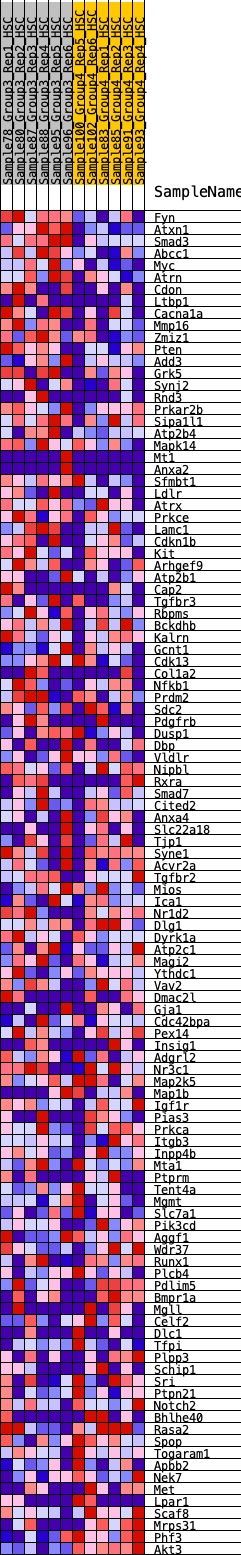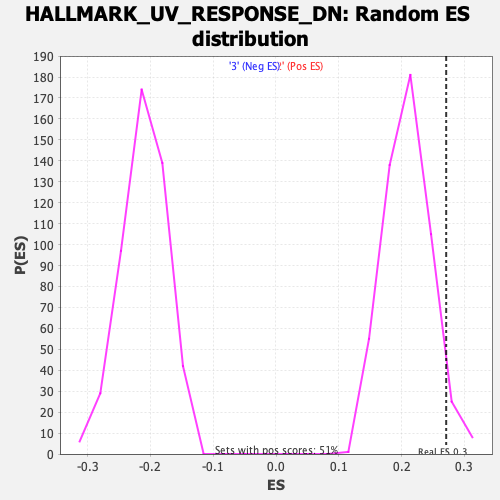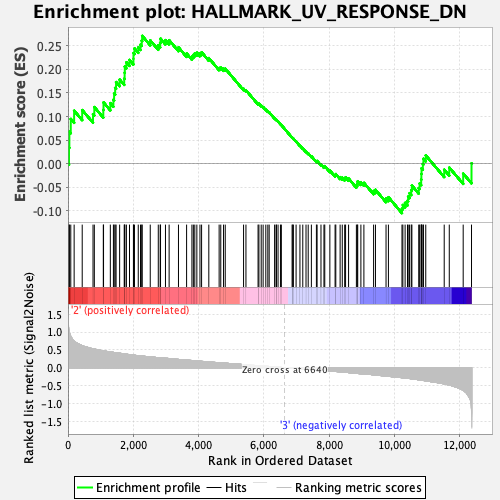
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | HSC.HSC_Pheno.cls#Group3_versus_Group4.HSC_Pheno.cls#Group3_versus_Group4_repos |
| Phenotype | HSC_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | 0.2711889 |
| Normalized Enrichment Score (NES) | 1.2880703 |
| Nominal p-value | 0.04288499 |
| FDR q-value | 0.41930905 |
| FWER p-Value | 0.794 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Fyn | 31 | 1.058 | 0.0343 | Yes |
| 2 | Atxn1 | 42 | 0.984 | 0.0677 | Yes |
| 3 | Smad3 | 85 | 0.879 | 0.0949 | Yes |
| 4 | Abcc1 | 188 | 0.743 | 0.1124 | Yes |
| 5 | Myc | 435 | 0.615 | 0.1137 | Yes |
| 6 | Atrn | 766 | 0.528 | 0.1052 | Yes |
| 7 | Cdon | 806 | 0.521 | 0.1201 | Yes |
| 8 | Ltbp1 | 1081 | 0.471 | 0.1141 | Yes |
| 9 | Cacna1a | 1088 | 0.470 | 0.1300 | Yes |
| 10 | Mmp16 | 1296 | 0.441 | 0.1284 | Yes |
| 11 | Zmiz1 | 1390 | 0.429 | 0.1358 | Yes |
| 12 | Pten | 1412 | 0.425 | 0.1489 | Yes |
| 13 | Add3 | 1444 | 0.418 | 0.1609 | Yes |
| 14 | Grk5 | 1472 | 0.416 | 0.1732 | Yes |
| 15 | Synj2 | 1581 | 0.403 | 0.1784 | Yes |
| 16 | Rnd3 | 1721 | 0.386 | 0.1805 | Yes |
| 17 | Prkar2b | 1734 | 0.384 | 0.1929 | Yes |
| 18 | Sipa1l1 | 1739 | 0.384 | 0.2059 | Yes |
| 19 | Atp2b4 | 1788 | 0.377 | 0.2151 | Yes |
| 20 | Mapk14 | 1883 | 0.365 | 0.2201 | Yes |
| 21 | Mt1 | 2003 | 0.351 | 0.2226 | Yes |
| 22 | Anxa2 | 2005 | 0.351 | 0.2347 | Yes |
| 23 | Sfmbt1 | 2035 | 0.348 | 0.2445 | Yes |
| 24 | Ldlr | 2151 | 0.336 | 0.2468 | Yes |
| 25 | Atrx | 2220 | 0.328 | 0.2526 | Yes |
| 26 | Prkce | 2257 | 0.325 | 0.2610 | Yes |
| 27 | Lamc1 | 2271 | 0.324 | 0.2712 | Yes |
| 28 | Cdkn1b | 2518 | 0.301 | 0.2616 | No |
| 29 | Kit | 2764 | 0.282 | 0.2514 | No |
| 30 | Arhgef9 | 2822 | 0.277 | 0.2564 | No |
| 31 | Atp2b1 | 2833 | 0.277 | 0.2652 | No |
| 32 | Cap2 | 2984 | 0.263 | 0.2621 | No |
| 33 | Tgfbr3 | 3096 | 0.256 | 0.2620 | No |
| 34 | Rbpms | 3382 | 0.231 | 0.2468 | No |
| 35 | Bckdhb | 3631 | 0.209 | 0.2338 | No |
| 36 | Kalrn | 3794 | 0.200 | 0.2275 | No |
| 37 | Gcnt1 | 3838 | 0.196 | 0.2309 | No |
| 38 | Cdk13 | 3882 | 0.193 | 0.2341 | No |
| 39 | Col1a2 | 3945 | 0.189 | 0.2356 | No |
| 40 | Nfkb1 | 4040 | 0.182 | 0.2342 | No |
| 41 | Prdm2 | 4090 | 0.177 | 0.2364 | No |
| 42 | Sdc2 | 4311 | 0.161 | 0.2240 | No |
| 43 | Pdgfrb | 4627 | 0.138 | 0.2031 | No |
| 44 | Dusp1 | 4677 | 0.134 | 0.2038 | No |
| 45 | Dbp | 4762 | 0.129 | 0.2014 | No |
| 46 | Vldlr | 4814 | 0.125 | 0.2016 | No |
| 47 | Nipbl | 5375 | 0.085 | 0.1589 | No |
| 48 | Rxra | 5448 | 0.081 | 0.1558 | No |
| 49 | Smad7 | 5819 | 0.054 | 0.1275 | No |
| 50 | Cited2 | 5845 | 0.052 | 0.1272 | No |
| 51 | Anxa4 | 5915 | 0.047 | 0.1232 | No |
| 52 | Slc22a18 | 5974 | 0.044 | 0.1200 | No |
| 53 | Tjp1 | 6056 | 0.040 | 0.1148 | No |
| 54 | Syne1 | 6118 | 0.036 | 0.1111 | No |
| 55 | Acvr2a | 6159 | 0.034 | 0.1090 | No |
| 56 | Tgfbr2 | 6325 | 0.021 | 0.0962 | No |
| 57 | Mios | 6370 | 0.018 | 0.0933 | No |
| 58 | Ica1 | 6382 | 0.017 | 0.0930 | No |
| 59 | Nr1d2 | 6431 | 0.013 | 0.0895 | No |
| 60 | Dlg1 | 6505 | 0.008 | 0.0838 | No |
| 61 | Dyrk1a | 6532 | 0.007 | 0.0819 | No |
| 62 | Atp2c1 | 6861 | -0.008 | 0.0554 | No |
| 63 | Magi2 | 6880 | -0.010 | 0.0543 | No |
| 64 | Ythdc1 | 6900 | -0.011 | 0.0532 | No |
| 65 | Vav2 | 6983 | -0.017 | 0.0470 | No |
| 66 | Dmac2l | 7101 | -0.025 | 0.0384 | No |
| 67 | Gja1 | 7188 | -0.029 | 0.0324 | No |
| 68 | Cdc42bpa | 7287 | -0.038 | 0.0257 | No |
| 69 | Pex14 | 7354 | -0.042 | 0.0218 | No |
| 70 | Insig1 | 7451 | -0.050 | 0.0157 | No |
| 71 | Adgrl2 | 7605 | -0.060 | 0.0053 | No |
| 72 | Nr3c1 | 7626 | -0.061 | 0.0058 | No |
| 73 | Map2k5 | 7745 | -0.071 | -0.0014 | No |
| 74 | Map1b | 7836 | -0.077 | -0.0061 | No |
| 75 | Igf1r | 7861 | -0.079 | -0.0053 | No |
| 76 | Pias3 | 8017 | -0.092 | -0.0147 | No |
| 77 | Prkca | 8177 | -0.105 | -0.0240 | No |
| 78 | Itgb3 | 8199 | -0.106 | -0.0221 | No |
| 79 | Inpp4b | 8331 | -0.116 | -0.0287 | No |
| 80 | Mta1 | 8397 | -0.122 | -0.0297 | No |
| 81 | Ptprm | 8471 | -0.128 | -0.0313 | No |
| 82 | Tent4a | 8498 | -0.129 | -0.0289 | No |
| 83 | Mgmt | 8590 | -0.137 | -0.0316 | No |
| 84 | Slc7a1 | 8829 | -0.159 | -0.0454 | No |
| 85 | Pik3cd | 8856 | -0.160 | -0.0420 | No |
| 86 | Aggf1 | 8870 | -0.162 | -0.0374 | No |
| 87 | Wdr37 | 8967 | -0.169 | -0.0394 | No |
| 88 | Runx1 | 9059 | -0.175 | -0.0407 | No |
| 89 | Plcb4 | 9356 | -0.198 | -0.0580 | No |
| 90 | Pdlim5 | 9413 | -0.204 | -0.0554 | No |
| 91 | Bmpr1a | 9737 | -0.233 | -0.0737 | No |
| 92 | Mgll | 9810 | -0.240 | -0.0712 | No |
| 93 | Celf2 | 10225 | -0.279 | -0.0953 | No |
| 94 | Dlc1 | 10252 | -0.283 | -0.0876 | No |
| 95 | Tfpi | 10321 | -0.289 | -0.0830 | No |
| 96 | Plpp3 | 10394 | -0.296 | -0.0786 | No |
| 97 | Schip1 | 10414 | -0.298 | -0.0698 | No |
| 98 | Sri | 10457 | -0.303 | -0.0627 | No |
| 99 | Ptpn21 | 10509 | -0.312 | -0.0559 | No |
| 100 | Notch2 | 10528 | -0.314 | -0.0465 | No |
| 101 | Bhlhe40 | 10738 | -0.339 | -0.0518 | No |
| 102 | Rasa2 | 10766 | -0.343 | -0.0421 | No |
| 103 | Spop | 10813 | -0.351 | -0.0336 | No |
| 104 | Togaram1 | 10819 | -0.351 | -0.0218 | No |
| 105 | Apbb2 | 10824 | -0.352 | -0.0099 | No |
| 106 | Nek7 | 10864 | -0.356 | -0.0007 | No |
| 107 | Met | 10882 | -0.359 | 0.0104 | No |
| 108 | Lpar1 | 10953 | -0.369 | 0.0175 | No |
| 109 | Scaf8 | 11518 | -0.454 | -0.0127 | No |
| 110 | Mrps31 | 11673 | -0.486 | -0.0083 | No |
| 111 | Phf3 | 12099 | -0.634 | -0.0210 | No |
| 112 | Akt3 | 12354 | -1.220 | 0.0007 | No |