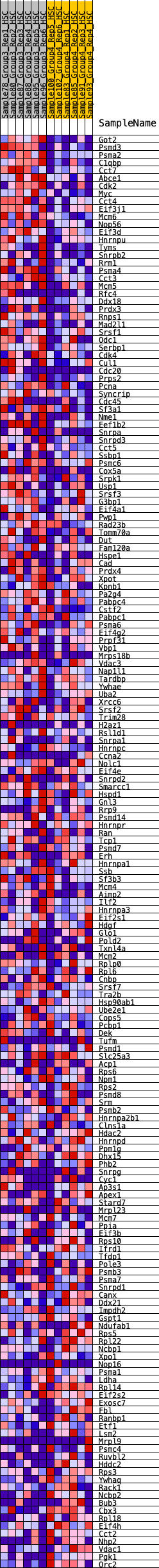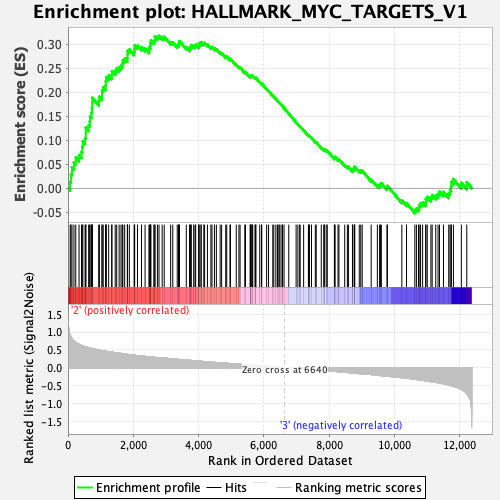
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | HSC.HSC_Pheno.cls#Group3_versus_Group4.HSC_Pheno.cls#Group3_versus_Group4_repos |
| Phenotype | HSC_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
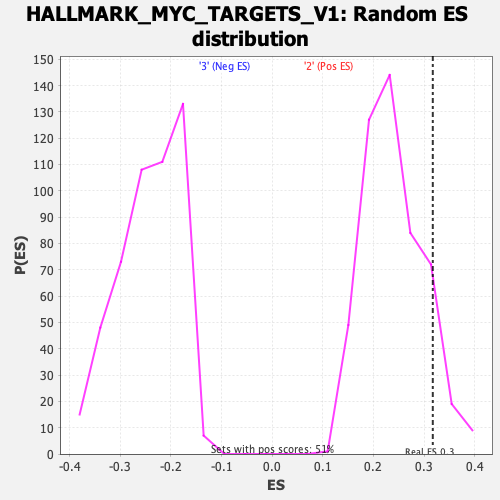
| GeneSet | HALLMARK_MYC_TARGETS_V1 |
| Enrichment Score (ES) | 0.31727716 |
| Normalized Enrichment Score (NES) | 1.3238132 |
| Nominal p-value | 0.12673268 |
| FDR q-value | 0.37919244 |
| FWER p-Value | 0.726 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Got2 | 66 | 0.922 | 0.0134 | Yes |
| 2 | Psmd3 | 91 | 0.867 | 0.0291 | Yes |
| 3 | Psma2 | 122 | 0.816 | 0.0434 | Yes |
| 4 | C1qbp | 182 | 0.747 | 0.0538 | Yes |
| 5 | Cct7 | 237 | 0.710 | 0.0638 | Yes |
| 6 | Abce1 | 338 | 0.658 | 0.0691 | Yes |
| 7 | Cdk2 | 414 | 0.625 | 0.0757 | Yes |
| 8 | Myc | 435 | 0.615 | 0.0866 | Yes |
| 9 | Cct4 | 455 | 0.608 | 0.0974 | Yes |
| 10 | Eif3j1 | 523 | 0.590 | 0.1040 | Yes |
| 11 | Mcm6 | 544 | 0.582 | 0.1142 | Yes |
| 12 | Nop56 | 545 | 0.581 | 0.1261 | Yes |
| 13 | Eif3d | 632 | 0.561 | 0.1305 | Yes |
| 14 | Hnrnpu | 665 | 0.554 | 0.1392 | Yes |
| 15 | Tyms | 675 | 0.552 | 0.1497 | Yes |
| 16 | Snrpb2 | 717 | 0.539 | 0.1574 | Yes |
| 17 | Rrm1 | 733 | 0.537 | 0.1671 | Yes |
| 18 | Psma4 | 743 | 0.535 | 0.1773 | Yes |
| 19 | Cct3 | 744 | 0.535 | 0.1882 | Yes |
| 20 | Mcm5 | 940 | 0.495 | 0.1823 | Yes |
| 21 | Rfc4 | 961 | 0.492 | 0.1907 | Yes |
| 22 | Ddx18 | 1041 | 0.477 | 0.1940 | Yes |
| 23 | Prdx3 | 1042 | 0.477 | 0.2037 | Yes |
| 24 | Rnps1 | 1078 | 0.472 | 0.2105 | Yes |
| 25 | Mad2l1 | 1152 | 0.461 | 0.2139 | Yes |
| 26 | Srsf1 | 1157 | 0.460 | 0.2230 | Yes |
| 27 | Odc1 | 1173 | 0.458 | 0.2311 | Yes |
| 28 | Serbp1 | 1245 | 0.447 | 0.2344 | Yes |
| 29 | Cdk4 | 1337 | 0.436 | 0.2359 | Yes |
| 30 | Cul1 | 1346 | 0.434 | 0.2441 | Yes |
| 31 | Cdc20 | 1445 | 0.418 | 0.2446 | Yes |
| 32 | Prps2 | 1491 | 0.416 | 0.2494 | Yes |
| 33 | Pcna | 1566 | 0.406 | 0.2516 | Yes |
| 34 | Syncrip | 1623 | 0.400 | 0.2551 | Yes |
| 35 | Cdc45 | 1663 | 0.394 | 0.2600 | Yes |
| 36 | Sf3a1 | 1680 | 0.391 | 0.2667 | Yes |
| 37 | Nme1 | 1735 | 0.384 | 0.2701 | Yes |
| 38 | Eef1b2 | 1817 | 0.374 | 0.2711 | Yes |
| 39 | Snrpa | 1821 | 0.374 | 0.2785 | Yes |
| 40 | Snrpd3 | 1823 | 0.374 | 0.2860 | Yes |
| 41 | Cct5 | 1878 | 0.366 | 0.2890 | Yes |
| 42 | Ssbp1 | 2024 | 0.349 | 0.2843 | Yes |
| 43 | Psmc6 | 2036 | 0.348 | 0.2905 | Yes |
| 44 | Cox5a | 2041 | 0.347 | 0.2972 | Yes |
| 45 | Srpk1 | 2127 | 0.338 | 0.2971 | Yes |
| 46 | Usp1 | 2250 | 0.326 | 0.2938 | Yes |
| 47 | Srsf3 | 2360 | 0.316 | 0.2913 | Yes |
| 48 | G3bp1 | 2482 | 0.305 | 0.2876 | Yes |
| 49 | Eif4a1 | 2489 | 0.304 | 0.2933 | Yes |
| 50 | Pwp1 | 2521 | 0.301 | 0.2969 | Yes |
| 51 | Rad23b | 2524 | 0.300 | 0.3029 | Yes |
| 52 | Tomm70a | 2539 | 0.299 | 0.3078 | Yes |
| 53 | Dut | 2627 | 0.292 | 0.3066 | Yes |
| 54 | Fam120a | 2660 | 0.289 | 0.3099 | Yes |
| 55 | Hspe1 | 2661 | 0.289 | 0.3158 | Yes |
| 56 | Cad | 2738 | 0.285 | 0.3154 | Yes |
| 57 | Prdx4 | 2786 | 0.281 | 0.3173 | Yes |
| 58 | Xpot | 2883 | 0.273 | 0.3150 | No |
| 59 | Kpnb1 | 2946 | 0.266 | 0.3153 | No |
| 60 | Pa2g4 | 3143 | 0.251 | 0.3044 | No |
| 61 | Pabpc4 | 3208 | 0.246 | 0.3041 | No |
| 62 | Cstf2 | 3344 | 0.234 | 0.2978 | No |
| 63 | Pabpc1 | 3384 | 0.231 | 0.2993 | No |
| 64 | Psma6 | 3395 | 0.230 | 0.3032 | No |
| 65 | Eif4g2 | 3414 | 0.228 | 0.3064 | No |
| 66 | Prpf31 | 3621 | 0.210 | 0.2938 | No |
| 67 | Vbp1 | 3722 | 0.205 | 0.2897 | No |
| 68 | Mrps18b | 3741 | 0.204 | 0.2924 | No |
| 69 | Vdac3 | 3758 | 0.203 | 0.2953 | No |
| 70 | Nap1l1 | 3774 | 0.202 | 0.2982 | No |
| 71 | Tardbp | 3848 | 0.195 | 0.2961 | No |
| 72 | Ywhae | 3889 | 0.192 | 0.2968 | No |
| 73 | Uba2 | 3909 | 0.190 | 0.2991 | No |
| 74 | Xrcc6 | 4005 | 0.184 | 0.2951 | No |
| 75 | Srsf2 | 4008 | 0.184 | 0.2987 | No |
| 76 | Trim28 | 4010 | 0.184 | 0.3024 | No |
| 77 | H2az1 | 4073 | 0.179 | 0.3009 | No |
| 78 | Rsl1d1 | 4075 | 0.178 | 0.3045 | No |
| 79 | Snrpa1 | 4158 | 0.173 | 0.3013 | No |
| 80 | Hnrnpc | 4182 | 0.171 | 0.3029 | No |
| 81 | Ccna2 | 4269 | 0.164 | 0.2992 | No |
| 82 | Nolc1 | 4375 | 0.156 | 0.2937 | No |
| 83 | Eif4e | 4408 | 0.153 | 0.2942 | No |
| 84 | Snrpd2 | 4483 | 0.148 | 0.2912 | No |
| 85 | Smarcc1 | 4542 | 0.144 | 0.2893 | No |
| 86 | Hspd1 | 4658 | 0.136 | 0.2827 | No |
| 87 | Gnl3 | 4703 | 0.132 | 0.2818 | No |
| 88 | Rrp9 | 4829 | 0.125 | 0.2740 | No |
| 89 | Psmd14 | 4859 | 0.123 | 0.2742 | No |
| 90 | Hnrnpr | 4959 | 0.115 | 0.2684 | No |
| 91 | Ran | 4975 | 0.114 | 0.2695 | No |
| 92 | Tcp1 | 5151 | 0.102 | 0.2572 | No |
| 93 | Psmd7 | 5233 | 0.097 | 0.2525 | No |
| 94 | Erh | 5268 | 0.093 | 0.2516 | No |
| 95 | Hnrnpa1 | 5413 | 0.082 | 0.2415 | No |
| 96 | Ssb | 5433 | 0.082 | 0.2416 | No |
| 97 | Sf3b3 | 5577 | 0.072 | 0.2313 | No |
| 98 | Mcm4 | 5585 | 0.071 | 0.2322 | No |
| 99 | Aimp2 | 5587 | 0.071 | 0.2336 | No |
| 100 | Ilf2 | 5598 | 0.070 | 0.2342 | No |
| 101 | Hnrnpa3 | 5611 | 0.069 | 0.2346 | No |
| 102 | Eif2s1 | 5641 | 0.066 | 0.2336 | No |
| 103 | Hdgf | 5656 | 0.065 | 0.2338 | No |
| 104 | Glo1 | 5725 | 0.061 | 0.2294 | No |
| 105 | Pold2 | 5744 | 0.060 | 0.2292 | No |
| 106 | Txnl4a | 5752 | 0.059 | 0.2298 | No |
| 107 | Mcm2 | 5871 | 0.050 | 0.2211 | No |
| 108 | Rplp0 | 5928 | 0.046 | 0.2175 | No |
| 109 | Rpl6 | 5929 | 0.046 | 0.2184 | No |
| 110 | Cnbp | 6081 | 0.039 | 0.2068 | No |
| 111 | Srsf7 | 6140 | 0.035 | 0.2028 | No |
| 112 | Tra2b | 6270 | 0.025 | 0.1927 | No |
| 113 | Hsp90ab1 | 6301 | 0.023 | 0.1907 | No |
| 114 | Ube2e1 | 6364 | 0.018 | 0.1860 | No |
| 115 | Cops5 | 6413 | 0.014 | 0.1823 | No |
| 116 | Pcbp1 | 6459 | 0.011 | 0.1788 | No |
| 117 | Dek | 6465 | 0.011 | 0.1787 | No |
| 118 | Tufm | 6521 | 0.007 | 0.1743 | No |
| 119 | Psmd1 | 6563 | 0.005 | 0.1710 | No |
| 120 | Slc25a3 | 6569 | 0.005 | 0.1707 | No |
| 121 | Acp1 | 6614 | 0.002 | 0.1671 | No |
| 122 | Rps6 | 6759 | -0.001 | 0.1553 | No |
| 123 | Npm1 | 6984 | -0.017 | 0.1373 | No |
| 124 | Rps2 | 7038 | -0.021 | 0.1334 | No |
| 125 | Psmd8 | 7094 | -0.024 | 0.1293 | No |
| 126 | Srm | 7105 | -0.025 | 0.1290 | No |
| 127 | Psmb2 | 7214 | -0.032 | 0.1208 | No |
| 128 | Hnrnpa2b1 | 7360 | -0.043 | 0.1098 | No |
| 129 | Clns1a | 7369 | -0.043 | 0.1100 | No |
| 130 | Hdac2 | 7392 | -0.046 | 0.1091 | No |
| 131 | Hnrnpd | 7455 | -0.050 | 0.1051 | No |
| 132 | Ppm1g | 7572 | -0.058 | 0.0967 | No |
| 133 | Dhx15 | 7599 | -0.060 | 0.0958 | No |
| 134 | Phb2 | 7754 | -0.072 | 0.0846 | No |
| 135 | Snrpg | 7833 | -0.077 | 0.0798 | No |
| 136 | Cyc1 | 7841 | -0.078 | 0.0808 | No |
| 137 | Ap3s1 | 7863 | -0.079 | 0.0807 | No |
| 138 | Apex1 | 7925 | -0.085 | 0.0774 | No |
| 139 | Stard7 | 7934 | -0.086 | 0.0785 | No |
| 140 | Mrpl23 | 8155 | -0.103 | 0.0626 | No |
| 141 | Mcm7 | 8166 | -0.104 | 0.0639 | No |
| 142 | Ppia | 8175 | -0.105 | 0.0654 | No |
| 143 | Eif3b | 8267 | -0.111 | 0.0601 | No |
| 144 | Rps10 | 8295 | -0.113 | 0.0602 | No |
| 145 | Ifrd1 | 8472 | -0.128 | 0.0484 | No |
| 146 | Tfdp1 | 8553 | -0.133 | 0.0445 | No |
| 147 | Pole3 | 8586 | -0.136 | 0.0447 | No |
| 148 | Psmb3 | 8709 | -0.147 | 0.0377 | No |
| 149 | Psma7 | 8718 | -0.148 | 0.0400 | No |
| 150 | Snrpd1 | 8765 | -0.153 | 0.0394 | No |
| 151 | Canx | 8770 | -0.153 | 0.0422 | No |
| 152 | Ddx21 | 8780 | -0.155 | 0.0446 | No |
| 153 | Impdh2 | 8917 | -0.166 | 0.0368 | No |
| 154 | Gspt1 | 8963 | -0.169 | 0.0366 | No |
| 155 | Ndufab1 | 9007 | -0.172 | 0.0366 | No |
| 156 | Rps5 | 9282 | -0.192 | 0.0180 | No |
| 157 | Rpl22 | 9471 | -0.210 | 0.0068 | No |
| 158 | Ncbp1 | 9540 | -0.217 | 0.0057 | No |
| 159 | Xpo1 | 9566 | -0.219 | 0.0081 | No |
| 160 | Nop16 | 9593 | -0.222 | 0.0105 | No |
| 161 | Psma1 | 9767 | -0.236 | 0.0011 | No |
| 162 | Ldha | 9777 | -0.236 | 0.0052 | No |
| 163 | Rpl14 | 10221 | -0.279 | -0.0254 | No |
| 164 | Eif2s2 | 10364 | -0.294 | -0.0311 | No |
| 165 | Exosc7 | 10619 | -0.322 | -0.0454 | No |
| 166 | Fbl | 10667 | -0.328 | -0.0425 | No |
| 167 | Ranbp1 | 10730 | -0.338 | -0.0407 | No |
| 168 | Etf1 | 10751 | -0.341 | -0.0354 | No |
| 169 | Lsm2 | 10794 | -0.347 | -0.0318 | No |
| 170 | Mrpl9 | 10857 | -0.356 | -0.0296 | No |
| 171 | Psmc4 | 10951 | -0.369 | -0.0297 | No |
| 172 | Ruvbl2 | 10956 | -0.369 | -0.0225 | No |
| 173 | Hddc2 | 10999 | -0.376 | -0.0182 | No |
| 174 | Rps3 | 11111 | -0.392 | -0.0193 | No |
| 175 | Ywhaq | 11149 | -0.394 | -0.0143 | No |
| 176 | Rack1 | 11259 | -0.411 | -0.0149 | No |
| 177 | Ncbp2 | 11329 | -0.421 | -0.0120 | No |
| 178 | Bub3 | 11373 | -0.428 | -0.0067 | No |
| 179 | Cbx3 | 11495 | -0.451 | -0.0075 | No |
| 180 | Rpl18 | 11656 | -0.483 | -0.0107 | No |
| 181 | Eif4h | 11696 | -0.493 | -0.0039 | No |
| 182 | Cct2 | 11728 | -0.499 | 0.0038 | No |
| 183 | Nhp2 | 11736 | -0.502 | 0.0134 | No |
| 184 | Vdac1 | 11800 | -0.520 | 0.0189 | No |
| 185 | Pgk1 | 12042 | -0.605 | 0.0115 | No |
| 186 | Orc2 | 12209 | -0.726 | 0.0126 | No |