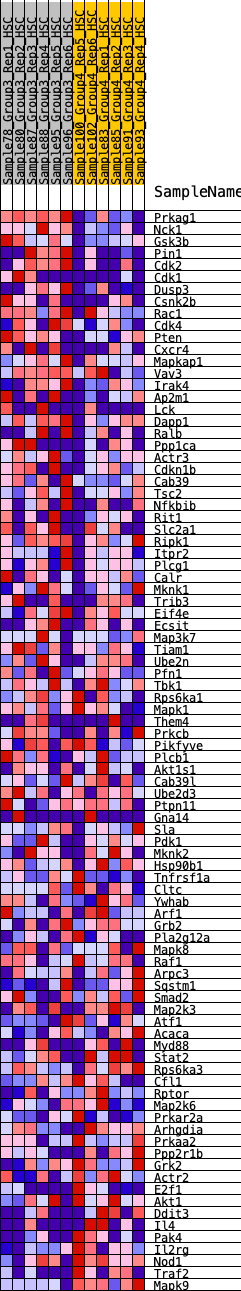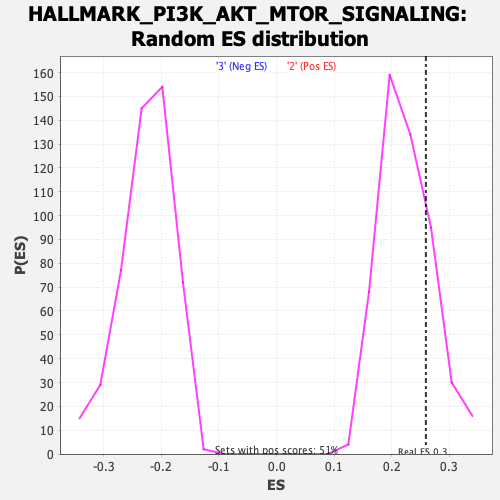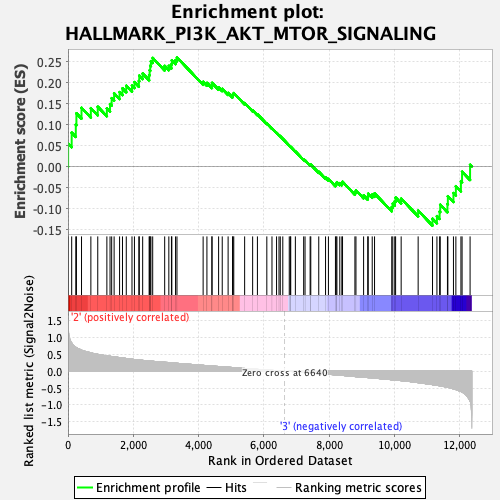
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | HSC.HSC_Pheno.cls#Group3_versus_Group4.HSC_Pheno.cls#Group3_versus_Group4_repos |
| Phenotype | HSC_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_PI3K_AKT_MTOR_SIGNALING |
| Enrichment Score (ES) | 0.2597325 |
| Normalized Enrichment Score (NES) | 1.1518139 |
| Nominal p-value | 0.2193676 |
| FDR q-value | 0.46592915 |
| FWER p-Value | 0.962 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Prkag1 | 4 | 1.343 | 0.0548 | Yes |
| 2 | Nck1 | 113 | 0.844 | 0.0807 | Yes |
| 3 | Gsk3b | 239 | 0.708 | 0.0996 | Yes |
| 4 | Pin1 | 258 | 0.698 | 0.1268 | Yes |
| 5 | Cdk2 | 414 | 0.625 | 0.1398 | Yes |
| 6 | Cdk1 | 700 | 0.544 | 0.1390 | Yes |
| 7 | Dusp3 | 911 | 0.500 | 0.1424 | Yes |
| 8 | Csnk2b | 1190 | 0.456 | 0.1385 | Yes |
| 9 | Rac1 | 1287 | 0.443 | 0.1488 | Yes |
| 10 | Cdk4 | 1337 | 0.436 | 0.1627 | Yes |
| 11 | Pten | 1412 | 0.425 | 0.1742 | Yes |
| 12 | Cxcr4 | 1579 | 0.404 | 0.1772 | Yes |
| 13 | Mapkap1 | 1668 | 0.394 | 0.1862 | Yes |
| 14 | Vav3 | 1784 | 0.377 | 0.1923 | Yes |
| 15 | Irak4 | 1959 | 0.356 | 0.1928 | Yes |
| 16 | Ap2m1 | 2034 | 0.348 | 0.2011 | Yes |
| 17 | Lck | 2170 | 0.334 | 0.2038 | Yes |
| 18 | Dapp1 | 2182 | 0.333 | 0.2166 | Yes |
| 19 | Ralb | 2286 | 0.322 | 0.2214 | Yes |
| 20 | Ppp1ca | 2484 | 0.305 | 0.2178 | Yes |
| 21 | Actr3 | 2499 | 0.303 | 0.2291 | Yes |
| 22 | Cdkn1b | 2518 | 0.301 | 0.2401 | Yes |
| 23 | Cab39 | 2538 | 0.299 | 0.2508 | Yes |
| 24 | Tsc2 | 2593 | 0.294 | 0.2585 | Yes |
| 25 | Nfkbib | 2960 | 0.265 | 0.2395 | Yes |
| 26 | Rit1 | 3087 | 0.257 | 0.2398 | Yes |
| 27 | Slc2a1 | 3167 | 0.250 | 0.2436 | Yes |
| 28 | Ripk1 | 3179 | 0.249 | 0.2530 | Yes |
| 29 | Itpr2 | 3293 | 0.238 | 0.2535 | Yes |
| 30 | Plcg1 | 3336 | 0.234 | 0.2597 | Yes |
| 31 | Calr | 4137 | 0.174 | 0.2017 | No |
| 32 | Mknk1 | 4253 | 0.165 | 0.1991 | No |
| 33 | Trib3 | 4407 | 0.153 | 0.1930 | No |
| 34 | Eif4e | 4408 | 0.153 | 0.1993 | No |
| 35 | Ecsit | 4611 | 0.139 | 0.1885 | No |
| 36 | Map3k7 | 4722 | 0.132 | 0.1850 | No |
| 37 | Tiam1 | 4902 | 0.120 | 0.1753 | No |
| 38 | Ube2n | 5037 | 0.110 | 0.1689 | No |
| 39 | Pfn1 | 5050 | 0.109 | 0.1724 | No |
| 40 | Tbk1 | 5075 | 0.108 | 0.1749 | No |
| 41 | Rps6ka1 | 5408 | 0.083 | 0.1513 | No |
| 42 | Mapk1 | 5653 | 0.065 | 0.1341 | No |
| 43 | Them4 | 5799 | 0.055 | 0.1245 | No |
| 44 | Prkcb | 6087 | 0.038 | 0.1027 | No |
| 45 | Pikfyve | 6245 | 0.027 | 0.0910 | No |
| 46 | Plcb1 | 6385 | 0.016 | 0.0804 | No |
| 47 | Akt1s1 | 6460 | 0.011 | 0.0748 | No |
| 48 | Cab39l | 6499 | 0.008 | 0.0721 | No |
| 49 | Ube2d3 | 6576 | 0.005 | 0.0661 | No |
| 50 | Ptpn11 | 6774 | -0.002 | 0.0501 | No |
| 51 | Gna14 | 6807 | -0.005 | 0.0477 | No |
| 52 | Sla | 6819 | -0.006 | 0.0470 | No |
| 53 | Pdk1 | 6961 | -0.015 | 0.0361 | No |
| 54 | Mknk2 | 7215 | -0.032 | 0.0168 | No |
| 55 | Hsp90b1 | 7257 | -0.036 | 0.0150 | No |
| 56 | Tnfrsf1a | 7413 | -0.047 | 0.0043 | No |
| 57 | Cltc | 7433 | -0.049 | 0.0048 | No |
| 58 | Ywhab | 7677 | -0.066 | -0.0123 | No |
| 59 | Arf1 | 7885 | -0.081 | -0.0259 | No |
| 60 | Grb2 | 7973 | -0.089 | -0.0293 | No |
| 61 | Pla2g12a | 8188 | -0.106 | -0.0424 | No |
| 62 | Mapk8 | 8210 | -0.107 | -0.0397 | No |
| 63 | Raf1 | 8233 | -0.108 | -0.0371 | No |
| 64 | Arpc3 | 8321 | -0.116 | -0.0394 | No |
| 65 | Sqstm1 | 8384 | -0.121 | -0.0395 | No |
| 66 | Smad2 | 8405 | -0.122 | -0.0361 | No |
| 67 | Map2k3 | 8784 | -0.156 | -0.0605 | No |
| 68 | Atf1 | 8816 | -0.158 | -0.0565 | No |
| 69 | Acaca | 9050 | -0.174 | -0.0683 | No |
| 70 | Myd88 | 9177 | -0.187 | -0.0709 | No |
| 71 | Stat2 | 9191 | -0.188 | -0.0643 | No |
| 72 | Rps6ka3 | 9314 | -0.195 | -0.0662 | No |
| 73 | Cfl1 | 9385 | -0.201 | -0.0637 | No |
| 74 | Rptor | 9912 | -0.251 | -0.0962 | No |
| 75 | Map2k6 | 9944 | -0.255 | -0.0883 | No |
| 76 | Prkar2a | 10004 | -0.259 | -0.0824 | No |
| 77 | Arhgdia | 10030 | -0.261 | -0.0737 | No |
| 78 | Prkaa2 | 10200 | -0.277 | -0.0761 | No |
| 79 | Ppp2r1b | 10720 | -0.337 | -0.1046 | No |
| 80 | Grk2 | 11160 | -0.396 | -0.1241 | No |
| 81 | Actr2 | 11295 | -0.417 | -0.1179 | No |
| 82 | E2f1 | 11378 | -0.429 | -0.1069 | No |
| 83 | Akt1 | 11396 | -0.430 | -0.0907 | No |
| 84 | Ddit3 | 11617 | -0.474 | -0.0891 | No |
| 85 | Il4 | 11630 | -0.478 | -0.0705 | No |
| 86 | Pak4 | 11801 | -0.521 | -0.0629 | No |
| 87 | Il2rg | 11875 | -0.542 | -0.0466 | No |
| 88 | Nod1 | 12030 | -0.597 | -0.0347 | No |
| 89 | Traf2 | 12065 | -0.617 | -0.0121 | No |
| 90 | Mapk9 | 12310 | -0.883 | 0.0043 | No |