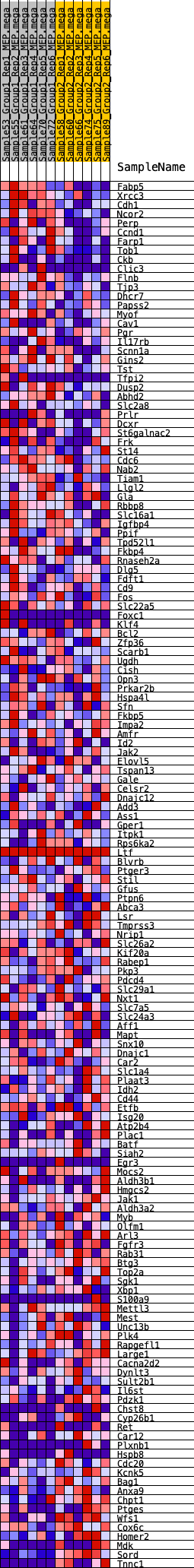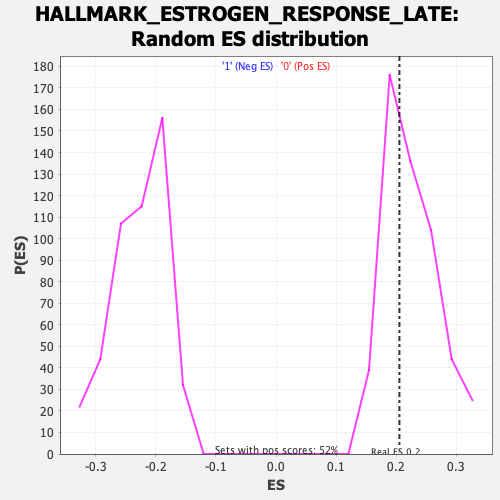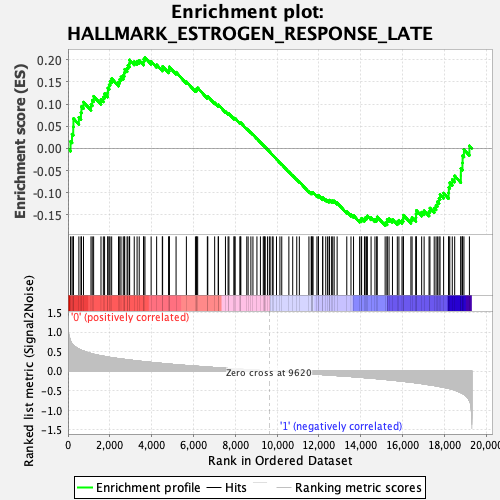
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.mega_Pheno.cls #Group1_versus_Group2.MEP.mega_Pheno.cls #Group1_versus_Group2_repos |
| Phenotype | MEP.mega_Pheno.cls#Group1_versus_Group2_repos |
| Upregulated in class | 0 |
| GeneSet | HALLMARK_ESTROGEN_RESPONSE_LATE |
| Enrichment Score (ES) | 0.20547189 |
| Normalized Enrichment Score (NES) | 0.9144846 |
| Nominal p-value | 0.60687023 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.994 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Fabp5 | 122 | 0.766 | 0.0159 | Yes |
| 2 | Xrcc3 | 193 | 0.695 | 0.0325 | Yes |
| 3 | Cdh1 | 250 | 0.661 | 0.0488 | Yes |
| 4 | Ncor2 | 260 | 0.657 | 0.0675 | Yes |
| 5 | Perp | 520 | 0.560 | 0.0703 | Yes |
| 6 | Ccnd1 | 620 | 0.533 | 0.0806 | Yes |
| 7 | Farp1 | 643 | 0.528 | 0.0948 | Yes |
| 8 | Tob1 | 740 | 0.510 | 0.1047 | Yes |
| 9 | Ckb | 1099 | 0.449 | 0.0990 | Yes |
| 10 | Clic3 | 1159 | 0.438 | 0.1087 | Yes |
| 11 | Flnb | 1225 | 0.429 | 0.1178 | Yes |
| 12 | Tjp3 | 1577 | 0.389 | 0.1108 | Yes |
| 13 | Dhcr7 | 1690 | 0.378 | 0.1160 | Yes |
| 14 | Papss2 | 1749 | 0.374 | 0.1238 | Yes |
| 15 | Myof | 1902 | 0.359 | 0.1263 | Yes |
| 16 | Cav1 | 1907 | 0.359 | 0.1366 | Yes |
| 17 | Pgr | 1968 | 0.353 | 0.1437 | Yes |
| 18 | Il17rb | 2010 | 0.349 | 0.1517 | Yes |
| 19 | Scnn1a | 2088 | 0.341 | 0.1576 | Yes |
| 20 | Gins2 | 2415 | 0.316 | 0.1498 | Yes |
| 21 | Tst | 2473 | 0.312 | 0.1559 | Yes |
| 22 | Tfpi2 | 2534 | 0.307 | 0.1617 | Yes |
| 23 | Dusp2 | 2642 | 0.302 | 0.1649 | Yes |
| 24 | Abhd2 | 2692 | 0.298 | 0.1710 | Yes |
| 25 | Slc2a8 | 2712 | 0.297 | 0.1787 | Yes |
| 26 | Prlr | 2825 | 0.290 | 0.1813 | Yes |
| 27 | Dcxr | 2865 | 0.286 | 0.1876 | Yes |
| 28 | St6galnac2 | 2940 | 0.282 | 0.1919 | Yes |
| 29 | Frk | 2947 | 0.282 | 0.1998 | Yes |
| 30 | St14 | 3167 | 0.269 | 0.1962 | Yes |
| 31 | Cdc6 | 3303 | 0.259 | 0.1967 | Yes |
| 32 | Nab2 | 3405 | 0.253 | 0.1988 | Yes |
| 33 | Tiam1 | 3612 | 0.240 | 0.1950 | Yes |
| 34 | Llgl2 | 3623 | 0.240 | 0.2015 | Yes |
| 35 | Gla | 3679 | 0.236 | 0.2055 | Yes |
| 36 | Rbbp8 | 3975 | 0.221 | 0.1965 | No |
| 37 | Slc16a1 | 4238 | 0.206 | 0.1888 | No |
| 38 | Igfbp4 | 4509 | 0.194 | 0.1804 | No |
| 39 | Ppif | 4524 | 0.193 | 0.1853 | No |
| 40 | Tpd52l1 | 4810 | 0.179 | 0.1756 | No |
| 41 | Fkbp4 | 4840 | 0.178 | 0.1793 | No |
| 42 | Rnaseh2a | 4843 | 0.178 | 0.1843 | No |
| 43 | Dlg5 | 5167 | 0.162 | 0.1722 | No |
| 44 | Fdft1 | 5660 | 0.142 | 0.1506 | No |
| 45 | Cd9 | 6093 | 0.124 | 0.1317 | No |
| 46 | Fos | 6145 | 0.122 | 0.1326 | No |
| 47 | Slc22a5 | 6158 | 0.121 | 0.1355 | No |
| 48 | Foxc1 | 6200 | 0.120 | 0.1368 | No |
| 49 | Klf4 | 6662 | 0.102 | 0.1157 | No |
| 50 | Bcl2 | 6683 | 0.101 | 0.1176 | No |
| 51 | Zfp36 | 7012 | 0.089 | 0.1031 | No |
| 52 | Scarb1 | 7182 | 0.083 | 0.0967 | No |
| 53 | Ugdh | 7193 | 0.083 | 0.0986 | No |
| 54 | Cish | 7528 | 0.071 | 0.0832 | No |
| 55 | Opn3 | 7654 | 0.066 | 0.0786 | No |
| 56 | Prkar2b | 7694 | 0.065 | 0.0785 | No |
| 57 | Hspa4l | 7934 | 0.056 | 0.0676 | No |
| 58 | Sfn | 7949 | 0.056 | 0.0685 | No |
| 59 | Fkbp5 | 7993 | 0.054 | 0.0678 | No |
| 60 | Impa2 | 8220 | 0.047 | 0.0574 | No |
| 61 | Amfr | 8239 | 0.046 | 0.0578 | No |
| 62 | Id2 | 8264 | 0.045 | 0.0578 | No |
| 63 | Jak2 | 8547 | 0.037 | 0.0442 | No |
| 64 | Elovl5 | 8612 | 0.034 | 0.0419 | No |
| 65 | Tspan13 | 8744 | 0.030 | 0.0359 | No |
| 66 | Gale | 8842 | 0.026 | 0.0316 | No |
| 67 | Celsr2 | 9033 | 0.019 | 0.0222 | No |
| 68 | Dnajc12 | 9200 | 0.014 | 0.0140 | No |
| 69 | Add3 | 9336 | 0.010 | 0.0072 | No |
| 70 | Ass1 | 9380 | 0.008 | 0.0052 | No |
| 71 | Gper1 | 9387 | 0.008 | 0.0051 | No |
| 72 | Itpk1 | 9430 | 0.006 | 0.0031 | No |
| 73 | Rps6ka2 | 9537 | 0.003 | -0.0023 | No |
| 74 | Ltf | 9625 | 0.000 | -0.0069 | No |
| 75 | Blvrb | 9664 | -0.000 | -0.0088 | No |
| 76 | Ptger3 | 9767 | -0.003 | -0.0141 | No |
| 77 | Stil | 9807 | -0.005 | -0.0160 | No |
| 78 | Gfus | 9969 | -0.011 | -0.0240 | No |
| 79 | Ptpn6 | 10132 | -0.016 | -0.0320 | No |
| 80 | Abca3 | 10220 | -0.019 | -0.0360 | No |
| 81 | Lsr | 10560 | -0.030 | -0.0528 | No |
| 82 | Tmprss3 | 10744 | -0.036 | -0.0613 | No |
| 83 | Nrip1 | 10937 | -0.042 | -0.0701 | No |
| 84 | Slc26a2 | 11061 | -0.047 | -0.0752 | No |
| 85 | Kif20a | 11519 | -0.064 | -0.0972 | No |
| 86 | Rabep1 | 11629 | -0.068 | -0.1009 | No |
| 87 | Pkp3 | 11640 | -0.068 | -0.0994 | No |
| 88 | Pdcd4 | 11701 | -0.070 | -0.1005 | No |
| 89 | Slc29a1 | 11710 | -0.070 | -0.0989 | No |
| 90 | Nxt1 | 11902 | -0.078 | -0.1066 | No |
| 91 | Slc7a5 | 11970 | -0.081 | -0.1078 | No |
| 92 | Slc24a3 | 11973 | -0.081 | -0.1055 | No |
| 93 | Aff1 | 12175 | -0.089 | -0.1134 | No |
| 94 | Mapt | 12180 | -0.089 | -0.1111 | No |
| 95 | Snx10 | 12324 | -0.093 | -0.1158 | No |
| 96 | Dnajc1 | 12412 | -0.096 | -0.1175 | No |
| 97 | Car2 | 12485 | -0.099 | -0.1184 | No |
| 98 | Slc1a4 | 12506 | -0.100 | -0.1166 | No |
| 99 | Plaat3 | 12604 | -0.105 | -0.1186 | No |
| 100 | Idh2 | 12657 | -0.107 | -0.1182 | No |
| 101 | Cd44 | 12726 | -0.109 | -0.1186 | No |
| 102 | Etfb | 12868 | -0.114 | -0.1226 | No |
| 103 | Isg20 | 13329 | -0.131 | -0.1428 | No |
| 104 | Atp2b4 | 13530 | -0.138 | -0.1492 | No |
| 105 | Plac1 | 13652 | -0.143 | -0.1514 | No |
| 106 | Batf | 13943 | -0.157 | -0.1620 | No |
| 107 | Siah2 | 14017 | -0.160 | -0.1611 | No |
| 108 | Egr3 | 14038 | -0.161 | -0.1575 | No |
| 109 | Mocs2 | 14174 | -0.166 | -0.1597 | No |
| 110 | Aldh3b1 | 14207 | -0.168 | -0.1565 | No |
| 111 | Hmgcs2 | 14275 | -0.170 | -0.1550 | No |
| 112 | Jak1 | 14314 | -0.172 | -0.1520 | No |
| 113 | Aldh3a2 | 14491 | -0.180 | -0.1559 | No |
| 114 | Myb | 14676 | -0.188 | -0.1601 | No |
| 115 | Olfm1 | 14758 | -0.192 | -0.1587 | No |
| 116 | Arl3 | 14784 | -0.193 | -0.1544 | No |
| 117 | Fgfr3 | 15163 | -0.213 | -0.1680 | No |
| 118 | Rab31 | 15255 | -0.217 | -0.1664 | No |
| 119 | Btg3 | 15259 | -0.217 | -0.1602 | No |
| 120 | Top2a | 15350 | -0.222 | -0.1585 | No |
| 121 | Sgk1 | 15513 | -0.230 | -0.1602 | No |
| 122 | Xbp1 | 15745 | -0.243 | -0.1652 | No |
| 123 | S100a9 | 15821 | -0.246 | -0.1620 | No |
| 124 | Mettl3 | 15969 | -0.255 | -0.1622 | No |
| 125 | Mest | 16028 | -0.259 | -0.1577 | No |
| 126 | Unc13b | 16037 | -0.260 | -0.1506 | No |
| 127 | Plk4 | 16395 | -0.283 | -0.1610 | No |
| 128 | Rapgefl1 | 16451 | -0.287 | -0.1555 | No |
| 129 | Large1 | 16634 | -0.299 | -0.1563 | No |
| 130 | Cacna2d2 | 16640 | -0.300 | -0.1479 | No |
| 131 | Dynlt3 | 16656 | -0.301 | -0.1399 | No |
| 132 | Sult2b1 | 16909 | -0.317 | -0.1438 | No |
| 133 | Il6st | 17027 | -0.328 | -0.1404 | No |
| 134 | Pdzk1 | 17263 | -0.347 | -0.1425 | No |
| 135 | Chst8 | 17305 | -0.350 | -0.1345 | No |
| 136 | Cyp26b1 | 17509 | -0.367 | -0.1344 | No |
| 137 | Ret | 17597 | -0.376 | -0.1280 | No |
| 138 | Car12 | 17661 | -0.382 | -0.1201 | No |
| 139 | Plxnb1 | 17739 | -0.390 | -0.1128 | No |
| 140 | Hspb8 | 17788 | -0.396 | -0.1038 | No |
| 141 | Cdc20 | 17959 | -0.412 | -0.1007 | No |
| 142 | Kcnk5 | 18198 | -0.438 | -0.1004 | No |
| 143 | Bag1 | 18208 | -0.440 | -0.0880 | No |
| 144 | Anxa9 | 18254 | -0.447 | -0.0774 | No |
| 145 | Chpt1 | 18373 | -0.463 | -0.0700 | No |
| 146 | Ptges | 18488 | -0.484 | -0.0619 | No |
| 147 | Wfs1 | 18783 | -0.549 | -0.0613 | No |
| 148 | Cox6c | 18784 | -0.550 | -0.0453 | No |
| 149 | Homer2 | 18859 | -0.572 | -0.0325 | No |
| 150 | Mdk | 18867 | -0.574 | -0.0162 | No |
| 151 | Sord | 18931 | -0.591 | -0.0023 | No |
| 152 | Tnnc1 | 19192 | -0.750 | 0.0060 | No |