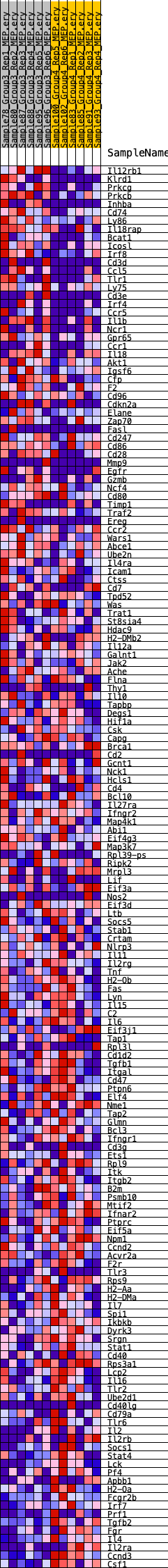
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | MEP.MEP.ery_Pheno.cls #Group3_versus_Group4.MEP.ery_Pheno.cls #Group3_versus_Group4_repos |
| Phenotype | MEP.ery_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
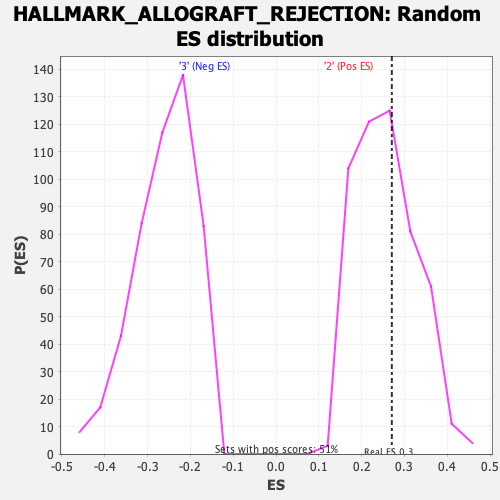
| GeneSet | HALLMARK_ALLOGRAFT_REJECTION |
| Enrichment Score (ES) | 0.27047488 |
| Normalized Enrichment Score (NES) | 1.0512054 |
| Nominal p-value | 0.39019608 |
| FDR q-value | 0.8724557 |
| FWER p-Value | 0.998 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Il12rb1 | 112 | 0.929 | 0.0192 | Yes |
| 2 | Klrd1 | 357 | 0.739 | 0.0287 | Yes |
| 3 | Prkcg | 429 | 0.706 | 0.0438 | Yes |
| 4 | Prkcb | 455 | 0.698 | 0.0603 | Yes |
| 5 | Inhba | 558 | 0.664 | 0.0732 | Yes |
| 6 | Cd74 | 934 | 0.581 | 0.0741 | Yes |
| 7 | Ly86 | 1110 | 0.550 | 0.0814 | Yes |
| 8 | Il18rap | 1132 | 0.546 | 0.0943 | Yes |
| 9 | Bcat1 | 1213 | 0.531 | 0.1047 | Yes |
| 10 | Icosl | 1277 | 0.520 | 0.1153 | Yes |
| 11 | Irf8 | 1418 | 0.505 | 0.1229 | Yes |
| 12 | Cd3d | 1506 | 0.496 | 0.1321 | Yes |
| 13 | Ccl5 | 1515 | 0.496 | 0.1442 | Yes |
| 14 | Tlr1 | 1633 | 0.483 | 0.1520 | Yes |
| 15 | Ly75 | 1710 | 0.472 | 0.1610 | Yes |
| 16 | Cd3e | 1797 | 0.468 | 0.1696 | Yes |
| 17 | Irf4 | 1893 | 0.459 | 0.1776 | Yes |
| 18 | Ccr5 | 1962 | 0.449 | 0.1864 | Yes |
| 19 | Il1b | 1965 | 0.449 | 0.1975 | Yes |
| 20 | Ncr1 | 2099 | 0.434 | 0.2035 | Yes |
| 21 | Gpr65 | 2300 | 0.423 | 0.2068 | Yes |
| 22 | Ccr1 | 2558 | 0.412 | 0.2077 | Yes |
| 23 | Il18 | 2694 | 0.403 | 0.2129 | Yes |
| 24 | Akt1 | 2997 | 0.381 | 0.2114 | Yes |
| 25 | Igsf6 | 3110 | 0.371 | 0.2166 | Yes |
| 26 | Cfp | 3314 | 0.355 | 0.2181 | Yes |
| 27 | F2 | 3653 | 0.335 | 0.2141 | Yes |
| 28 | Cd96 | 3660 | 0.334 | 0.2223 | Yes |
| 29 | Cdkn2a | 3692 | 0.332 | 0.2294 | Yes |
| 30 | Elane | 3799 | 0.325 | 0.2337 | Yes |
| 31 | Zap70 | 3898 | 0.318 | 0.2381 | Yes |
| 32 | Fasl | 4228 | 0.300 | 0.2336 | Yes |
| 33 | Cd247 | 4359 | 0.298 | 0.2363 | Yes |
| 34 | Cd86 | 4395 | 0.296 | 0.2424 | Yes |
| 35 | Cd28 | 4405 | 0.295 | 0.2495 | Yes |
| 36 | Mmp9 | 4454 | 0.292 | 0.2550 | Yes |
| 37 | Egfr | 4608 | 0.286 | 0.2566 | Yes |
| 38 | Gzmb | 4612 | 0.286 | 0.2636 | Yes |
| 39 | Ncf4 | 4621 | 0.286 | 0.2705 | Yes |
| 40 | Cd80 | 4934 | 0.271 | 0.2659 | No |
| 41 | Timp1 | 5673 | 0.240 | 0.2450 | No |
| 42 | Traf2 | 6263 | 0.221 | 0.2290 | No |
| 43 | Ereg | 6395 | 0.216 | 0.2296 | No |
| 44 | Ccr2 | 6690 | 0.212 | 0.2242 | No |
| 45 | Wars1 | 7074 | 0.196 | 0.2152 | No |
| 46 | Abce1 | 7133 | 0.194 | 0.2179 | No |
| 47 | Ube2n | 8105 | 0.159 | 0.1865 | No |
| 48 | Il4ra | 8280 | 0.152 | 0.1839 | No |
| 49 | Icam1 | 8360 | 0.149 | 0.1848 | No |
| 50 | Ctss | 8367 | 0.149 | 0.1883 | No |
| 51 | Cd7 | 8544 | 0.144 | 0.1855 | No |
| 52 | Tpd52 | 8767 | 0.137 | 0.1808 | No |
| 53 | Was | 9419 | 0.116 | 0.1600 | No |
| 54 | Trat1 | 10323 | 0.092 | 0.1294 | No |
| 55 | St8sia4 | 10572 | 0.086 | 0.1225 | No |
| 56 | Hdac9 | 10575 | 0.086 | 0.1245 | No |
| 57 | H2-DMb2 | 10768 | 0.081 | 0.1195 | No |
| 58 | Il12a | 10847 | 0.079 | 0.1187 | No |
| 59 | Galnt1 | 11011 | 0.075 | 0.1146 | No |
| 60 | Jak2 | 11062 | 0.073 | 0.1146 | No |
| 61 | Ache | 11076 | 0.073 | 0.1159 | No |
| 62 | Flna | 11118 | 0.072 | 0.1162 | No |
| 63 | Thy1 | 11292 | 0.068 | 0.1116 | No |
| 64 | Il10 | 11306 | 0.067 | 0.1128 | No |
| 65 | Tapbp | 11414 | 0.064 | 0.1105 | No |
| 66 | Degs1 | 11438 | 0.063 | 0.1113 | No |
| 67 | Hif1a | 11809 | 0.053 | 0.0991 | No |
| 68 | Csk | 11823 | 0.053 | 0.1000 | No |
| 69 | Capg | 11829 | 0.053 | 0.1011 | No |
| 70 | Brca1 | 11833 | 0.053 | 0.1023 | No |
| 71 | Cd2 | 11919 | 0.050 | 0.1005 | No |
| 72 | Gcnt1 | 12193 | 0.043 | 0.0916 | No |
| 73 | Nck1 | 12480 | 0.036 | 0.0821 | No |
| 74 | Hcls1 | 12698 | 0.030 | 0.0749 | No |
| 75 | Cd4 | 12862 | 0.026 | 0.0696 | No |
| 76 | Bcl10 | 13078 | 0.021 | 0.0623 | No |
| 77 | Il27ra | 13081 | 0.021 | 0.0627 | No |
| 78 | Ifngr2 | 13173 | 0.019 | 0.0599 | No |
| 79 | Map4k1 | 13239 | 0.017 | 0.0579 | No |
| 80 | Abi1 | 13990 | 0.002 | 0.0307 | No |
| 81 | Eif4g3 | 14003 | 0.002 | 0.0303 | No |
| 82 | Map3k7 | 14942 | -0.003 | -0.0038 | No |
| 83 | Rpl39-ps | 15012 | -0.004 | -0.0062 | No |
| 84 | Ripk2 | 15066 | -0.006 | -0.0080 | No |
| 85 | Mrpl3 | 15100 | -0.007 | -0.0090 | No |
| 86 | Lif | 15156 | -0.008 | -0.0109 | No |
| 87 | Eif3a | 15245 | -0.010 | -0.0138 | No |
| 88 | Nos2 | 15284 | -0.011 | -0.0149 | No |
| 89 | Eif3d | 15577 | -0.016 | -0.0251 | No |
| 90 | Ltb | 15624 | -0.017 | -0.0264 | No |
| 91 | Socs5 | 15832 | -0.023 | -0.0334 | No |
| 92 | Stab1 | 16048 | -0.028 | -0.0405 | No |
| 93 | Crtam | 16051 | -0.028 | -0.0399 | No |
| 94 | Nlrp3 | 16073 | -0.029 | -0.0399 | No |
| 95 | Il11 | 16090 | -0.029 | -0.0397 | No |
| 96 | Il2rg | 16387 | -0.037 | -0.0496 | No |
| 97 | Tnf | 16482 | -0.039 | -0.0520 | No |
| 98 | H2-Ob | 16666 | -0.044 | -0.0576 | No |
| 99 | Fas | 16681 | -0.045 | -0.0570 | No |
| 100 | Lyn | 17244 | -0.059 | -0.0760 | No |
| 101 | Il15 | 17404 | -0.064 | -0.0802 | No |
| 102 | C2 | 17452 | -0.065 | -0.0803 | No |
| 103 | Il6 | 17789 | -0.072 | -0.0908 | No |
| 104 | Eif3j1 | 17799 | -0.072 | -0.0893 | No |
| 105 | Tap1 | 17898 | -0.074 | -0.0910 | No |
| 106 | Rpl3l | 18127 | -0.080 | -0.0973 | No |
| 107 | Cd1d2 | 18144 | -0.081 | -0.0959 | No |
| 108 | Tgfb1 | 18170 | -0.082 | -0.0947 | No |
| 109 | Itgal | 18512 | -0.090 | -0.1049 | No |
| 110 | Cd47 | 18591 | -0.092 | -0.1055 | No |
| 111 | Ptpn6 | 18908 | -0.100 | -0.1145 | No |
| 112 | Elf4 | 19396 | -0.115 | -0.1294 | No |
| 113 | Nme1 | 19513 | -0.117 | -0.1307 | No |
| 114 | Tap2 | 19562 | -0.119 | -0.1294 | No |
| 115 | Glmn | 19593 | -0.120 | -0.1275 | No |
| 116 | Bcl3 | 19598 | -0.120 | -0.1247 | No |
| 117 | Ifngr1 | 19612 | -0.121 | -0.1221 | No |
| 118 | Cd3g | 19636 | -0.121 | -0.1199 | No |
| 119 | Ets1 | 19655 | -0.122 | -0.1176 | No |
| 120 | Rpl9 | 19930 | -0.129 | -0.1243 | No |
| 121 | Itk | 20024 | -0.133 | -0.1244 | No |
| 122 | Itgb2 | 20093 | -0.135 | -0.1235 | No |
| 123 | B2m | 20334 | -0.143 | -0.1286 | No |
| 124 | Psmb10 | 20530 | -0.149 | -0.1320 | No |
| 125 | Mtif2 | 20556 | -0.150 | -0.1292 | No |
| 126 | Ifnar2 | 20870 | -0.159 | -0.1366 | No |
| 127 | Ptprc | 20884 | -0.160 | -0.1331 | No |
| 128 | Eif5a | 20999 | -0.164 | -0.1332 | No |
| 129 | Npm1 | 21414 | -0.178 | -0.1438 | No |
| 130 | Ccnd2 | 21480 | -0.181 | -0.1416 | No |
| 131 | Acvr2a | 21742 | -0.191 | -0.1464 | No |
| 132 | F2r | 21855 | -0.195 | -0.1456 | No |
| 133 | Tlr3 | 22285 | -0.210 | -0.1560 | No |
| 134 | Rps9 | 22403 | -0.215 | -0.1548 | No |
| 135 | H2-Aa | 22459 | -0.217 | -0.1514 | No |
| 136 | H2-DMa | 22485 | -0.218 | -0.1469 | No |
| 137 | Il7 | 22517 | -0.219 | -0.1426 | No |
| 138 | Spi1 | 22849 | -0.224 | -0.1490 | No |
| 139 | Ikbkb | 23295 | -0.244 | -0.1592 | No |
| 140 | Dyrk3 | 23623 | -0.258 | -0.1646 | No |
| 141 | Srgn | 23947 | -0.268 | -0.1697 | No |
| 142 | Stat1 | 23990 | -0.270 | -0.1645 | No |
| 143 | Cd40 | 24114 | -0.276 | -0.1621 | No |
| 144 | Rps3a1 | 24251 | -0.284 | -0.1599 | No |
| 145 | Lcp2 | 24389 | -0.290 | -0.1577 | No |
| 146 | Il16 | 24441 | -0.292 | -0.1522 | No |
| 147 | Tlr2 | 24525 | -0.297 | -0.1478 | No |
| 148 | Ube2d1 | 24654 | -0.304 | -0.1449 | No |
| 149 | Cd40lg | 24841 | -0.314 | -0.1438 | No |
| 150 | Cd79a | 25169 | -0.330 | -0.1475 | No |
| 151 | Tlr6 | 25277 | -0.336 | -0.1430 | No |
| 152 | Il2 | 25433 | -0.349 | -0.1399 | No |
| 153 | Il2rb | 26024 | -0.397 | -0.1515 | No |
| 154 | Socs1 | 26435 | -0.433 | -0.1556 | No |
| 155 | Stat4 | 26502 | -0.441 | -0.1470 | No |
| 156 | Lck | 26580 | -0.449 | -0.1385 | No |
| 157 | Pf4 | 26623 | -0.454 | -0.1287 | No |
| 158 | Apbb1 | 26715 | -0.466 | -0.1204 | No |
| 159 | H2-Oa | 26719 | -0.466 | -0.1089 | No |
| 160 | Fcgr2b | 26759 | -0.472 | -0.0985 | No |
| 161 | Irf7 | 26817 | -0.483 | -0.0885 | No |
| 162 | Prf1 | 27048 | -0.525 | -0.0838 | No |
| 163 | Tgfb2 | 27246 | -0.578 | -0.0765 | No |
| 164 | Fgr | 27298 | -0.596 | -0.0634 | No |
| 165 | Il4 | 27384 | -0.645 | -0.0504 | No |
| 166 | Il2ra | 27486 | -0.732 | -0.0358 | No |
| 167 | Ccnd3 | 27529 | -0.794 | -0.0175 | No |
| 168 | Csf1 | 27536 | -0.805 | 0.0024 | No |