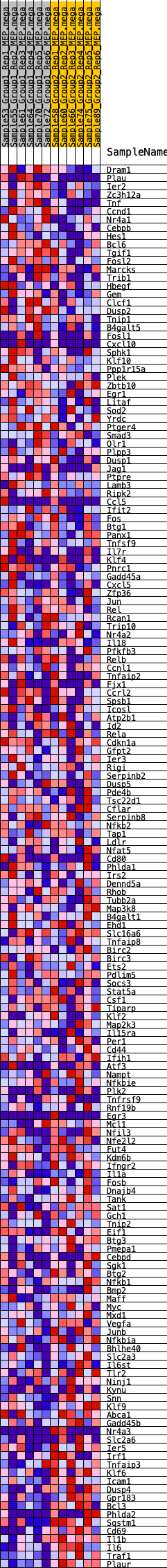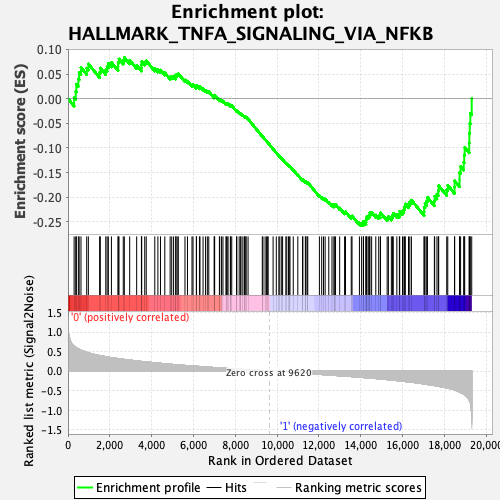
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group1_versus_Group2.MEP.mega_Pheno.cls #Group1_versus_Group2_repos |
| Phenotype | MEP.mega_Pheno.cls#Group1_versus_Group2_repos |
| Upregulated in class | 1 |
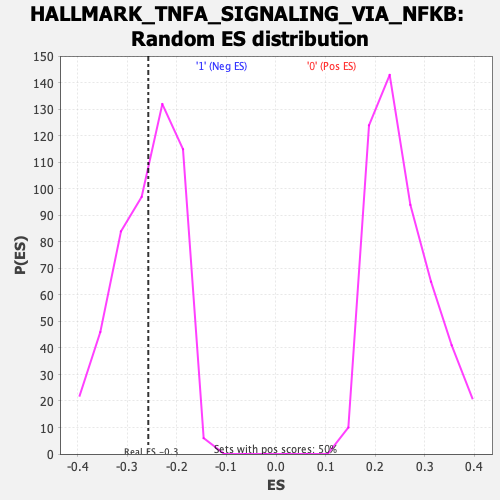
| GeneSet | HALLMARK_TNFA_SIGNALING_VIA_NFKB |
| Enrichment Score (ES) | -0.2578975 |
| Normalized Enrichment Score (NES) | -0.9885482 |
| Nominal p-value | 0.4621514 |
| FDR q-value | 0.96273905 |
| FWER p-Value | 0.985 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Dram1 | 293 | 0.643 | 0.0022 | No |
| 2 | Plau | 370 | 0.611 | 0.0148 | No |
| 3 | Ier2 | 401 | 0.596 | 0.0295 | No |
| 4 | Zc3h12a | 495 | 0.567 | 0.0400 | No |
| 5 | Tnf | 533 | 0.557 | 0.0532 | No |
| 6 | Ccnd1 | 620 | 0.533 | 0.0632 | No |
| 7 | Nr4a1 | 894 | 0.480 | 0.0620 | No |
| 8 | Cebpb | 973 | 0.467 | 0.0707 | No |
| 9 | Hes1 | 1512 | 0.396 | 0.0533 | No |
| 10 | Bcl6 | 1544 | 0.392 | 0.0624 | No |
| 11 | Tgif1 | 1806 | 0.368 | 0.0588 | No |
| 12 | Fosl2 | 1871 | 0.362 | 0.0652 | No |
| 13 | Marcks | 1929 | 0.356 | 0.0720 | No |
| 14 | Trib1 | 2076 | 0.342 | 0.0736 | No |
| 15 | Hbegf | 2396 | 0.318 | 0.0656 | No |
| 16 | Gem | 2398 | 0.317 | 0.0742 | No |
| 17 | Clcf1 | 2438 | 0.314 | 0.0807 | No |
| 18 | Dusp2 | 2642 | 0.302 | 0.0783 | No |
| 19 | Tnip1 | 2693 | 0.298 | 0.0838 | No |
| 20 | B4galt5 | 2949 | 0.281 | 0.0781 | No |
| 21 | Fosl1 | 3283 | 0.260 | 0.0678 | No |
| 22 | Cxcl10 | 3511 | 0.246 | 0.0627 | No |
| 23 | Sphk1 | 3514 | 0.246 | 0.0693 | No |
| 24 | Klf10 | 3524 | 0.246 | 0.0755 | No |
| 25 | Ppp1r15a | 3669 | 0.237 | 0.0744 | No |
| 26 | Plek | 3740 | 0.233 | 0.0771 | No |
| 27 | Zbtb10 | 4149 | 0.211 | 0.0615 | No |
| 28 | Egr1 | 4298 | 0.204 | 0.0593 | No |
| 29 | Litaf | 4423 | 0.198 | 0.0582 | No |
| 30 | Sod2 | 4628 | 0.188 | 0.0527 | No |
| 31 | Yrdc | 4883 | 0.176 | 0.0442 | No |
| 32 | Ptger4 | 4959 | 0.173 | 0.0450 | No |
| 33 | Smad3 | 5048 | 0.168 | 0.0450 | No |
| 34 | Olr1 | 5153 | 0.163 | 0.0440 | No |
| 35 | Plpp3 | 5156 | 0.163 | 0.0483 | No |
| 36 | Dusp1 | 5219 | 0.160 | 0.0494 | No |
| 37 | Jag1 | 5274 | 0.158 | 0.0509 | No |
| 38 | Ptpre | 5593 | 0.144 | 0.0382 | No |
| 39 | Lamb3 | 5712 | 0.139 | 0.0358 | No |
| 40 | Ripk2 | 5928 | 0.130 | 0.0281 | No |
| 41 | Ccl5 | 5978 | 0.128 | 0.0290 | No |
| 42 | Ifit2 | 6136 | 0.122 | 0.0241 | No |
| 43 | Fos | 6145 | 0.122 | 0.0270 | No |
| 44 | Btg1 | 6285 | 0.117 | 0.0229 | No |
| 45 | Panx1 | 6310 | 0.117 | 0.0249 | No |
| 46 | Tnfsf9 | 6457 | 0.110 | 0.0202 | No |
| 47 | Il7r | 6577 | 0.105 | 0.0169 | No |
| 48 | Klf4 | 6662 | 0.102 | 0.0153 | No |
| 49 | Pnrc1 | 6731 | 0.100 | 0.0144 | No |
| 50 | Gadd45a | 6987 | 0.090 | 0.0036 | No |
| 51 | Cxcl5 | 7004 | 0.090 | 0.0052 | No |
| 52 | Zfp36 | 7012 | 0.089 | 0.0072 | No |
| 53 | Jun | 7239 | 0.081 | -0.0024 | No |
| 54 | Rel | 7254 | 0.080 | -0.0009 | No |
| 55 | Rcan1 | 7342 | 0.077 | -0.0034 | No |
| 56 | Trip10 | 7409 | 0.074 | -0.0048 | No |
| 57 | Nr4a2 | 7556 | 0.070 | -0.0105 | No |
| 58 | Il18 | 7569 | 0.069 | -0.0093 | No |
| 59 | Pfkfb3 | 7608 | 0.068 | -0.0094 | No |
| 60 | Relb | 7652 | 0.067 | -0.0099 | No |
| 61 | Ccnl1 | 7762 | 0.063 | -0.0138 | No |
| 62 | Tnfaip2 | 7775 | 0.062 | -0.0128 | No |
| 63 | Fjx1 | 7829 | 0.060 | -0.0139 | No |
| 64 | Ccrl2 | 8060 | 0.052 | -0.0245 | No |
| 65 | Spsb1 | 8071 | 0.051 | -0.0237 | No |
| 66 | Icosl | 8185 | 0.048 | -0.0283 | No |
| 67 | Atp2b1 | 8260 | 0.045 | -0.0309 | No |
| 68 | Id2 | 8264 | 0.045 | -0.0299 | No |
| 69 | Rela | 8360 | 0.042 | -0.0337 | No |
| 70 | Cdkn1a | 8434 | 0.040 | -0.0364 | No |
| 71 | Gfpt2 | 8458 | 0.039 | -0.0365 | No |
| 72 | Ier3 | 8467 | 0.039 | -0.0359 | No |
| 73 | Rigi | 8520 | 0.037 | -0.0376 | No |
| 74 | Serpinb2 | 8609 | 0.035 | -0.0412 | No |
| 75 | Dusp5 | 9289 | 0.011 | -0.0764 | No |
| 76 | Pde4b | 9321 | 0.010 | -0.0778 | No |
| 77 | Tsc22d1 | 9423 | 0.007 | -0.0829 | No |
| 78 | Cflar | 9494 | 0.005 | -0.0864 | No |
| 79 | Serpinb8 | 9531 | 0.003 | -0.0882 | No |
| 80 | Nfkb2 | 9571 | 0.002 | -0.0902 | No |
| 81 | Tap1 | 9806 | -0.005 | -0.1023 | No |
| 82 | Ldlr | 9959 | -0.011 | -0.1099 | No |
| 83 | Nfat5 | 10081 | -0.014 | -0.1158 | No |
| 84 | Cd80 | 10128 | -0.016 | -0.1178 | No |
| 85 | Phlda1 | 10218 | -0.019 | -0.1219 | No |
| 86 | Irs2 | 10261 | -0.021 | -0.1236 | No |
| 87 | Dennd5a | 10416 | -0.026 | -0.1309 | No |
| 88 | Rhob | 10460 | -0.027 | -0.1324 | No |
| 89 | Tubb2a | 10545 | -0.030 | -0.1360 | No |
| 90 | Map3k8 | 10577 | -0.031 | -0.1368 | No |
| 91 | B4galt1 | 10613 | -0.032 | -0.1377 | No |
| 92 | Ehd1 | 10770 | -0.037 | -0.1448 | No |
| 93 | Slc16a6 | 10988 | -0.044 | -0.1550 | No |
| 94 | Tnfaip8 | 11216 | -0.052 | -0.1654 | No |
| 95 | Birc2 | 11228 | -0.053 | -0.1645 | No |
| 96 | Birc3 | 11348 | -0.057 | -0.1692 | No |
| 97 | Ets2 | 11370 | -0.058 | -0.1687 | No |
| 98 | Pdlim5 | 11433 | -0.061 | -0.1703 | No |
| 99 | Socs3 | 11463 | -0.062 | -0.1702 | No |
| 100 | Stat5a | 12020 | -0.083 | -0.1970 | No |
| 101 | Csf1 | 12129 | -0.087 | -0.2002 | No |
| 102 | Tiparp | 12217 | -0.090 | -0.2023 | No |
| 103 | Klf2 | 12295 | -0.092 | -0.2039 | No |
| 104 | Map2k3 | 12468 | -0.098 | -0.2102 | No |
| 105 | Il15ra | 12622 | -0.105 | -0.2153 | No |
| 106 | Per1 | 12720 | -0.109 | -0.2174 | No |
| 107 | Cd44 | 12726 | -0.109 | -0.2147 | No |
| 108 | Ifih1 | 12791 | -0.111 | -0.2150 | No |
| 109 | Atf3 | 12989 | -0.119 | -0.2221 | No |
| 110 | Nampt | 13224 | -0.126 | -0.2309 | No |
| 111 | Nfkbie | 13268 | -0.128 | -0.2296 | No |
| 112 | Plk2 | 13542 | -0.139 | -0.2401 | No |
| 113 | Tnfrsf9 | 13580 | -0.140 | -0.2383 | No |
| 114 | Rnf19b | 13934 | -0.156 | -0.2524 | No |
| 115 | Egr3 | 14038 | -0.161 | -0.2535 | Yes |
| 116 | Mcl1 | 14124 | -0.163 | -0.2535 | Yes |
| 117 | Nfil3 | 14126 | -0.163 | -0.2491 | Yes |
| 118 | Nfe2l2 | 14246 | -0.169 | -0.2507 | Yes |
| 119 | Fut4 | 14254 | -0.170 | -0.2464 | Yes |
| 120 | Kdm6b | 14261 | -0.170 | -0.2421 | Yes |
| 121 | Ifngr2 | 14296 | -0.171 | -0.2392 | Yes |
| 122 | Il1a | 14376 | -0.176 | -0.2386 | Yes |
| 123 | Fosb | 14426 | -0.177 | -0.2363 | Yes |
| 124 | Dnajb4 | 14430 | -0.177 | -0.2316 | Yes |
| 125 | Tank | 14518 | -0.182 | -0.2313 | Yes |
| 126 | Sat1 | 14716 | -0.190 | -0.2364 | Yes |
| 127 | Gch1 | 14855 | -0.196 | -0.2383 | Yes |
| 128 | Tnip2 | 14933 | -0.201 | -0.2368 | Yes |
| 129 | Eif1 | 14942 | -0.201 | -0.2318 | Yes |
| 130 | Btg3 | 15259 | -0.217 | -0.2424 | Yes |
| 131 | Pmepa1 | 15319 | -0.220 | -0.2395 | Yes |
| 132 | Cebpd | 15460 | -0.228 | -0.2406 | Yes |
| 133 | Sgk1 | 15513 | -0.230 | -0.2370 | Yes |
| 134 | Btg2 | 15551 | -0.232 | -0.2327 | Yes |
| 135 | Nfkb1 | 15723 | -0.242 | -0.2350 | Yes |
| 136 | Bmp2 | 15845 | -0.247 | -0.2346 | Yes |
| 137 | Maff | 15859 | -0.248 | -0.2285 | Yes |
| 138 | Myc | 15993 | -0.257 | -0.2285 | Yes |
| 139 | Mxd1 | 16060 | -0.262 | -0.2248 | Yes |
| 140 | Vegfa | 16093 | -0.264 | -0.2193 | Yes |
| 141 | Junb | 16128 | -0.266 | -0.2138 | Yes |
| 142 | Nfkbia | 16280 | -0.275 | -0.2142 | Yes |
| 143 | Bhlhe40 | 16341 | -0.279 | -0.2098 | Yes |
| 144 | Slc2a3 | 16419 | -0.284 | -0.2061 | Yes |
| 145 | Il6st | 17027 | -0.328 | -0.2289 | Yes |
| 146 | Tlr2 | 17030 | -0.328 | -0.2200 | Yes |
| 147 | Ninj1 | 17071 | -0.332 | -0.2131 | Yes |
| 148 | Kynu | 17150 | -0.338 | -0.2080 | Yes |
| 149 | Snn | 17186 | -0.342 | -0.2005 | Yes |
| 150 | Klf9 | 17517 | -0.368 | -0.2077 | Yes |
| 151 | Abca1 | 17531 | -0.369 | -0.1984 | Yes |
| 152 | Gadd45b | 17641 | -0.380 | -0.1937 | Yes |
| 153 | Nr4a3 | 17708 | -0.387 | -0.1866 | Yes |
| 154 | Slc2a6 | 17723 | -0.389 | -0.1768 | Yes |
| 155 | Ier5 | 18108 | -0.428 | -0.1852 | Yes |
| 156 | Irf1 | 18157 | -0.434 | -0.1759 | Yes |
| 157 | Tnfaip3 | 18481 | -0.480 | -0.1797 | Yes |
| 158 | Klf6 | 18487 | -0.482 | -0.1669 | Yes |
| 159 | Icam1 | 18721 | -0.534 | -0.1645 | Yes |
| 160 | Dusp4 | 18722 | -0.535 | -0.1500 | Yes |
| 161 | Gpr183 | 18771 | -0.545 | -0.1377 | Yes |
| 162 | Bcl3 | 18920 | -0.589 | -0.1294 | Yes |
| 163 | Phlda2 | 18943 | -0.595 | -0.1143 | Yes |
| 164 | Sqstm1 | 18967 | -0.605 | -0.0991 | Yes |
| 165 | Cd69 | 19181 | -0.739 | -0.0901 | Yes |
| 166 | Il1b | 19189 | -0.746 | -0.0702 | Yes |
| 167 | Il6 | 19208 | -0.767 | -0.0503 | Yes |
| 168 | Traf1 | 19235 | -0.805 | -0.0297 | Yes |
| 169 | Plaur | 19303 | -1.228 | 0.0002 | Yes |