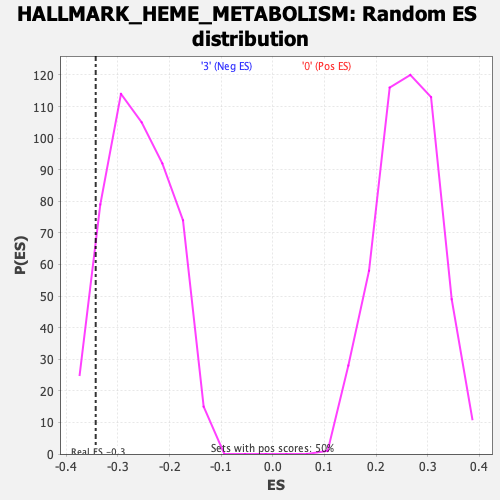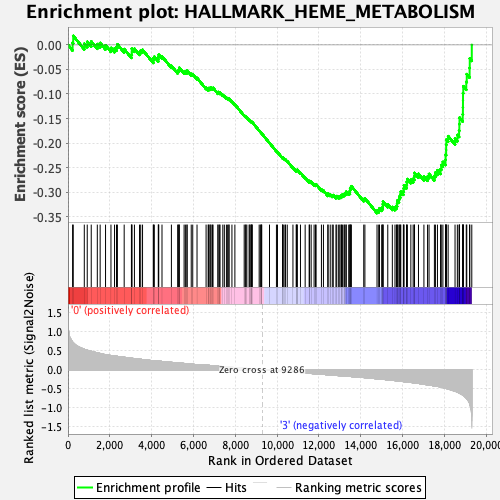
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group1_versus_Group4.MEP.mega_Pheno.cls #Group1_versus_Group4_repos |
| Phenotype | MEP.mega_Pheno.cls#Group1_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_HEME_METABOLISM |
| Enrichment Score (ES) | -0.34297374 |
| Normalized Enrichment Score (NES) | -1.3176128 |
| Nominal p-value | 0.0952381 |
| FDR q-value | 0.5449292 |
| FWER p-Value | 0.73 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Bmp2k | 220 | 0.726 | 0.0044 | No |
| 2 | P4ha2 | 250 | 0.707 | 0.0184 | No |
| 3 | Igsf3 | 779 | 0.536 | 0.0026 | No |
| 4 | Osbp2 | 920 | 0.511 | 0.0065 | No |
| 5 | Acsl6 | 1111 | 0.482 | 0.0071 | No |
| 6 | Gmps | 1401 | 0.441 | 0.0017 | No |
| 7 | Hebp1 | 1537 | 0.427 | 0.0040 | No |
| 8 | Ncoa4 | 1795 | 0.395 | -0.0007 | No |
| 9 | Tent5c | 2054 | 0.371 | -0.0061 | No |
| 10 | Nfe2l1 | 2225 | 0.359 | -0.0071 | No |
| 11 | Endod1 | 2320 | 0.352 | -0.0043 | No |
| 12 | Mboat2 | 2362 | 0.350 | 0.0012 | No |
| 13 | Abcg2 | 2686 | 0.327 | -0.0085 | No |
| 14 | Trak2 | 3032 | 0.301 | -0.0199 | No |
| 15 | Sidt2 | 3051 | 0.299 | -0.0143 | No |
| 16 | Dcaf11 | 3052 | 0.299 | -0.0077 | No |
| 17 | Fbxo34 | 3176 | 0.291 | -0.0077 | No |
| 18 | C3 | 3420 | 0.275 | -0.0144 | No |
| 19 | Add2 | 3477 | 0.271 | -0.0114 | No |
| 20 | Rnf19a | 3564 | 0.267 | -0.0100 | No |
| 21 | Mpp1 | 4078 | 0.235 | -0.0317 | No |
| 22 | Fbxo7 | 4086 | 0.234 | -0.0269 | No |
| 23 | Ell2 | 4127 | 0.232 | -0.0239 | No |
| 24 | Spta1 | 4319 | 0.226 | -0.0289 | No |
| 25 | Mgst3 | 4324 | 0.226 | -0.0242 | No |
| 26 | Khnyn | 4339 | 0.225 | -0.0200 | No |
| 27 | Bach1 | 4494 | 0.216 | -0.0233 | No |
| 28 | Marchf8 | 4944 | 0.192 | -0.0426 | No |
| 29 | Atp6v0a1 | 5245 | 0.180 | -0.0543 | No |
| 30 | Ermap | 5275 | 0.178 | -0.0519 | No |
| 31 | Rnf123 | 5312 | 0.176 | -0.0499 | No |
| 32 | Gclc | 5320 | 0.175 | -0.0465 | No |
| 33 | Sec14l1 | 5551 | 0.163 | -0.0549 | No |
| 34 | Slc6a8 | 5612 | 0.160 | -0.0545 | No |
| 35 | Gde1 | 5686 | 0.156 | -0.0549 | No |
| 36 | Picalm | 5695 | 0.156 | -0.0519 | No |
| 37 | Smox | 5892 | 0.145 | -0.0590 | No |
| 38 | Map2k3 | 5964 | 0.142 | -0.0596 | No |
| 39 | Slc25a38 | 6169 | 0.131 | -0.0674 | No |
| 40 | Bnip3l | 6602 | 0.119 | -0.0873 | No |
| 41 | Trim58 | 6704 | 0.115 | -0.0901 | No |
| 42 | Abcb6 | 6713 | 0.114 | -0.0880 | No |
| 43 | Synj1 | 6789 | 0.110 | -0.0895 | No |
| 44 | Ctsb | 6794 | 0.110 | -0.0873 | No |
| 45 | Mospd1 | 6842 | 0.108 | -0.0874 | No |
| 46 | Fbxo9 | 6907 | 0.105 | -0.0884 | No |
| 47 | Aldh1l1 | 6934 | 0.103 | -0.0875 | No |
| 48 | Slc6a9 | 7170 | 0.092 | -0.0978 | No |
| 49 | Foxo3 | 7174 | 0.092 | -0.0959 | No |
| 50 | Mocos | 7252 | 0.088 | -0.0980 | No |
| 51 | Eif2ak1 | 7264 | 0.088 | -0.0967 | No |
| 52 | E2f2 | 7402 | 0.081 | -0.1021 | No |
| 53 | Usp15 | 7475 | 0.079 | -0.1041 | No |
| 54 | Tns1 | 7574 | 0.074 | -0.1076 | No |
| 55 | Clcn3 | 7633 | 0.071 | -0.1091 | No |
| 56 | Tfrc | 7690 | 0.069 | -0.1105 | No |
| 57 | Nr3c1 | 7700 | 0.068 | -0.1095 | No |
| 58 | Ubac1 | 7838 | 0.061 | -0.1153 | No |
| 59 | Ranbp10 | 7979 | 0.054 | -0.1214 | No |
| 60 | Gapvd1 | 8424 | 0.037 | -0.1438 | No |
| 61 | Aldh6a1 | 8479 | 0.034 | -0.1459 | No |
| 62 | Agpat4 | 8508 | 0.033 | -0.1466 | No |
| 63 | Rcl1 | 8551 | 0.032 | -0.1481 | No |
| 64 | Nek7 | 8659 | 0.027 | -0.1531 | No |
| 65 | Hmbs | 8702 | 0.026 | -0.1547 | No |
| 66 | Rbm5 | 8705 | 0.026 | -0.1543 | No |
| 67 | Sptb | 8777 | 0.022 | -0.1575 | No |
| 68 | Nudt4 | 8787 | 0.022 | -0.1575 | No |
| 69 | Optn | 8789 | 0.022 | -0.1571 | No |
| 70 | Kdm7a | 8805 | 0.021 | -0.1574 | No |
| 71 | Mark3 | 9140 | 0.006 | -0.1747 | No |
| 72 | Ppox | 9195 | 0.004 | -0.1774 | No |
| 73 | Cdc27 | 9234 | 0.002 | -0.1794 | No |
| 74 | Xpo7 | 9244 | 0.002 | -0.1798 | No |
| 75 | Ezh1 | 9263 | 0.001 | -0.1807 | No |
| 76 | Asns | 9634 | -0.009 | -0.1999 | No |
| 77 | Minpp1 | 9966 | -0.024 | -0.2166 | No |
| 78 | Btg2 | 10015 | -0.028 | -0.2185 | No |
| 79 | Epb41 | 10256 | -0.039 | -0.2302 | No |
| 80 | Snca | 10258 | -0.039 | -0.2294 | No |
| 81 | Tmem9b | 10293 | -0.040 | -0.2303 | No |
| 82 | Riok3 | 10371 | -0.043 | -0.2334 | No |
| 83 | Slc11a2 | 10402 | -0.044 | -0.2340 | No |
| 84 | Vezf1 | 10483 | -0.048 | -0.2371 | No |
| 85 | Tspan5 | 10755 | -0.060 | -0.2500 | No |
| 86 | Ypel5 | 10910 | -0.067 | -0.2565 | No |
| 87 | Narf | 10925 | -0.068 | -0.2558 | No |
| 88 | Dcun1d1 | 10952 | -0.069 | -0.2556 | No |
| 89 | Daam1 | 10960 | -0.070 | -0.2545 | No |
| 90 | Nnt | 11115 | -0.076 | -0.2609 | No |
| 91 | Arhgef12 | 11339 | -0.086 | -0.2706 | No |
| 92 | Dcaf10 | 11530 | -0.095 | -0.2785 | No |
| 93 | Add1 | 11549 | -0.096 | -0.2773 | No |
| 94 | Lpin2 | 11626 | -0.099 | -0.2791 | No |
| 95 | Rhag | 11766 | -0.106 | -0.2841 | No |
| 96 | Slc30a1 | 11832 | -0.109 | -0.2851 | No |
| 97 | Cast | 11862 | -0.110 | -0.2842 | No |
| 98 | Ucp2 | 12117 | -0.121 | -0.2948 | No |
| 99 | Pcx | 12213 | -0.125 | -0.2970 | No |
| 100 | Klf3 | 12423 | -0.133 | -0.3050 | No |
| 101 | Mfhas1 | 12435 | -0.133 | -0.3027 | No |
| 102 | Fech | 12538 | -0.138 | -0.3050 | No |
| 103 | Foxj2 | 12635 | -0.142 | -0.3069 | No |
| 104 | Slc7a11 | 12682 | -0.144 | -0.3061 | No |
| 105 | Lamp2 | 12812 | -0.150 | -0.3096 | No |
| 106 | Btrc | 12850 | -0.152 | -0.3082 | No |
| 107 | Blvra | 12941 | -0.156 | -0.3095 | No |
| 108 | Tcea1 | 12995 | -0.158 | -0.3088 | No |
| 109 | Lmo2 | 13057 | -0.160 | -0.3085 | No |
| 110 | Mxi1 | 13078 | -0.161 | -0.3060 | No |
| 111 | Tnrc6b | 13129 | -0.163 | -0.3050 | No |
| 112 | Cir1 | 13198 | -0.167 | -0.3049 | No |
| 113 | Epb42 | 13229 | -0.168 | -0.3028 | No |
| 114 | Rbm38 | 13290 | -0.171 | -0.3022 | No |
| 115 | Cpox | 13302 | -0.171 | -0.2990 | No |
| 116 | Kat2b | 13422 | -0.175 | -0.3014 | No |
| 117 | Gclm | 13478 | -0.177 | -0.3004 | No |
| 118 | Pigq | 13482 | -0.178 | -0.2966 | No |
| 119 | Blvrb | 13491 | -0.178 | -0.2931 | No |
| 120 | Sdcbp | 13519 | -0.180 | -0.2906 | No |
| 121 | Psmd9 | 13548 | -0.181 | -0.2881 | No |
| 122 | Atg4a | 14140 | -0.207 | -0.3145 | No |
| 123 | Top1 | 14195 | -0.209 | -0.3127 | No |
| 124 | Ctns | 14775 | -0.240 | -0.3377 | Yes |
| 125 | Kel | 14856 | -0.245 | -0.3365 | Yes |
| 126 | Ackr1 | 14893 | -0.247 | -0.3330 | Yes |
| 127 | Slc4a1 | 14998 | -0.252 | -0.3329 | Yes |
| 128 | Pdzk1ip1 | 15049 | -0.254 | -0.3299 | Yes |
| 129 | Tspo2 | 15054 | -0.255 | -0.3245 | Yes |
| 130 | Mkrn1 | 15063 | -0.255 | -0.3193 | Yes |
| 131 | Urod | 15289 | -0.269 | -0.3252 | Yes |
| 132 | Myl4 | 15507 | -0.279 | -0.3304 | Yes |
| 133 | Ppp2r5b | 15634 | -0.287 | -0.3307 | Yes |
| 134 | Ank1 | 15704 | -0.290 | -0.3280 | Yes |
| 135 | Dmtn | 15737 | -0.292 | -0.3232 | Yes |
| 136 | Slc22a4 | 15745 | -0.293 | -0.3172 | Yes |
| 137 | Gypc | 15824 | -0.297 | -0.3147 | Yes |
| 138 | Arl2bp | 15841 | -0.299 | -0.3090 | Yes |
| 139 | Hagh | 15891 | -0.302 | -0.3049 | Yes |
| 140 | Tmcc2 | 15911 | -0.304 | -0.2992 | Yes |
| 141 | Tal1 | 16034 | -0.313 | -0.2988 | Yes |
| 142 | Htatip2 | 16048 | -0.314 | -0.2926 | Yes |
| 143 | Tfdp2 | 16065 | -0.315 | -0.2865 | Yes |
| 144 | Pgls | 16185 | -0.322 | -0.2856 | Yes |
| 145 | Car2 | 16199 | -0.323 | -0.2792 | Yes |
| 146 | Acp5 | 16226 | -0.325 | -0.2735 | Yes |
| 147 | Glrx5 | 16392 | -0.336 | -0.2747 | Yes |
| 148 | Xk | 16502 | -0.344 | -0.2729 | Yes |
| 149 | Cdr2 | 16556 | -0.347 | -0.2680 | Yes |
| 150 | Alad | 16568 | -0.348 | -0.2610 | Yes |
| 151 | Rap1gap | 16757 | -0.362 | -0.2628 | Yes |
| 152 | Fn3k | 17017 | -0.383 | -0.2680 | Yes |
| 153 | Nfe2 | 17191 | -0.398 | -0.2683 | Yes |
| 154 | Icam4 | 17262 | -0.405 | -0.2630 | Yes |
| 155 | Klf1 | 17518 | -0.427 | -0.2670 | Yes |
| 156 | Marchf2 | 17567 | -0.431 | -0.2601 | Yes |
| 157 | Adipor1 | 17664 | -0.439 | -0.2555 | Yes |
| 158 | Lrp10 | 17812 | -0.456 | -0.2531 | Yes |
| 159 | Bpgm | 17846 | -0.460 | -0.2448 | Yes |
| 160 | Hdgf | 17924 | -0.471 | -0.2384 | Yes |
| 161 | Slc10a3 | 18049 | -0.490 | -0.2342 | Yes |
| 162 | Epor | 18054 | -0.490 | -0.2236 | Yes |
| 163 | Gata1 | 18079 | -0.493 | -0.2141 | Yes |
| 164 | Slc66a2 | 18081 | -0.494 | -0.2033 | Yes |
| 165 | H1f0 | 18090 | -0.495 | -0.1928 | Yes |
| 166 | Bsg | 18175 | -0.508 | -0.1861 | Yes |
| 167 | Selenbp1 | 18505 | -0.568 | -0.1908 | Yes |
| 168 | Rad23a | 18616 | -0.592 | -0.1836 | Yes |
| 169 | Car1 | 18699 | -0.614 | -0.1744 | Yes |
| 170 | Tyr | 18712 | -0.616 | -0.1615 | Yes |
| 171 | Htra2 | 18719 | -0.618 | -0.1483 | Yes |
| 172 | Ccnd3 | 18873 | -0.663 | -0.1417 | Yes |
| 173 | Trim10 | 18882 | -0.668 | -0.1275 | Yes |
| 174 | Rhd | 18885 | -0.668 | -0.1129 | Yes |
| 175 | Ccdc28a | 18887 | -0.668 | -0.0983 | Yes |
| 176 | Slc2a1 | 18900 | -0.673 | -0.0842 | Yes |
| 177 | Cat | 19042 | -0.748 | -0.0751 | Yes |
| 178 | Alas2 | 19066 | -0.765 | -0.0596 | Yes |
| 179 | Uros | 19202 | -0.891 | -0.0471 | Yes |
| 180 | Ctse | 19209 | -0.897 | -0.0277 | Yes |
| 181 | Prdx2 | 19306 | -1.493 | 0.0001 | Yes |