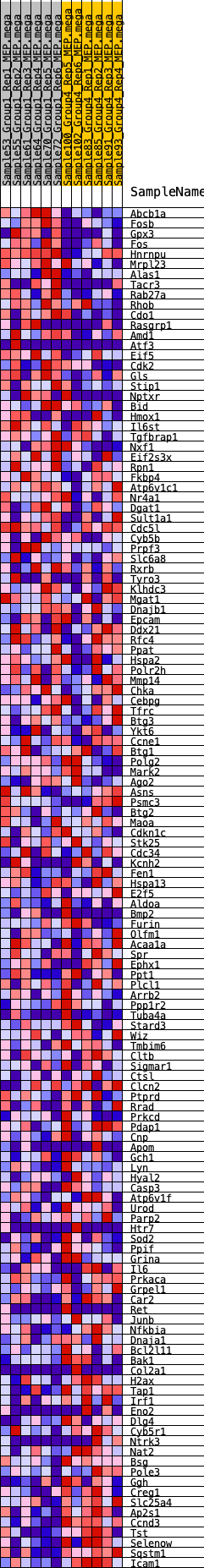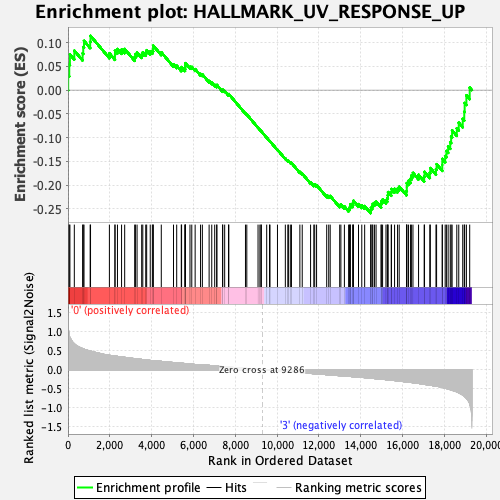
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group1_versus_Group4.MEP.mega_Pheno.cls #Group1_versus_Group4_repos |
| Phenotype | MEP.mega_Pheno.cls#Group1_versus_Group4_repos |
| Upregulated in class | 3 |
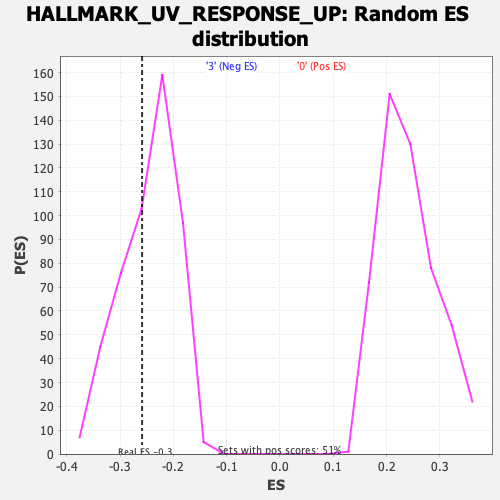
| GeneSet | HALLMARK_UV_RESPONSE_UP |
| Enrichment Score (ES) | -0.2587291 |
| Normalized Enrichment Score (NES) | -1.0525678 |
| Nominal p-value | 0.34756097 |
| FDR q-value | 0.56289333 |
| FWER p-Value | 0.97 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Abcb1a | 9 | 1.167 | 0.0313 | No |
| 2 | Fosb | 60 | 0.924 | 0.0538 | No |
| 3 | Gpx3 | 91 | 0.847 | 0.0753 | No |
| 4 | Fos | 302 | 0.683 | 0.0830 | No |
| 5 | Hnrnpu | 699 | 0.553 | 0.0774 | No |
| 6 | Mrpl23 | 729 | 0.548 | 0.0908 | No |
| 7 | Alas1 | 765 | 0.538 | 0.1036 | No |
| 8 | Tacr3 | 1060 | 0.490 | 0.1017 | No |
| 9 | Rab27a | 1077 | 0.486 | 0.1141 | No |
| 10 | Rhob | 1979 | 0.380 | 0.0774 | No |
| 11 | Cdo1 | 2245 | 0.357 | 0.0733 | No |
| 12 | Rasgrp1 | 2247 | 0.357 | 0.0830 | No |
| 13 | Amd1 | 2365 | 0.349 | 0.0864 | No |
| 14 | Atf3 | 2561 | 0.334 | 0.0853 | No |
| 15 | Eif5 | 2710 | 0.325 | 0.0865 | No |
| 16 | Cdk2 | 3189 | 0.290 | 0.0694 | No |
| 17 | Gls | 3223 | 0.288 | 0.0755 | No |
| 18 | Stip1 | 3302 | 0.281 | 0.0791 | No |
| 19 | Nptxr | 3523 | 0.270 | 0.0750 | No |
| 20 | Bid | 3576 | 0.266 | 0.0795 | No |
| 21 | Hmox1 | 3717 | 0.256 | 0.0792 | No |
| 22 | Il6st | 3761 | 0.254 | 0.0839 | No |
| 23 | Tgfbrap1 | 3932 | 0.243 | 0.0816 | No |
| 24 | Nxf1 | 4033 | 0.237 | 0.0828 | No |
| 25 | Eif2s3x | 4070 | 0.236 | 0.0874 | No |
| 26 | Rpn1 | 4073 | 0.235 | 0.0937 | No |
| 27 | Fkbp4 | 4460 | 0.218 | 0.0795 | No |
| 28 | Atp6v1c1 | 5044 | 0.188 | 0.0542 | No |
| 29 | Nr4a1 | 5197 | 0.182 | 0.0512 | No |
| 30 | Dgat1 | 5416 | 0.170 | 0.0445 | No |
| 31 | Sult1a1 | 5431 | 0.169 | 0.0484 | No |
| 32 | Cdc5l | 5592 | 0.161 | 0.0444 | No |
| 33 | Cyb5b | 5604 | 0.160 | 0.0482 | No |
| 34 | Prpf3 | 5606 | 0.160 | 0.0525 | No |
| 35 | Slc6a8 | 5612 | 0.160 | 0.0566 | No |
| 36 | Rxrb | 5832 | 0.148 | 0.0492 | No |
| 37 | Tyro3 | 5918 | 0.144 | 0.0487 | No |
| 38 | Klhdc3 | 6079 | 0.136 | 0.0441 | No |
| 39 | Mgat1 | 6337 | 0.125 | 0.0341 | No |
| 40 | Dnajb1 | 6424 | 0.122 | 0.0329 | No |
| 41 | Epcam | 6747 | 0.113 | 0.0192 | No |
| 42 | Ddx21 | 6873 | 0.106 | 0.0156 | No |
| 43 | Rfc4 | 7014 | 0.099 | 0.0110 | No |
| 44 | Ppat | 7111 | 0.095 | 0.0086 | No |
| 45 | Hspa2 | 7117 | 0.095 | 0.0109 | No |
| 46 | Polr2h | 7394 | 0.082 | -0.0013 | No |
| 47 | Mmp14 | 7395 | 0.082 | 0.0009 | No |
| 48 | Chka | 7491 | 0.078 | -0.0019 | No |
| 49 | Cebpg | 7674 | 0.069 | -0.0095 | No |
| 50 | Tfrc | 7690 | 0.069 | -0.0084 | No |
| 51 | Btg3 | 8485 | 0.034 | -0.0489 | No |
| 52 | Ykt6 | 8549 | 0.032 | -0.0513 | No |
| 53 | Ccne1 | 9086 | 0.009 | -0.0790 | No |
| 54 | Btg1 | 9176 | 0.005 | -0.0835 | No |
| 55 | Polg2 | 9226 | 0.003 | -0.0860 | No |
| 56 | Mark2 | 9245 | 0.002 | -0.0869 | No |
| 57 | Ago2 | 9500 | -0.004 | -0.1000 | No |
| 58 | Asns | 9634 | -0.009 | -0.1067 | No |
| 59 | Psmc3 | 9658 | -0.010 | -0.1077 | No |
| 60 | Btg2 | 10015 | -0.028 | -0.1255 | No |
| 61 | Maoa | 10391 | -0.044 | -0.1438 | No |
| 62 | Cdkn1c | 10513 | -0.049 | -0.1488 | No |
| 63 | Stk25 | 10536 | -0.050 | -0.1486 | No |
| 64 | Cdc34 | 10629 | -0.054 | -0.1519 | No |
| 65 | Kcnh2 | 10684 | -0.056 | -0.1532 | No |
| 66 | Fen1 | 11086 | -0.075 | -0.1721 | No |
| 67 | Hspa13 | 11199 | -0.080 | -0.1757 | No |
| 68 | E2f5 | 11598 | -0.098 | -0.1938 | No |
| 69 | Aldoa | 11753 | -0.105 | -0.1990 | No |
| 70 | Bmp2 | 11796 | -0.107 | -0.1983 | No |
| 71 | Furin | 11885 | -0.111 | -0.1998 | No |
| 72 | Olfm1 | 12369 | -0.131 | -0.2215 | No |
| 73 | Acaa1a | 12466 | -0.135 | -0.2228 | No |
| 74 | Spr | 12536 | -0.138 | -0.2226 | No |
| 75 | Ephx1 | 12990 | -0.157 | -0.2420 | No |
| 76 | Ppt1 | 13047 | -0.160 | -0.2405 | No |
| 77 | Plcl1 | 13215 | -0.168 | -0.2447 | No |
| 78 | Arrb2 | 13417 | -0.175 | -0.2504 | No |
| 79 | Ppp1r2 | 13461 | -0.177 | -0.2478 | No |
| 80 | Tuba4a | 13487 | -0.178 | -0.2443 | No |
| 81 | Stard3 | 13499 | -0.178 | -0.2400 | No |
| 82 | Wiz | 13613 | -0.183 | -0.2409 | No |
| 83 | Tmbim6 | 13632 | -0.184 | -0.2368 | No |
| 84 | Cltb | 13648 | -0.185 | -0.2326 | No |
| 85 | Sigmar1 | 13890 | -0.196 | -0.2398 | No |
| 86 | Ctsl | 14046 | -0.204 | -0.2423 | No |
| 87 | Clcn2 | 14184 | -0.209 | -0.2438 | No |
| 88 | Ptprd | 14471 | -0.224 | -0.2526 | Yes |
| 89 | Rrad | 14487 | -0.225 | -0.2473 | Yes |
| 90 | Prkcd | 14544 | -0.228 | -0.2440 | Yes |
| 91 | Pdap1 | 14574 | -0.229 | -0.2393 | Yes |
| 92 | Cnp | 14651 | -0.233 | -0.2369 | Yes |
| 93 | Apom | 14729 | -0.237 | -0.2345 | Yes |
| 94 | Gch1 | 14969 | -0.249 | -0.2401 | Yes |
| 95 | Lyn | 14980 | -0.250 | -0.2339 | Yes |
| 96 | Hyal2 | 15041 | -0.254 | -0.2301 | Yes |
| 97 | Casp3 | 15203 | -0.263 | -0.2313 | Yes |
| 98 | Atp6v1f | 15269 | -0.267 | -0.2274 | Yes |
| 99 | Urod | 15289 | -0.269 | -0.2211 | Yes |
| 100 | Parp2 | 15303 | -0.269 | -0.2145 | Yes |
| 101 | Htr7 | 15454 | -0.276 | -0.2148 | Yes |
| 102 | Sod2 | 15468 | -0.277 | -0.2079 | Yes |
| 103 | Ppif | 15614 | -0.286 | -0.2077 | Yes |
| 104 | Grina | 15757 | -0.293 | -0.2071 | Yes |
| 105 | Il6 | 15835 | -0.298 | -0.2030 | Yes |
| 106 | Prkaca | 16184 | -0.322 | -0.2124 | Yes |
| 107 | Grpel1 | 16198 | -0.323 | -0.2043 | Yes |
| 108 | Car2 | 16199 | -0.323 | -0.1955 | Yes |
| 109 | Ret | 16278 | -0.329 | -0.1906 | Yes |
| 110 | Junb | 16373 | -0.335 | -0.1864 | Yes |
| 111 | Nfkbia | 16418 | -0.337 | -0.1795 | Yes |
| 112 | Dnaja1 | 16493 | -0.343 | -0.1740 | Yes |
| 113 | Bcl2l11 | 16761 | -0.363 | -0.1781 | Yes |
| 114 | Bak1 | 17029 | -0.384 | -0.1816 | Yes |
| 115 | Col2a1 | 17046 | -0.386 | -0.1719 | Yes |
| 116 | H2ax | 17299 | -0.408 | -0.1739 | Yes |
| 117 | Tap1 | 17323 | -0.411 | -0.1639 | Yes |
| 118 | Irf1 | 17594 | -0.433 | -0.1662 | Yes |
| 119 | Eno2 | 17623 | -0.434 | -0.1559 | Yes |
| 120 | Dlg4 | 17894 | -0.466 | -0.1573 | Yes |
| 121 | Cyb5r1 | 17896 | -0.466 | -0.1446 | Yes |
| 122 | Ntrk3 | 18033 | -0.487 | -0.1385 | Yes |
| 123 | Nat2 | 18092 | -0.496 | -0.1280 | Yes |
| 124 | Bsg | 18175 | -0.508 | -0.1184 | Yes |
| 125 | Pole3 | 18275 | -0.525 | -0.1093 | Yes |
| 126 | Ggh | 18319 | -0.534 | -0.0970 | Yes |
| 127 | Creg1 | 18365 | -0.542 | -0.0846 | Yes |
| 128 | Slc25a4 | 18589 | -0.584 | -0.0803 | Yes |
| 129 | Ap2s1 | 18683 | -0.608 | -0.0686 | Yes |
| 130 | Ccnd3 | 18873 | -0.663 | -0.0604 | Yes |
| 131 | Tst | 18946 | -0.691 | -0.0453 | Yes |
| 132 | Selenow | 18965 | -0.706 | -0.0271 | Yes |
| 133 | Sqstm1 | 19048 | -0.750 | -0.0109 | Yes |
| 134 | Icam1 | 19206 | -0.896 | 0.0053 | Yes |