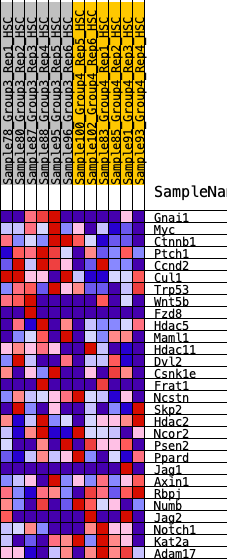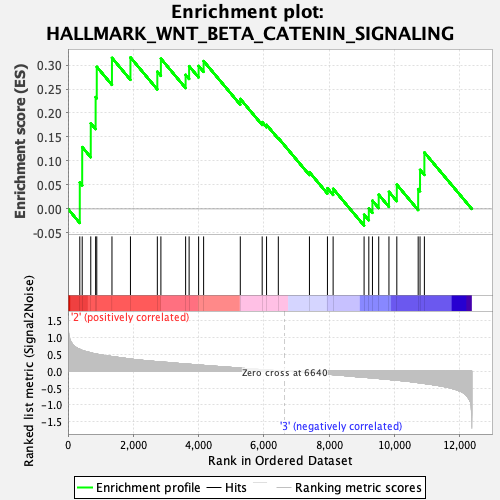
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | HSC.HSC_Pheno.cls#Group3_versus_Group4.HSC_Pheno.cls#Group3_versus_Group4_repos |
| Phenotype | HSC_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_WNT_BETA_CATENIN_SIGNALING |
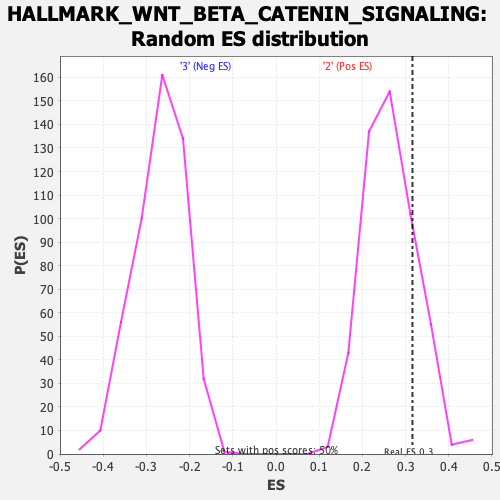
| Enrichment Score (ES) | 0.31587094 |
| Normalized Enrichment Score (NES) | 1.1946342 |
| Nominal p-value | 0.20039682 |
| FDR q-value | 0.39549696 |
| FWER p-Value | 0.942 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Gnai1 | 361 | 0.650 | 0.0550 | Yes |
| 2 | Myc | 435 | 0.615 | 0.1289 | Yes |
| 3 | Ctnnb1 | 697 | 0.545 | 0.1784 | Yes |
| 4 | Ptch1 | 846 | 0.514 | 0.2332 | Yes |
| 5 | Ccnd2 | 881 | 0.508 | 0.2963 | Yes |
| 6 | Cul1 | 1346 | 0.434 | 0.3150 | Yes |
| 7 | Trp53 | 1912 | 0.360 | 0.3159 | Yes |
| 8 | Wnt5b | 2735 | 0.285 | 0.2862 | No |
| 9 | Fzd8 | 2844 | 0.276 | 0.3133 | No |
| 10 | Hdac5 | 3601 | 0.212 | 0.2795 | No |
| 11 | Maml1 | 3708 | 0.206 | 0.2977 | No |
| 12 | Hdac11 | 3999 | 0.185 | 0.2981 | No |
| 13 | Dvl2 | 4152 | 0.173 | 0.3083 | No |
| 14 | Csnk1e | 5275 | 0.093 | 0.2293 | No |
| 15 | Frat1 | 5945 | 0.046 | 0.1810 | No |
| 16 | Ncstn | 6079 | 0.039 | 0.1753 | No |
| 17 | Skp2 | 6439 | 0.013 | 0.1478 | No |
| 18 | Hdac2 | 7392 | -0.046 | 0.0766 | No |
| 19 | Ncor2 | 7943 | -0.086 | 0.0432 | No |
| 20 | Psen2 | 8117 | -0.099 | 0.0420 | No |
| 21 | Ppard | 9065 | -0.176 | -0.0119 | No |
| 22 | Jag1 | 9210 | -0.188 | 0.0008 | No |
| 23 | Axin1 | 9321 | -0.196 | 0.0173 | No |
| 24 | Rbpj | 9513 | -0.215 | 0.0297 | No |
| 25 | Numb | 9826 | -0.241 | 0.0357 | No |
| 26 | Jag2 | 10067 | -0.265 | 0.0506 | No |
| 27 | Notch1 | 10721 | -0.337 | 0.0413 | No |
| 28 | Kat2a | 10778 | -0.345 | 0.0815 | No |
| 29 | Adam17 | 10911 | -0.361 | 0.1177 | No |