jun | 437 | 0.555 | 0.0493 | Yes |
| 5 | Sap30 | 542 | 0.526 | 0.0580 | Yes |
| 6 | P4ha1 | 567 | 0.522 | 0.0707 | Yes |
| 7 | Ets1 | 801 | 0.474 | 0.0712 | Yes |
| 8 | Plaur | 827 | 0.469 | 0.0825 | Yes |
| 9 | Aldoc | 993 | 0.440 | 0.0857 | Yes |
| 10 | Pgk1 | 1001 | 0.439 | 0.0971 | Yes |
| 11 | Slc2a3 | 1169 | 0.416 | 0.0996 | Yes |
| 12 | Atf3 | 1187 | 0.414 | 0.1098 | Yes |
| 13 | Tnfaip3 | 1239 | 0.409 | 0.1181 | Yes |
| 14 | Klf7 | 1241 | 0.409 | 0.1290 | Yes |
| 15 | Mxi1 | 1431 | 0.391 | 0.1296 | Yes |
| 16 | Gpc4 | 1690 | 0.366 | 0.1260 | Yes |
| 17 | Myh9 | 1697 | 0.365 | 0.1355 | Yes |
| 18 | B4galnt2 | 1721 | 0.362 | 0.1440 | Yes |
| 19 | Dusp1 | 1797 | 0.357 | 0.1496 | Yes |
| 20 | Il6 | 2038 | 0.339 | 0.1462 | Yes |
| 21 | Pgm2 | 2050 | 0.338 | 0.1547 | Yes |
| 22 | Fos | 2111 | 0.332 | 0.1604 | Yes |
| 23 | Klf6 | 2309 | 0.317 | 0.1587 | Yes |
| 24 | Kdm3a | 2548 | 0.302 | 0.1543 | Yes |
| 25 | Cp | 2732 | 0.288 | 0.1525 | Yes |
| 26 | Sdc3 | 2906 | 0.276 | 0.1509 | Yes |
| 27 | Pim1 | 2968 | 0.272 | 0.1550 | Yes |
| 28 | Cdkn1b | 3003 | 0.270 | 0.1604 | Yes |
| 29 | P4ha2 | 3089 | 0.265 | 0.1631 | Yes |
| 30 | Plac8 | 3144 | 0.261 | 0.1673 | Yes |
| 31 | Aldoa | 3164 | 0.260 | 0.1733 | Yes |
| 32 | Ppfia4 | 3244 | 0.255 | 0.1760 | Yes |
| 33 | Ampd3 | 3290 | 0.252 | 0.1804 | Yes |
| 34 | Bhlhe40 | 3324 | 0.250 | 0.1854 | Yes |
| 35 | Pfkp | 3402 | 0.245 | 0.1879 | Yes |
| 36 | Csrp2 | 3418 | 0.244 | 0.1937 | Yes |
| 37 | Hdlbp | 3452 | 0.242 | 0.1985 | Yes |
| 38 | Tmem45a | 3466 | 0.241 | 0.2043 | Yes |
| 39 | Cavin1 | 3714 | 0.228 | 0.1975 | Yes |
| 40 | Foxo3 | 3868 | 0.219 | 0.1954 | Yes |
| 41 | Cavin3 | 4025 | 0.212 | 0.1929 | Yes |
| 42 | Ccng2 | 4091 | 0.210 | 0.1952 | Yes |
| 43 | Rora | 4143 | 0.207 | 0.1981 | Yes |
| 44 | Errfi1 | 4182 | 0.205 | 0.2016 | Yes |
| 45 | Tgfb3 | 4237 | 0.203 | 0.2042 | Yes |
| 46 | Klhl24 | 4354 | 0.197 | 0.2034 | Yes |
| 47 | Tiparp | 4361 | 0.196 | 0.2084 | Yes |
| 48 | Pfkfb3 | 4409 | 0.194 | 0.2111 | Yes |
| 49 | Stc2 | 4418 | 0.194 | 0.2159 | Yes |
| 50 | Gpc1 | 4485 | 0.190 | 0.2176 | Yes |
| 51 | Prkca | 4486 | 0.190 | 0.2226 | Yes |
| 52 | Pam | 4542 | 0.186 | 0.2248 | Yes |
| 53 | Dcn | 4547 | 0.186 | 0.2296 | Yes |
| 54 | Maff | 4592 | 0.184 | 0.2322 | Yes |
| 55 | Tpd52 | 4606 | 0.183 | 0.2364 | Yes |
| 56 | Vhl | 4736 | 0.178 | 0.2345 | Yes |
| 57 | Ext1 | 4770 | 0.176 | 0.2375 | Yes |
| 58 | Slc25a1 | 4955 | 0.169 | 0.2324 | No |
| 59 | Sult2b1 | 4980 | 0.167 | 0.2356 | No |
| 60 | Inha | 5090 | 0.162 | 0.2343 | No |
| 61 | Casp6 | 5149 | 0.161 | 0.2355 | No |
| 62 | Kdelr3 | 5234 | 0.157 | 0.2354 | No |
| 63 | Map3k1 | 5766 | 0.135 | 0.2113 | No |
| 64 | Gck | 5858 | 0.131 | 0.2100 | No |
| 65 | Plin2 | 5927 | 0.128 | 0.2099 | No |
| 66 | Zfp36 | 5932 | 0.128 | 0.2131 | No |
| 67 | Ndrg1 | 5950 | 0.127 | 0.2156 | No |
| 68 | Ugp2 | 5952 | 0.127 | 0.2190 | No |
| 69 | Eno1b | 6034 | 0.123 | 0.2180 | No |
| 70 | Hmox1 | 6069 | 0.121 | 0.2195 | No |
| 71 | Wsb1 | 6150 | 0.118 | 0.2185 | No |
| 72 | Gpi1 | 6499 | 0.103 | 0.2031 | No |
| 73 | Cav1 | 6643 | 0.098 | 0.1982 | No |
| 74 | Hk1 | 7061 | 0.081 | 0.1786 | No |
| 75 | Gbe1 | 7063 | 0.081 | 0.1807 | No |
| 76 | Hk2 | 7327 | 0.071 | 0.1689 | No |
| 77 | Jmjd6 | 7476 | 0.066 | 0.1629 | No |
| 78 | Prdx5 | 7598 | 0.060 | 0.1582 | No |
| 79 | Mt1 | 7647 | 0.059 | 0.1573 | No |
| 80 | Hs3st1 | 7667 | 0.058 | 0.1579 | No |
| 81 | Zfp292 | 7723 | 0.056 | 0.1565 | No |
| 82 | Large1 | 7768 | 0.055 | 0.1557 | No |
| 83 | Btg1 | 7809 | 0.053 | 0.1550 | No |
| 84 | Xpnpep1 | 7935 | 0.049 | 0.1498 | No |
| 85 | Bnip3l | 7965 | 0.047 | 0.1496 | No |
| 86 | Nfil3 | 8164 | 0.040 | 0.1403 | No |
| 87 | Ilvbl | 8213 | 0.037 | 0.1388 | No |
| 88 | Bcl2 | 8238 | 0.036 | 0.1385 | No |
| 89 | Slc37a4 | 8298 | 0.034 | 0.1363 | No |
| 90 | Tgm2 | 8478 | 0.028 | 0.1277 | No |
| 91 | Rbpj | 8534 | 0.026 | 0.1256 | No |
| 92 | Ankzf1 | 8623 | 0.022 | 0.1216 | No |
| 93 | Fbp1 | 8732 | 0.019 | 0.1164 | No |
| 94 | Sdc2 | 8986 | 0.009 | 0.1035 | No |
| 95 | Siah2 | 8991 | 0.009 | 0.1035 | No |
| 96 | Gapdhs | 9051 | 0.007 | 0.1006 | No |
| 97 | Ero1a | 9150 | 0.003 | 0.0956 | No |
| 98 | Nedd4l | 9156 | 0.003 | 0.0954 | No |
| 99 | Atp7a | 9289 | -0.000 | 0.0885 | No |
| 100 | Pklr | 9292 | -0.000 | 0.0884 | No |
| 101 | Ndst2 | 9459 | -0.006 | 0.0799 | No |
| 102 | Slc6a6 | 9859 | -0.020 | 0.0596 | No |
| 103 | Gpc3 | 10121 | -0.030 | 0.0467 | No |
| 104 | Pdk1 | 10277 | -0.036 | 0.0396 | No |
| 105 | Nagk | 10376 | -0.040 | 0.0356 | No |
| 106 | Ier3 | 10398 | -0.041 | 0.0356 | No |
| 107 | Pdk3 | 10403 | -0.041 | 0.0365 | No |
| 108 | Akap12 | 10467 | -0.044 | 0.0344 | No |
| 109 | Vegfa | 10517 | -0.046 | 0.0330 | No |
| 110 | Col5a1 | 10645 | -0.051 | 0.0278 | No |
| 111 | Selenbp1 | 10825 | -0.057 | 0.0199 | No |
| 112 | Nr3c1 | 10864 | -0.059 | 0.0195 | No |
| 113 | Isg20 | 11043 | -0.066 | 0.0120 | No |
| 114 | Dtna | 11447 | -0.080 | -0.0069 | No |
| 115 | Tpst2 | 11812 | -0.096 | -0.0233 | No |
| 116 | Hspa5 | 11927 | -0.101 | -0.0266 | No |
| 117 | Irs2 | 11962 | -0.103 | -0.0256 | No |
| 118 | Ddit3 | 12116 | -0.109 | -0.0307 | No |
| 119 | Slc2a5 | 12704 | -0.134 | -0.0577 | No |
| 120 | Gaa | 12721 | -0.135 | -0.0550 | No |
| 121 | Phkg1 | 13120 | -0.153 | -0.0716 | No |
| 122 | Stbd1 | 13499 | -0.168 | -0.0869 | No |
| 123 | Anxa2 | 13513 | -0.169 | -0.0830 | No |
| 124 | Pnrc1 | 13623 | -0.174 | -0.0841 | No |
| 125 | Eno2 | 13738 | -0.179 | -0.0852 | No |
| 126 | Kif5a | 13807 | -0.181 | -0.0839 | No |
| 127 | Pygm | 14392 | -0.210 | -0.1088 | No |
| 128 | Pgm1 | 14559 | -0.218 | -0.1116 | No |
| 129 | Ids | 14591 | -0.220 | -0.1073 | No |
| 130 | Cdkn1a | 14702 | -0.226 | -0.1070 | No |
| 131 | Tes | 14770 | -0.230 | -0.1043 | No |
| 132 | Ppp1r15a | 14807 | -0.231 | -0.1000 | No |
| 133 | Fosl2 | 15190 | -0.253 | -0.1132 | No |
| 134 | Gys1 | 15536 | -0.271 | -0.1239 | No |
| 135 | Galk1 | 15639 | -0.276 | -0.1218 | No |
| 136 | Ndst1 | 15828 | -0.288 | -0.1239 | No |
| 137 | Angptl4 | 15917 | -0.296 | -0.1206 | No |
| 138 | Hexa | 16000 | -0.302 | -0.1168 | No |
| 139 | Car12 | 16124 | -0.310 | -0.1149 | No |
| 140 | Grhpr | 16206 | -0.315 | -0.1107 | No |
| 141 | Pfkl | 16486 | -0.334 | -0.1163 | No |
| 142 | Scarb1 | 16534 | -0.337 | -0.1097 | No |
| 143 | Tpi1 | 16599 | -0.341 | -0.1039 | No |
| 144 | Gcnt2 | 16616 | -0.342 | -0.0956 | No |
| 145 | Cxcr4 | 16820 | -0.357 | -0.0966 | No |
| 146 | Fam162a | 16885 | -0.362 | -0.0902 | No |
| 147 | Glrx | 16903 | -0.363 | -0.0814 | No |
| 148 | Ddit4 | 16906 | -0.364 | -0.0717 | No |
| 149 | Mif | 17149 | -0.385 | -0.0740 | No |
| 150 | Pdgfb | 17675 | -0.430 | -0.0899 | No |
| 151 | B3galt6 | 17734 | -0.436 | -0.0812 | No |
| 152 | Slc2a1 | 17741 | -0.436 | -0.0699 | No |
| 153 | Cited2 | 17889 | -0.452 | -0.0654 | No |
| 154 | Cdkn1c | 17985 | -0.464 | -0.0579 | No |
| 155 | Adora2b | 18064 | -0.475 | -0.0493 | No |
| 156 | Eno3 | 18105 | -0.482 | -0.0384 | No |
| 157 | Bgn | 18119 | -0.483 | -0.0261 | No |
| 158 | Noct | 18127 | -0.485 | -0.0135 | No |
| 159 | Lxn | 18223 | -0.500 | -0.0050 | No |
| 160 | S100a4 | 18225 | -0.500 | 0.0083 | No |
| 161 | Ldha | 18254 | -0.504 | 0.0204 | No |
| 162 | Rragd | 18256 | -0.504 | 0.0339 | No |
| 163 | Ak4 | 19105 | -0.782 | 0.0106 | No |