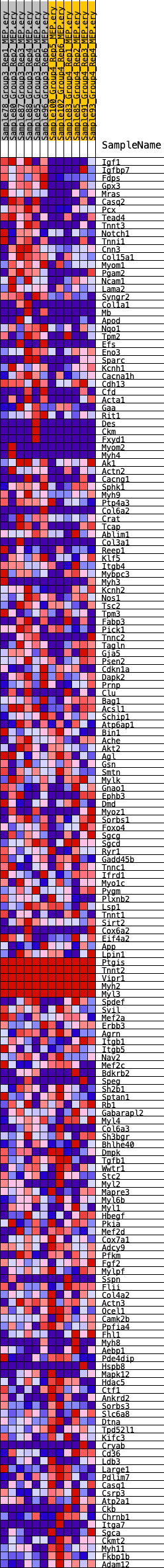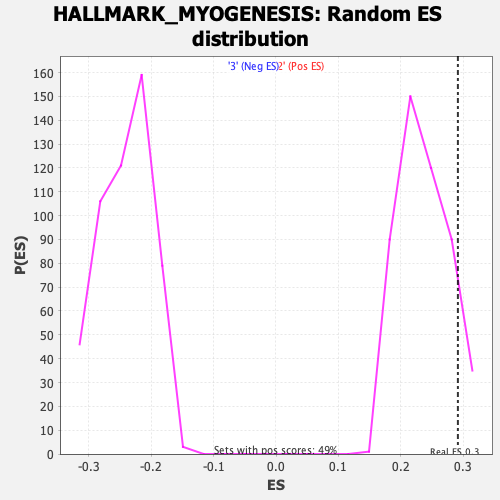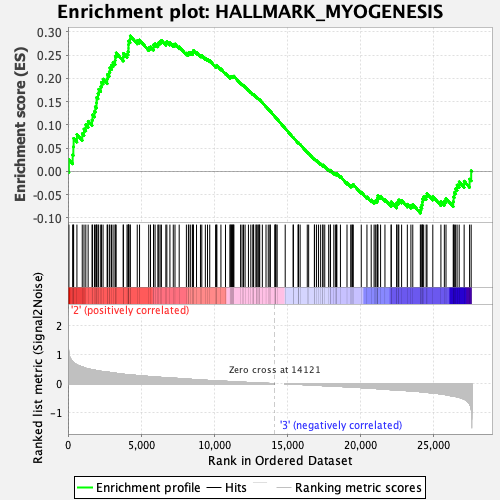
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.ery_Pheno.cls #Group3_versus_Group4.MEP.ery_Pheno.cls #Group3_versus_Group4_repos |
| Phenotype | MEP.ery_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_MYOGENESIS |
| Enrichment Score (ES) | 0.2914589 |
| Normalized Enrichment Score (NES) | 1.2229674 |
| Nominal p-value | 0.10288066 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.95 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Igf1 | 62 | 1.068 | 0.0254 | Yes |
| 2 | Igfbp7 | 319 | 0.756 | 0.0356 | Yes |
| 3 | Fdps | 365 | 0.737 | 0.0530 | Yes |
| 4 | Gpx3 | 385 | 0.726 | 0.0710 | Yes |
| 5 | Mras | 610 | 0.649 | 0.0796 | Yes |
| 6 | Casq2 | 974 | 0.574 | 0.0812 | Yes |
| 7 | Pcx | 1093 | 0.554 | 0.0912 | Yes |
| 8 | Tead4 | 1218 | 0.530 | 0.1004 | Yes |
| 9 | Tnnt3 | 1376 | 0.510 | 0.1079 | Yes |
| 10 | Notch1 | 1643 | 0.481 | 0.1106 | Yes |
| 11 | Tnni1 | 1678 | 0.477 | 0.1217 | Yes |
| 12 | Cnn3 | 1810 | 0.468 | 0.1290 | Yes |
| 13 | Col15a1 | 1862 | 0.463 | 0.1391 | Yes |
| 14 | Myom1 | 1949 | 0.452 | 0.1477 | Yes |
| 15 | Pgam2 | 1957 | 0.450 | 0.1590 | Yes |
| 16 | Ncam1 | 2050 | 0.438 | 0.1670 | Yes |
| 17 | Lama2 | 2097 | 0.434 | 0.1765 | Yes |
| 18 | Syngr2 | 2238 | 0.427 | 0.1825 | Yes |
| 19 | Col1a1 | 2287 | 0.424 | 0.1917 | Yes |
| 20 | Mb | 2397 | 0.420 | 0.1986 | Yes |
| 21 | Apod | 2677 | 0.405 | 0.1988 | Yes |
| 22 | Nqo1 | 2696 | 0.403 | 0.2086 | Yes |
| 23 | Tpm2 | 2819 | 0.392 | 0.2143 | Yes |
| 24 | Efs | 2863 | 0.387 | 0.2227 | Yes |
| 25 | Eno3 | 2975 | 0.384 | 0.2286 | Yes |
| 26 | Sparc | 3080 | 0.374 | 0.2345 | Yes |
| 27 | Kcnh1 | 3221 | 0.363 | 0.2387 | Yes |
| 28 | Cacna1h | 3230 | 0.362 | 0.2478 | Yes |
| 29 | Cdh13 | 3291 | 0.357 | 0.2549 | Yes |
| 30 | Cfd | 3776 | 0.326 | 0.2456 | Yes |
| 31 | Acta1 | 3784 | 0.326 | 0.2538 | Yes |
| 32 | Gaa | 4035 | 0.308 | 0.2527 | Yes |
| 33 | Rit1 | 4104 | 0.302 | 0.2580 | Yes |
| 34 | Des | 4134 | 0.300 | 0.2647 | Yes |
| 35 | Ckm | 4135 | 0.300 | 0.2724 | Yes |
| 36 | Fxyd1 | 4139 | 0.300 | 0.2801 | Yes |
| 37 | Myom2 | 4241 | 0.300 | 0.2841 | Yes |
| 38 | Myh4 | 4254 | 0.300 | 0.2915 | Yes |
| 39 | Ak1 | 4733 | 0.280 | 0.2813 | No |
| 40 | Actn2 | 4890 | 0.274 | 0.2826 | No |
| 41 | Cacng1 | 5519 | 0.246 | 0.2661 | No |
| 42 | Sphk1 | 5639 | 0.241 | 0.2680 | No |
| 43 | Myh9 | 5847 | 0.233 | 0.2665 | No |
| 44 | Ptp4a3 | 5848 | 0.232 | 0.2725 | No |
| 45 | Col6a2 | 5952 | 0.227 | 0.2746 | No |
| 46 | Crat | 6143 | 0.227 | 0.2735 | No |
| 47 | Tcap | 6202 | 0.224 | 0.2772 | No |
| 48 | Ablim1 | 6297 | 0.219 | 0.2794 | No |
| 49 | Col3a1 | 6386 | 0.216 | 0.2818 | No |
| 50 | Reep1 | 6689 | 0.212 | 0.2763 | No |
| 51 | Klf5 | 6757 | 0.209 | 0.2792 | No |
| 52 | Itgb4 | 6966 | 0.200 | 0.2768 | No |
| 53 | Mybpc3 | 7196 | 0.192 | 0.2734 | No |
| 54 | Myh3 | 7313 | 0.187 | 0.2740 | No |
| 55 | Kcnh2 | 7600 | 0.177 | 0.2682 | No |
| 56 | Nos1 | 8092 | 0.160 | 0.2544 | No |
| 57 | Tsc2 | 8229 | 0.154 | 0.2534 | No |
| 58 | Tpm3 | 8264 | 0.152 | 0.2561 | No |
| 59 | Fabp3 | 8385 | 0.149 | 0.2556 | No |
| 60 | Pick1 | 8516 | 0.145 | 0.2546 | No |
| 61 | Tnnc2 | 8545 | 0.144 | 0.2573 | No |
| 62 | Tagln | 8571 | 0.144 | 0.2601 | No |
| 63 | Gja5 | 8785 | 0.136 | 0.2559 | No |
| 64 | Psen2 | 9054 | 0.128 | 0.2494 | No |
| 65 | Cdkn1a | 9146 | 0.125 | 0.2493 | No |
| 66 | Dapk2 | 9388 | 0.117 | 0.2435 | No |
| 67 | Prnp | 9536 | 0.113 | 0.2411 | No |
| 68 | Clu | 9688 | 0.108 | 0.2384 | No |
| 69 | Bag1 | 10088 | 0.097 | 0.2263 | No |
| 70 | Acsl1 | 10159 | 0.095 | 0.2263 | No |
| 71 | Schip1 | 10171 | 0.095 | 0.2283 | No |
| 72 | Atp6ap1 | 10453 | 0.089 | 0.2204 | No |
| 73 | Bin1 | 10765 | 0.081 | 0.2111 | No |
| 74 | Ache | 11076 | 0.073 | 0.2017 | No |
| 75 | Akt2 | 11086 | 0.073 | 0.2032 | No |
| 76 | Agl | 11146 | 0.071 | 0.2029 | No |
| 77 | Gsn | 11172 | 0.070 | 0.2038 | No |
| 78 | Smtn | 11241 | 0.069 | 0.2031 | No |
| 79 | Mylk | 11266 | 0.068 | 0.2040 | No |
| 80 | Gnao1 | 11318 | 0.067 | 0.2039 | No |
| 81 | Ephb3 | 11337 | 0.066 | 0.2049 | No |
| 82 | Dmd | 11798 | 0.053 | 0.1895 | No |
| 83 | Myoz1 | 11898 | 0.051 | 0.1872 | No |
| 84 | Sorbs1 | 11998 | 0.048 | 0.1849 | No |
| 85 | Foxo4 | 12104 | 0.045 | 0.1822 | No |
| 86 | Sgcg | 12334 | 0.039 | 0.1749 | No |
| 87 | Sgcd | 12496 | 0.035 | 0.1699 | No |
| 88 | Ryr1 | 12637 | 0.032 | 0.1656 | No |
| 89 | Gadd45b | 12665 | 0.031 | 0.1654 | No |
| 90 | Tnnc1 | 12691 | 0.030 | 0.1653 | No |
| 91 | Ifrd1 | 12855 | 0.026 | 0.1600 | No |
| 92 | Myo1c | 12866 | 0.026 | 0.1603 | No |
| 93 | Pygm | 12980 | 0.023 | 0.1568 | No |
| 94 | Plxnb2 | 13000 | 0.023 | 0.1567 | No |
| 95 | Lsp1 | 13079 | 0.021 | 0.1544 | No |
| 96 | Tnnt1 | 13105 | 0.020 | 0.1540 | No |
| 97 | Sirt2 | 13284 | 0.016 | 0.1479 | No |
| 98 | Cox6a2 | 13535 | 0.012 | 0.1391 | No |
| 99 | Eif4a2 | 13688 | 0.009 | 0.1338 | No |
| 100 | App | 13799 | 0.007 | 0.1300 | No |
| 101 | Lpin1 | 13830 | 0.006 | 0.1291 | No |
| 102 | Ptgis | 14136 | 0.000 | 0.1179 | No |
| 103 | Tnnt2 | 14168 | 0.000 | 0.1168 | No |
| 104 | Vipr1 | 14204 | 0.000 | 0.1155 | No |
| 105 | Myh2 | 14206 | 0.000 | 0.1155 | No |
| 106 | Myl3 | 14296 | 0.000 | 0.1123 | No |
| 107 | Spdef | 14847 | -0.001 | 0.0922 | No |
| 108 | Svil | 15387 | -0.012 | 0.0729 | No |
| 109 | Mef2a | 15408 | -0.013 | 0.0725 | No |
| 110 | Erbb3 | 15725 | -0.020 | 0.0615 | No |
| 111 | Agrn | 15751 | -0.021 | 0.0611 | No |
| 112 | Itgb1 | 15757 | -0.021 | 0.0614 | No |
| 113 | Itgb5 | 15867 | -0.024 | 0.0581 | No |
| 114 | Nav2 | 16355 | -0.037 | 0.0413 | No |
| 115 | Mef2c | 16450 | -0.039 | 0.0388 | No |
| 116 | Bdkrb2 | 16840 | -0.049 | 0.0259 | No |
| 117 | Speg | 16855 | -0.049 | 0.0267 | No |
| 118 | Sh2b1 | 16989 | -0.052 | 0.0232 | No |
| 119 | Sptan1 | 17126 | -0.056 | 0.0196 | No |
| 120 | Rb1 | 17269 | -0.060 | 0.0160 | No |
| 121 | Gabarapl2 | 17419 | -0.064 | 0.0122 | No |
| 122 | Myl4 | 17423 | -0.064 | 0.0138 | No |
| 123 | Col6a3 | 17550 | -0.067 | 0.0109 | No |
| 124 | Sh3bgr | 17818 | -0.072 | 0.0030 | No |
| 125 | Bhlhe40 | 17930 | -0.075 | 0.0009 | No |
| 126 | Dmpk | 17940 | -0.075 | 0.0025 | No |
| 127 | Tgfb1 | 18170 | -0.082 | -0.0037 | No |
| 128 | Wwtr1 | 18277 | -0.084 | -0.0054 | No |
| 129 | Stc2 | 18352 | -0.086 | -0.0059 | No |
| 130 | Myl2 | 18378 | -0.086 | -0.0046 | No |
| 131 | Mapre3 | 18622 | -0.093 | -0.0110 | No |
| 132 | Myl6b | 19071 | -0.105 | -0.0246 | No |
| 133 | Myl1 | 19325 | -0.113 | -0.0310 | No |
| 134 | Hbegf | 19399 | -0.115 | -0.0306 | No |
| 135 | Pkia | 19450 | -0.116 | -0.0295 | No |
| 136 | Mef2d | 19499 | -0.117 | -0.0282 | No |
| 137 | Cox7a1 | 20046 | -0.134 | -0.0447 | No |
| 138 | Adcy9 | 20434 | -0.146 | -0.0550 | No |
| 139 | Pfkm | 20725 | -0.155 | -0.0616 | No |
| 140 | Fgf2 | 20922 | -0.161 | -0.0645 | No |
| 141 | Mylpf | 21007 | -0.164 | -0.0634 | No |
| 142 | Sspn | 21119 | -0.168 | -0.0631 | No |
| 143 | Flii | 21153 | -0.169 | -0.0599 | No |
| 144 | Col4a2 | 21167 | -0.170 | -0.0560 | No |
| 145 | Actn3 | 21183 | -0.170 | -0.0521 | No |
| 146 | Ocel1 | 21366 | -0.177 | -0.0542 | No |
| 147 | Camk2b | 21665 | -0.188 | -0.0602 | No |
| 148 | Ppfia4 | 22089 | -0.203 | -0.0704 | No |
| 149 | Fhl1 | 22094 | -0.204 | -0.0653 | No |
| 150 | Myh8 | 22469 | -0.217 | -0.0733 | No |
| 151 | Aebp1 | 22474 | -0.217 | -0.0678 | No |
| 152 | Pde4dip | 22572 | -0.221 | -0.0657 | No |
| 153 | Hspb8 | 22613 | -0.221 | -0.0614 | No |
| 154 | Mapk12 | 22803 | -0.222 | -0.0626 | No |
| 155 | Hdac5 | 23194 | -0.239 | -0.0706 | No |
| 156 | Ctf1 | 23439 | -0.249 | -0.0731 | No |
| 157 | Ankrd2 | 23567 | -0.256 | -0.0711 | No |
| 158 | Sorbs3 | 24084 | -0.274 | -0.0828 | No |
| 159 | Slc6a8 | 24123 | -0.277 | -0.0770 | No |
| 160 | Dtna | 24191 | -0.280 | -0.0722 | No |
| 161 | Tpd52l1 | 24216 | -0.282 | -0.0658 | No |
| 162 | Kifc3 | 24239 | -0.283 | -0.0593 | No |
| 163 | Cryab | 24311 | -0.287 | -0.0545 | No |
| 164 | Cd36 | 24481 | -0.294 | -0.0530 | No |
| 165 | Ldb3 | 24539 | -0.298 | -0.0474 | No |
| 166 | Large1 | 24936 | -0.316 | -0.0537 | No |
| 167 | Pdlim7 | 25490 | -0.354 | -0.0647 | No |
| 168 | Casq1 | 25725 | -0.369 | -0.0637 | No |
| 169 | Csrp3 | 25838 | -0.378 | -0.0580 | No |
| 170 | Atp2a1 | 26332 | -0.424 | -0.0650 | No |
| 171 | Ckb | 26358 | -0.428 | -0.0549 | No |
| 172 | Chrnb1 | 26407 | -0.430 | -0.0455 | No |
| 173 | Itga7 | 26495 | -0.441 | -0.0373 | No |
| 174 | Sgca | 26596 | -0.450 | -0.0293 | No |
| 175 | Ckmt2 | 26739 | -0.467 | -0.0224 | No |
| 176 | Myh11 | 27078 | -0.530 | -0.0210 | No |
| 177 | Fkbp1b | 27450 | -0.693 | -0.0167 | No |
| 178 | Adam12 | 27559 | -0.858 | 0.0016 | No |