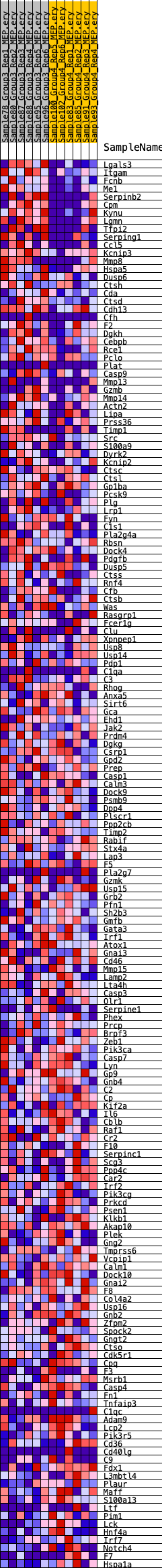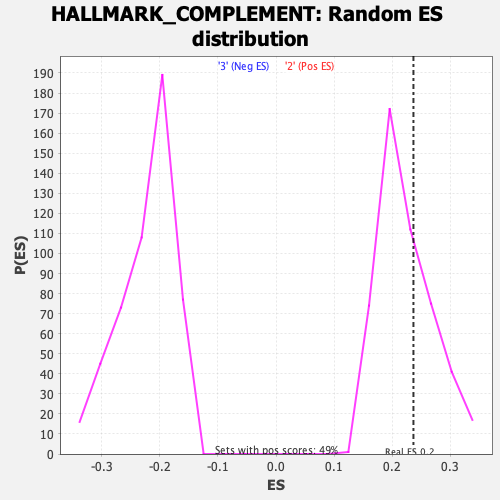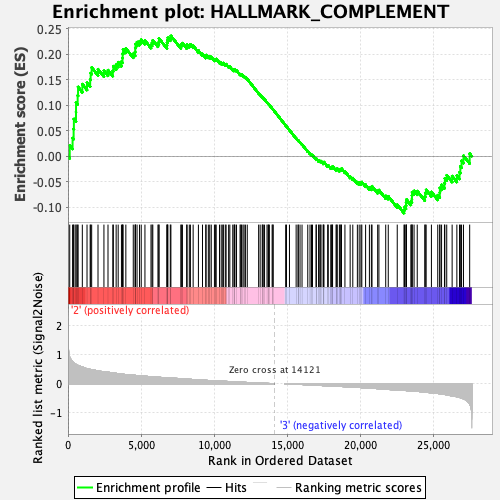
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.ery_Pheno.cls #Group3_versus_Group4.MEP.ery_Pheno.cls #Group3_versus_Group4_repos |
| Phenotype | MEP.ery_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_COMPLEMENT |
| Enrichment Score (ES) | 0.23636648 |
| Normalized Enrichment Score (NES) | 1.0582876 |
| Nominal p-value | 0.35365853 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.996 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Lgals3 | 118 | 0.914 | 0.0214 | Yes |
| 2 | Itgam | 313 | 0.761 | 0.0357 | Yes |
| 3 | Fcnb | 384 | 0.727 | 0.0536 | Yes |
| 4 | Me1 | 402 | 0.721 | 0.0733 | Yes |
| 5 | Serpinb2 | 546 | 0.668 | 0.0868 | Yes |
| 6 | Cpm | 550 | 0.666 | 0.1055 | Yes |
| 7 | Kynu | 660 | 0.637 | 0.1194 | Yes |
| 8 | Lgmn | 688 | 0.630 | 0.1361 | Yes |
| 9 | Tfpi2 | 979 | 0.573 | 0.1417 | Yes |
| 10 | Serping1 | 1300 | 0.517 | 0.1445 | Yes |
| 11 | Ccl5 | 1515 | 0.496 | 0.1507 | Yes |
| 12 | Kcnip3 | 1549 | 0.490 | 0.1632 | Yes |
| 13 | Mmp8 | 1618 | 0.484 | 0.1744 | Yes |
| 14 | Hspa5 | 2051 | 0.438 | 0.1709 | Yes |
| 15 | Dusp6 | 2456 | 0.420 | 0.1680 | Yes |
| 16 | Ctsh | 2734 | 0.399 | 0.1691 | Yes |
| 17 | Cda | 3064 | 0.376 | 0.1677 | Yes |
| 18 | Ctsd | 3099 | 0.372 | 0.1769 | Yes |
| 19 | Cdh13 | 3291 | 0.357 | 0.1800 | Yes |
| 20 | Cfh | 3430 | 0.346 | 0.1847 | Yes |
| 21 | F2 | 3653 | 0.335 | 0.1860 | Yes |
| 22 | Dgkh | 3723 | 0.329 | 0.1928 | Yes |
| 23 | Cebpb | 3735 | 0.328 | 0.2016 | Yes |
| 24 | Rce1 | 3764 | 0.327 | 0.2098 | Yes |
| 25 | Pclo | 3954 | 0.314 | 0.2117 | Yes |
| 26 | Plat | 4461 | 0.292 | 0.2015 | Yes |
| 27 | Casp9 | 4588 | 0.287 | 0.2049 | Yes |
| 28 | Mmp13 | 4598 | 0.287 | 0.2127 | Yes |
| 29 | Gzmb | 4612 | 0.286 | 0.2203 | Yes |
| 30 | Mmp14 | 4720 | 0.281 | 0.2243 | Yes |
| 31 | Actn2 | 4890 | 0.274 | 0.2258 | Yes |
| 32 | Lipa | 5006 | 0.267 | 0.2291 | Yes |
| 33 | Prss36 | 5259 | 0.261 | 0.2273 | Yes |
| 34 | Timp1 | 5673 | 0.240 | 0.2189 | Yes |
| 35 | Src | 5746 | 0.236 | 0.2230 | Yes |
| 36 | S100a9 | 5797 | 0.233 | 0.2277 | Yes |
| 37 | Dyrk2 | 6149 | 0.226 | 0.2213 | Yes |
| 38 | Kcnip2 | 6213 | 0.223 | 0.2252 | Yes |
| 39 | Ctsc | 6217 | 0.223 | 0.2314 | Yes |
| 40 | Ctsl | 6763 | 0.209 | 0.2174 | Yes |
| 41 | Gp1ba | 6771 | 0.208 | 0.2230 | Yes |
| 42 | Pcsk9 | 6799 | 0.207 | 0.2279 | Yes |
| 43 | Plg | 6828 | 0.206 | 0.2326 | Yes |
| 44 | Lrp1 | 6992 | 0.199 | 0.2323 | Yes |
| 45 | Fyn | 7033 | 0.198 | 0.2364 | Yes |
| 46 | C1s1 | 7714 | 0.172 | 0.2164 | No |
| 47 | Pla2g4a | 7750 | 0.171 | 0.2200 | No |
| 48 | Rbsn | 7822 | 0.168 | 0.2221 | No |
| 49 | Dock4 | 8113 | 0.159 | 0.2160 | No |
| 50 | Pdgfb | 8139 | 0.158 | 0.2195 | No |
| 51 | Dusp5 | 8291 | 0.151 | 0.2183 | No |
| 52 | Ctss | 8367 | 0.149 | 0.2197 | No |
| 53 | Rnf4 | 8554 | 0.144 | 0.2170 | No |
| 54 | Cfb | 8914 | 0.132 | 0.2076 | No |
| 55 | Ctsb | 9193 | 0.124 | 0.2010 | No |
| 56 | Was | 9419 | 0.116 | 0.1960 | No |
| 57 | Rasgrp1 | 9435 | 0.116 | 0.1987 | No |
| 58 | Fcer1g | 9592 | 0.111 | 0.1962 | No |
| 59 | Clu | 9688 | 0.108 | 0.1958 | No |
| 60 | Xpnpep1 | 9792 | 0.105 | 0.1950 | No |
| 61 | Usp8 | 10009 | 0.099 | 0.1899 | No |
| 62 | Usp14 | 10074 | 0.097 | 0.1903 | No |
| 63 | Pdp1 | 10129 | 0.096 | 0.1910 | No |
| 64 | C1qa | 10357 | 0.091 | 0.1853 | No |
| 65 | C3 | 10473 | 0.088 | 0.1836 | No |
| 66 | Rhog | 10579 | 0.085 | 0.1822 | No |
| 67 | Anxa5 | 10620 | 0.084 | 0.1831 | No |
| 68 | Sirt6 | 10751 | 0.081 | 0.1806 | No |
| 69 | Gca | 10806 | 0.080 | 0.1809 | No |
| 70 | Ehd1 | 10980 | 0.075 | 0.1767 | No |
| 71 | Jak2 | 11062 | 0.073 | 0.1758 | No |
| 72 | Prdm4 | 11262 | 0.068 | 0.1705 | No |
| 73 | Dgkg | 11347 | 0.066 | 0.1693 | No |
| 74 | Csrp1 | 11391 | 0.065 | 0.1695 | No |
| 75 | Gpd2 | 11405 | 0.064 | 0.1708 | No |
| 76 | Prep | 11519 | 0.061 | 0.1684 | No |
| 77 | Casp1 | 11761 | 0.054 | 0.1612 | No |
| 78 | Calm3 | 11811 | 0.053 | 0.1609 | No |
| 79 | Dock9 | 11862 | 0.052 | 0.1605 | No |
| 80 | Psmb9 | 11959 | 0.049 | 0.1584 | No |
| 81 | Dpp4 | 12036 | 0.047 | 0.1570 | No |
| 82 | Plscr1 | 12127 | 0.045 | 0.1549 | No |
| 83 | Ppp2cb | 12253 | 0.041 | 0.1515 | No |
| 84 | Timp2 | 13031 | 0.022 | 0.1238 | No |
| 85 | Rabif | 13134 | 0.019 | 0.1207 | No |
| 86 | Stx4a | 13275 | 0.016 | 0.1160 | No |
| 87 | Lap3 | 13317 | 0.015 | 0.1149 | No |
| 88 | F5 | 13337 | 0.015 | 0.1147 | No |
| 89 | Pla2g7 | 13434 | 0.013 | 0.1115 | No |
| 90 | Gzmk | 13604 | 0.011 | 0.1057 | No |
| 91 | Usp15 | 13686 | 0.009 | 0.1030 | No |
| 92 | Grb2 | 13701 | 0.009 | 0.1027 | No |
| 93 | Pfn1 | 13765 | 0.008 | 0.1007 | No |
| 94 | Sh2b3 | 13953 | 0.003 | 0.0939 | No |
| 95 | Gmfb | 14002 | 0.002 | 0.0923 | No |
| 96 | Gata3 | 14022 | 0.002 | 0.0916 | No |
| 97 | Irf1 | 14879 | -0.001 | 0.0604 | No |
| 98 | Atox1 | 14934 | -0.003 | 0.0586 | No |
| 99 | Gnai3 | 15138 | -0.008 | 0.0514 | No |
| 100 | Cd46 | 15594 | -0.017 | 0.0352 | No |
| 101 | Mmp15 | 15730 | -0.020 | 0.0309 | No |
| 102 | Lamp2 | 15736 | -0.020 | 0.0313 | No |
| 103 | Lta4h | 15860 | -0.024 | 0.0275 | No |
| 104 | Casp3 | 15997 | -0.027 | 0.0233 | No |
| 105 | Olr1 | 16389 | -0.037 | 0.0101 | No |
| 106 | Serpine1 | 16543 | -0.041 | 0.0056 | No |
| 107 | Phex | 16646 | -0.044 | 0.0031 | No |
| 108 | Prcp | 16712 | -0.045 | 0.0021 | No |
| 109 | Brpf3 | 16958 | -0.051 | -0.0054 | No |
| 110 | Zeb1 | 16973 | -0.052 | -0.0045 | No |
| 111 | Pik3ca | 17128 | -0.056 | -0.0085 | No |
| 112 | Casp7 | 17168 | -0.057 | -0.0084 | No |
| 113 | Lyn | 17244 | -0.059 | -0.0094 | No |
| 114 | Gp9 | 17295 | -0.060 | -0.0096 | No |
| 115 | Gnb4 | 17447 | -0.065 | -0.0133 | No |
| 116 | C2 | 17452 | -0.065 | -0.0116 | No |
| 117 | Cp | 17497 | -0.066 | -0.0113 | No |
| 118 | Kif2a | 17750 | -0.071 | -0.0185 | No |
| 119 | Il6 | 17789 | -0.072 | -0.0179 | No |
| 120 | Cblb | 17961 | -0.076 | -0.0220 | No |
| 121 | Raf1 | 18020 | -0.078 | -0.0219 | No |
| 122 | Cr2 | 18058 | -0.079 | -0.0211 | No |
| 123 | F10 | 18084 | -0.079 | -0.0197 | No |
| 124 | Serpinc1 | 18323 | -0.085 | -0.0260 | No |
| 125 | Scg3 | 18371 | -0.086 | -0.0253 | No |
| 126 | Ppp4c | 18401 | -0.087 | -0.0239 | No |
| 127 | Car2 | 18550 | -0.091 | -0.0268 | No |
| 128 | Irf2 | 18605 | -0.092 | -0.0261 | No |
| 129 | Pik3cg | 18657 | -0.094 | -0.0254 | No |
| 130 | Prkcd | 18695 | -0.095 | -0.0240 | No |
| 131 | Psen1 | 18929 | -0.101 | -0.0297 | No |
| 132 | Klkb1 | 19286 | -0.111 | -0.0395 | No |
| 133 | Akap10 | 19467 | -0.116 | -0.0428 | No |
| 134 | Plek | 19779 | -0.125 | -0.0507 | No |
| 135 | Gng2 | 19903 | -0.129 | -0.0515 | No |
| 136 | Tmprss6 | 20023 | -0.133 | -0.0521 | No |
| 137 | Vcpip1 | 20083 | -0.135 | -0.0505 | No |
| 138 | Calm1 | 20342 | -0.143 | -0.0559 | No |
| 139 | Dock10 | 20602 | -0.152 | -0.0611 | No |
| 140 | Gnai2 | 20737 | -0.156 | -0.0616 | No |
| 141 | F8 | 20780 | -0.157 | -0.0587 | No |
| 142 | Col4a2 | 21167 | -0.170 | -0.0680 | No |
| 143 | Usp16 | 21252 | -0.172 | -0.0662 | No |
| 144 | Gnb2 | 21717 | -0.190 | -0.0778 | No |
| 145 | Zfpm2 | 21891 | -0.197 | -0.0786 | No |
| 146 | Spock2 | 22505 | -0.219 | -0.0948 | No |
| 147 | Gngt2 | 22967 | -0.229 | -0.1051 | No |
| 148 | Ctso | 23005 | -0.231 | -0.1000 | No |
| 149 | Cdk5r1 | 23100 | -0.235 | -0.0968 | No |
| 150 | Cpq | 23112 | -0.236 | -0.0906 | No |
| 151 | F3 | 23137 | -0.237 | -0.0848 | No |
| 152 | Msrb1 | 23436 | -0.249 | -0.0886 | No |
| 153 | Casp4 | 23507 | -0.253 | -0.0841 | No |
| 154 | Fn1 | 23511 | -0.253 | -0.0771 | No |
| 155 | Tnfaip3 | 23521 | -0.254 | -0.0703 | No |
| 156 | C1qc | 23655 | -0.260 | -0.0678 | No |
| 157 | Adam9 | 23872 | -0.264 | -0.0683 | No |
| 158 | Lcp2 | 24389 | -0.290 | -0.0789 | No |
| 159 | Pik3r5 | 24419 | -0.291 | -0.0718 | No |
| 160 | Cd36 | 24481 | -0.294 | -0.0657 | No |
| 161 | Cd40lg | 24841 | -0.314 | -0.0700 | No |
| 162 | C9 | 25274 | -0.336 | -0.0763 | No |
| 163 | Fdx1 | 25412 | -0.348 | -0.0715 | No |
| 164 | L3mbtl4 | 25419 | -0.349 | -0.0619 | No |
| 165 | Plaur | 25535 | -0.358 | -0.0561 | No |
| 166 | Maff | 25733 | -0.370 | -0.0529 | No |
| 167 | S100a13 | 25759 | -0.372 | -0.0433 | No |
| 168 | Ltf | 25888 | -0.382 | -0.0372 | No |
| 169 | Pim1 | 26257 | -0.420 | -0.0388 | No |
| 170 | Lck | 26580 | -0.449 | -0.0379 | No |
| 171 | Hnf4a | 26763 | -0.472 | -0.0313 | No |
| 172 | Irf7 | 26817 | -0.483 | -0.0197 | No |
| 173 | Notch4 | 26900 | -0.495 | -0.0087 | No |
| 174 | F7 | 27032 | -0.521 | 0.0011 | No |
| 175 | Hspa1a | 27456 | -0.698 | 0.0053 | No |