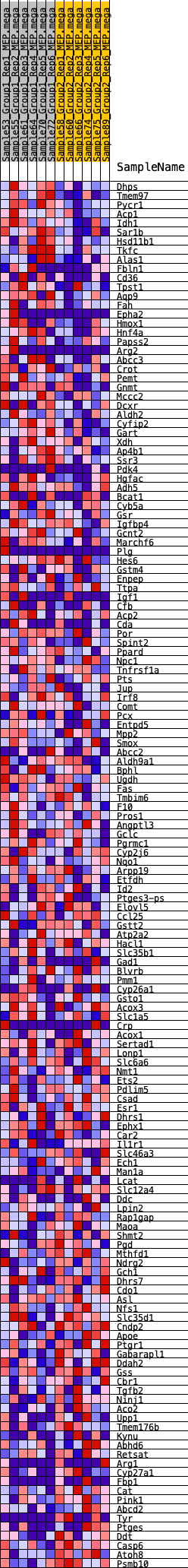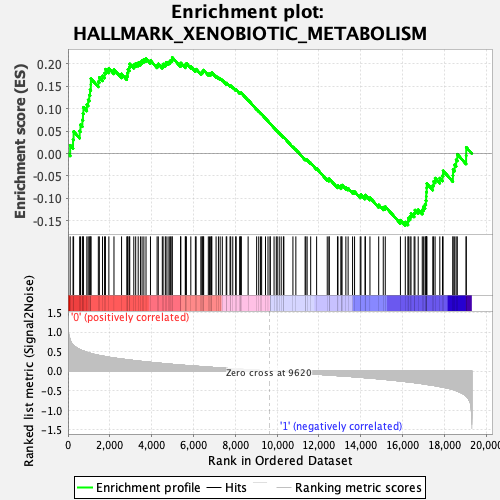
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group1_versus_Group2.MEP.mega_Pheno.cls #Group1_versus_Group2_repos |
| Phenotype | MEP.mega_Pheno.cls#Group1_versus_Group2_repos |
| Upregulated in class | 0 |
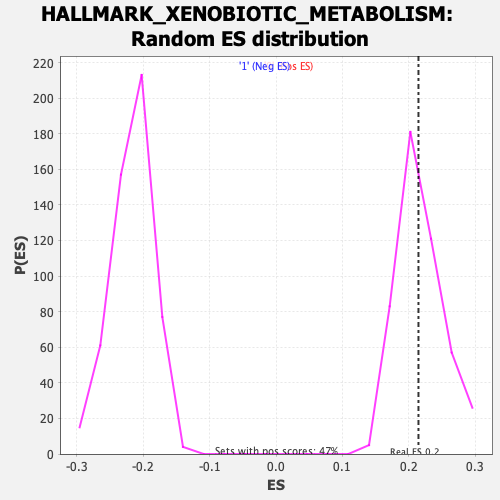
| GeneSet | HALLMARK_XENOBIOTIC_METABOLISM |
| Enrichment Score (ES) | 0.2142462 |
| Normalized Enrichment Score (NES) | 0.9892704 |
| Nominal p-value | 0.45665962 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.985 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Dhps | 104 | 0.802 | 0.0184 | Yes |
| 2 | Tmem97 | 241 | 0.666 | 0.0311 | Yes |
| 3 | Pycr1 | 268 | 0.653 | 0.0491 | Yes |
| 4 | Acp1 | 558 | 0.552 | 0.0504 | Yes |
| 5 | Idh1 | 597 | 0.538 | 0.0644 | Yes |
| 6 | Sar1b | 687 | 0.521 | 0.0753 | Yes |
| 7 | Hsd11b1 | 716 | 0.515 | 0.0891 | Yes |
| 8 | Tkfc | 737 | 0.510 | 0.1032 | Yes |
| 9 | Alas1 | 901 | 0.479 | 0.1090 | Yes |
| 10 | Fbln1 | 963 | 0.469 | 0.1197 | Yes |
| 11 | Cd36 | 1017 | 0.460 | 0.1306 | Yes |
| 12 | Tpst1 | 1054 | 0.454 | 0.1422 | Yes |
| 13 | Aqp9 | 1092 | 0.450 | 0.1537 | Yes |
| 14 | Fah | 1095 | 0.450 | 0.1669 | Yes |
| 15 | Epha2 | 1450 | 0.402 | 0.1604 | Yes |
| 16 | Hmox1 | 1498 | 0.398 | 0.1698 | Yes |
| 17 | Hnf4a | 1651 | 0.382 | 0.1732 | Yes |
| 18 | Papss2 | 1749 | 0.374 | 0.1792 | Yes |
| 19 | Arg2 | 1790 | 0.369 | 0.1881 | Yes |
| 20 | Abcc3 | 1954 | 0.354 | 0.1901 | Yes |
| 21 | Crot | 2198 | 0.334 | 0.1873 | Yes |
| 22 | Pemt | 2561 | 0.307 | 0.1775 | Yes |
| 23 | Gnmt | 2812 | 0.291 | 0.1731 | Yes |
| 24 | Mccc2 | 2851 | 0.288 | 0.1797 | Yes |
| 25 | Dcxr | 2865 | 0.286 | 0.1875 | Yes |
| 26 | Aldh2 | 2933 | 0.283 | 0.1924 | Yes |
| 27 | Cyfip2 | 2952 | 0.281 | 0.1998 | Yes |
| 28 | Gart | 3146 | 0.270 | 0.1978 | Yes |
| 29 | Xdh | 3236 | 0.263 | 0.2010 | Yes |
| 30 | Ap4b1 | 3356 | 0.255 | 0.2023 | Yes |
| 31 | Ssr3 | 3464 | 0.249 | 0.2042 | Yes |
| 32 | Pdk4 | 3541 | 0.245 | 0.2075 | Yes |
| 33 | Hgfac | 3628 | 0.239 | 0.2101 | Yes |
| 34 | Adh5 | 3730 | 0.233 | 0.2118 | Yes |
| 35 | Bcat1 | 3943 | 0.223 | 0.2073 | Yes |
| 36 | Cyb5a | 4260 | 0.205 | 0.1969 | Yes |
| 37 | Gsr | 4318 | 0.203 | 0.2000 | Yes |
| 38 | Igfbp4 | 4509 | 0.194 | 0.1958 | Yes |
| 39 | Gcnt2 | 4547 | 0.192 | 0.1996 | Yes |
| 40 | Marchf6 | 4652 | 0.187 | 0.1997 | Yes |
| 41 | Plg | 4690 | 0.185 | 0.2033 | Yes |
| 42 | Hes6 | 4784 | 0.181 | 0.2038 | Yes |
| 43 | Gstm4 | 4864 | 0.177 | 0.2049 | Yes |
| 44 | Enpep | 4907 | 0.175 | 0.2079 | Yes |
| 45 | Ttpa | 4972 | 0.172 | 0.2097 | Yes |
| 46 | Igf1 | 4983 | 0.171 | 0.2142 | Yes |
| 47 | Cfb | 5381 | 0.153 | 0.1981 | No |
| 48 | Acp2 | 5394 | 0.152 | 0.2019 | No |
| 49 | Cda | 5607 | 0.144 | 0.1951 | No |
| 50 | Por | 5619 | 0.143 | 0.1988 | No |
| 51 | Spint2 | 5661 | 0.142 | 0.2009 | No |
| 52 | Ppard | 5872 | 0.132 | 0.1939 | No |
| 53 | Npc1 | 6090 | 0.124 | 0.1862 | No |
| 54 | Tnfrsf1a | 6125 | 0.122 | 0.1881 | No |
| 55 | Pts | 6352 | 0.114 | 0.1797 | No |
| 56 | Jup | 6402 | 0.112 | 0.1805 | No |
| 57 | Irf8 | 6422 | 0.112 | 0.1828 | No |
| 58 | Comt | 6464 | 0.110 | 0.1839 | No |
| 59 | Pcx | 6486 | 0.110 | 0.1861 | No |
| 60 | Entpd5 | 6709 | 0.100 | 0.1775 | No |
| 61 | Mpp2 | 6763 | 0.098 | 0.1776 | No |
| 62 | Smox | 6820 | 0.096 | 0.1776 | No |
| 63 | Abcc2 | 6866 | 0.095 | 0.1780 | No |
| 64 | Aldh9a1 | 6874 | 0.095 | 0.1805 | No |
| 65 | Bphl | 7080 | 0.087 | 0.1724 | No |
| 66 | Ugdh | 7193 | 0.083 | 0.1690 | No |
| 67 | Fas | 7275 | 0.079 | 0.1671 | No |
| 68 | Tmbim6 | 7378 | 0.075 | 0.1640 | No |
| 69 | F10 | 7564 | 0.070 | 0.1564 | No |
| 70 | Pros1 | 7590 | 0.069 | 0.1572 | No |
| 71 | Angptl3 | 7748 | 0.063 | 0.1508 | No |
| 72 | Gclc | 7780 | 0.062 | 0.1511 | No |
| 73 | Pgrmc1 | 7878 | 0.058 | 0.1477 | No |
| 74 | Cyp2j6 | 8015 | 0.053 | 0.1422 | No |
| 75 | Nqo1 | 8040 | 0.052 | 0.1425 | No |
| 76 | Arpp19 | 8207 | 0.047 | 0.1352 | No |
| 77 | Etfdh | 8240 | 0.046 | 0.1349 | No |
| 78 | Id2 | 8264 | 0.045 | 0.1351 | No |
| 79 | Ptges3-ps | 8285 | 0.044 | 0.1353 | No |
| 80 | Elovl5 | 8612 | 0.034 | 0.1193 | No |
| 81 | Ccl25 | 9019 | 0.020 | 0.0987 | No |
| 82 | Gstt2 | 9119 | 0.017 | 0.0941 | No |
| 83 | Atp2a2 | 9201 | 0.014 | 0.0902 | No |
| 84 | Hacl1 | 9258 | 0.012 | 0.0877 | No |
| 85 | Slc35b1 | 9443 | 0.006 | 0.0783 | No |
| 86 | Gad1 | 9562 | 0.002 | 0.0722 | No |
| 87 | Blvrb | 9664 | -0.000 | 0.0669 | No |
| 88 | Pmm1 | 9667 | -0.000 | 0.0668 | No |
| 89 | Cyp26a1 | 9852 | -0.006 | 0.0574 | No |
| 90 | Gsto1 | 9958 | -0.011 | 0.0522 | No |
| 91 | Acox3 | 10003 | -0.012 | 0.0503 | No |
| 92 | Slc1a5 | 10099 | -0.015 | 0.0457 | No |
| 93 | Crp | 10194 | -0.019 | 0.0414 | No |
| 94 | Acox1 | 10299 | -0.022 | 0.0366 | No |
| 95 | Sertad1 | 10321 | -0.023 | 0.0362 | No |
| 96 | Lonp1 | 10754 | -0.037 | 0.0147 | No |
| 97 | Slc6a6 | 10894 | -0.041 | 0.0087 | No |
| 98 | Nmt1 | 11337 | -0.057 | -0.0127 | No |
| 99 | Ets2 | 11370 | -0.058 | -0.0126 | No |
| 100 | Pdlim5 | 11433 | -0.061 | -0.0141 | No |
| 101 | Csad | 11603 | -0.067 | -0.0209 | No |
| 102 | Esr1 | 11887 | -0.077 | -0.0334 | No |
| 103 | Dhrs1 | 12395 | -0.095 | -0.0570 | No |
| 104 | Ephx1 | 12482 | -0.099 | -0.0586 | No |
| 105 | Car2 | 12485 | -0.099 | -0.0557 | No |
| 106 | Il1r1 | 12884 | -0.114 | -0.0731 | No |
| 107 | Slc46a3 | 12908 | -0.115 | -0.0709 | No |
| 108 | Ech1 | 13056 | -0.120 | -0.0750 | No |
| 109 | Man1a | 13057 | -0.120 | -0.0715 | No |
| 110 | Lcat | 13103 | -0.122 | -0.0702 | No |
| 111 | Slc12a4 | 13285 | -0.129 | -0.0758 | No |
| 112 | Ddc | 13393 | -0.133 | -0.0774 | No |
| 113 | Lpin2 | 13606 | -0.142 | -0.0843 | No |
| 114 | Rap1gap | 13701 | -0.145 | -0.0849 | No |
| 115 | Maoa | 13972 | -0.158 | -0.0943 | No |
| 116 | Shmt2 | 14006 | -0.160 | -0.0913 | No |
| 117 | Pgd | 14202 | -0.167 | -0.0965 | No |
| 118 | Mthfd1 | 14220 | -0.168 | -0.0924 | No |
| 119 | Ndrg2 | 14437 | -0.178 | -0.0984 | No |
| 120 | Gch1 | 14855 | -0.196 | -0.1143 | No |
| 121 | Dhrs7 | 15073 | -0.208 | -0.1194 | No |
| 122 | Cdo1 | 15169 | -0.213 | -0.1181 | No |
| 123 | Asl | 15894 | -0.251 | -0.1484 | No |
| 124 | Nfs1 | 16127 | -0.266 | -0.1526 | No |
| 125 | Slc35d1 | 16256 | -0.274 | -0.1512 | No |
| 126 | Cndp2 | 16269 | -0.274 | -0.1436 | No |
| 127 | Apoe | 16354 | -0.280 | -0.1397 | No |
| 128 | Ptgr1 | 16400 | -0.283 | -0.1336 | No |
| 129 | Gabarapl1 | 16549 | -0.293 | -0.1327 | No |
| 130 | Ddah2 | 16588 | -0.296 | -0.1258 | No |
| 131 | Gss | 16744 | -0.306 | -0.1248 | No |
| 132 | Cbr1 | 16938 | -0.321 | -0.1254 | No |
| 133 | Tgfb2 | 16990 | -0.325 | -0.1184 | No |
| 134 | Ninj1 | 17071 | -0.332 | -0.1127 | No |
| 135 | Aco2 | 17110 | -0.335 | -0.1047 | No |
| 136 | Upp1 | 17127 | -0.337 | -0.0956 | No |
| 137 | Tmem176b | 17131 | -0.337 | -0.0857 | No |
| 138 | Kynu | 17150 | -0.338 | -0.0766 | No |
| 139 | Abhd6 | 17155 | -0.339 | -0.0667 | No |
| 140 | Retsat | 17442 | -0.363 | -0.0709 | No |
| 141 | Arg1 | 17480 | -0.366 | -0.0620 | No |
| 142 | Cyp27a1 | 17552 | -0.371 | -0.0546 | No |
| 143 | Fbp1 | 17777 | -0.395 | -0.0546 | No |
| 144 | Cat | 17906 | -0.407 | -0.0492 | No |
| 145 | Pink1 | 17933 | -0.410 | -0.0383 | No |
| 146 | Abcd2 | 18402 | -0.468 | -0.0489 | No |
| 147 | Tyr | 18411 | -0.469 | -0.0353 | No |
| 148 | Ptges | 18488 | -0.484 | -0.0249 | No |
| 149 | Ddt | 18557 | -0.497 | -0.0137 | No |
| 150 | Casp6 | 18613 | -0.509 | -0.0015 | No |
| 151 | Atoh8 | 19034 | -0.631 | -0.0046 | No |
| 152 | Psmb10 | 19040 | -0.634 | 0.0139 | No |