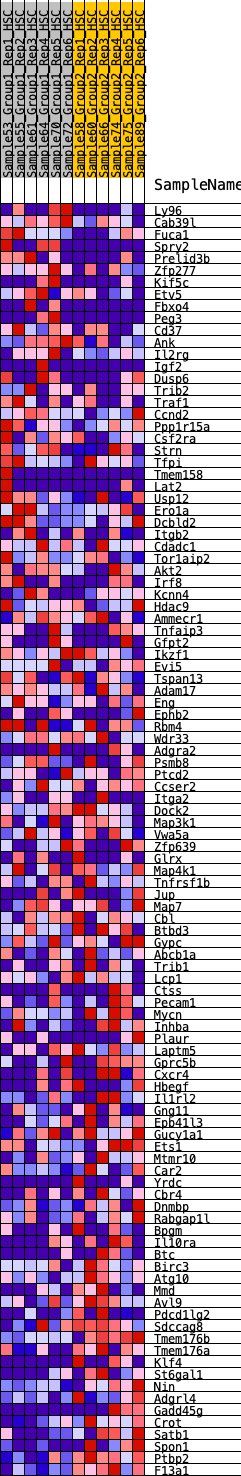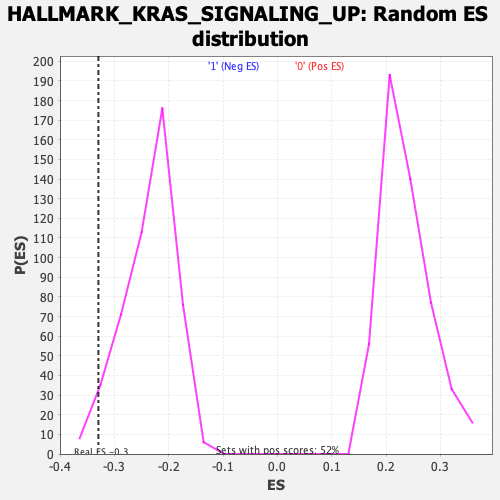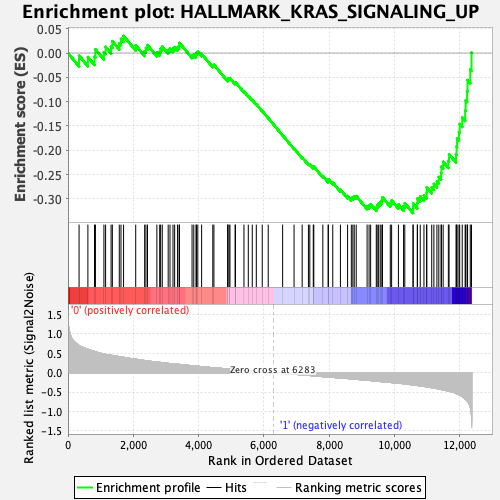
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | HSC.HSC_Pheno.cls#Group1_versus_Group2.HSC_Pheno.cls#Group1_versus_Group2_repos |
| Phenotype | HSC_Pheno.cls#Group1_versus_Group2_repos |
| Upregulated in class | 1 |
| GeneSet | HALLMARK_KRAS_SIGNALING_UP |
| Enrichment Score (ES) | -0.3295553 |
| Normalized Enrichment Score (NES) | -1.3937876 |
| Nominal p-value | 0.04742268 |
| FDR q-value | 0.84850204 |
| FWER p-Value | 0.524 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Ly96 | 339 | 0.697 | -0.0055 | No |
| 2 | Cab39l | 609 | 0.604 | -0.0083 | No |
| 3 | Fuca1 | 812 | 0.549 | -0.0073 | No |
| 4 | Spry2 | 840 | 0.539 | 0.0076 | No |
| 5 | Prelid3b | 1100 | 0.481 | 0.0018 | No |
| 6 | Zfp277 | 1151 | 0.472 | 0.0127 | No |
| 7 | Kif5c | 1319 | 0.449 | 0.0134 | No |
| 8 | Etv5 | 1358 | 0.444 | 0.0244 | No |
| 9 | Fbxo4 | 1570 | 0.413 | 0.0203 | No |
| 10 | Peg3 | 1622 | 0.406 | 0.0291 | No |
| 11 | Cd37 | 1701 | 0.396 | 0.0353 | No |
| 12 | Ank | 2073 | 0.346 | 0.0160 | No |
| 13 | Il2rg | 2347 | 0.314 | 0.0037 | No |
| 14 | Igf2 | 2387 | 0.310 | 0.0104 | No |
| 15 | Dusp6 | 2434 | 0.305 | 0.0163 | No |
| 16 | Trib2 | 2721 | 0.273 | 0.0017 | No |
| 17 | Traf1 | 2807 | 0.266 | 0.0032 | No |
| 18 | Ccnd2 | 2836 | 0.263 | 0.0092 | No |
| 19 | Ppp1r15a | 2886 | 0.257 | 0.0134 | No |
| 20 | Csf2ra | 3067 | 0.240 | 0.0064 | No |
| 21 | Strn | 3123 | 0.234 | 0.0093 | No |
| 22 | Tfpi | 3214 | 0.227 | 0.0092 | No |
| 23 | Tmem158 | 3265 | 0.224 | 0.0122 | No |
| 24 | Lat2 | 3354 | 0.217 | 0.0119 | No |
| 25 | Usp12 | 3400 | 0.214 | 0.0151 | No |
| 26 | Ero1a | 3414 | 0.212 | 0.0207 | No |
| 27 | Dcbld2 | 3798 | 0.179 | -0.0048 | No |
| 28 | Itgb2 | 3851 | 0.176 | -0.0035 | No |
| 29 | Cdadc1 | 3919 | 0.170 | -0.0035 | No |
| 30 | Tor1aip2 | 3935 | 0.169 | 0.0006 | No |
| 31 | Akt2 | 3974 | 0.166 | 0.0028 | No |
| 32 | Irf8 | 4091 | 0.156 | -0.0017 | No |
| 33 | Kcnn4 | 4433 | 0.129 | -0.0255 | No |
| 34 | Hdac9 | 4467 | 0.126 | -0.0241 | No |
| 35 | Ammecr1 | 4887 | 0.098 | -0.0552 | No |
| 36 | Tnfaip3 | 4895 | 0.098 | -0.0526 | No |
| 37 | Gfpt2 | 4931 | 0.096 | -0.0524 | No |
| 38 | Ikzf1 | 4958 | 0.093 | -0.0516 | No |
| 39 | Evi5 | 5116 | 0.082 | -0.0618 | No |
| 40 | Tspan13 | 5127 | 0.081 | -0.0600 | No |
| 41 | Adam17 | 5387 | 0.063 | -0.0792 | No |
| 42 | Eng | 5520 | 0.054 | -0.0882 | No |
| 43 | Ephb2 | 5641 | 0.047 | -0.0965 | No |
| 44 | Rbm4 | 5765 | 0.038 | -0.1053 | No |
| 45 | Wdr33 | 5947 | 0.023 | -0.1194 | No |
| 46 | Adgra2 | 6130 | 0.011 | -0.1339 | No |
| 47 | Psmb8 | 6571 | -0.017 | -0.1692 | No |
| 48 | Ptcd2 | 6921 | -0.042 | -0.1964 | No |
| 49 | Ccser2 | 7169 | -0.057 | -0.2147 | No |
| 50 | Itga2 | 7361 | -0.071 | -0.2280 | No |
| 51 | Dock2 | 7403 | -0.073 | -0.2291 | No |
| 52 | Map3k1 | 7509 | -0.080 | -0.2351 | No |
| 53 | Vwa5a | 7526 | -0.081 | -0.2338 | No |
| 54 | Zfp639 | 7802 | -0.102 | -0.2530 | No |
| 55 | Glrx | 7965 | -0.112 | -0.2627 | No |
| 56 | Map4k1 | 7971 | -0.113 | -0.2595 | No |
| 57 | Tnfrsf1b | 8106 | -0.122 | -0.2665 | No |
| 58 | Jup | 8340 | -0.138 | -0.2811 | No |
| 59 | Map7 | 8560 | -0.152 | -0.2942 | No |
| 60 | Cbl | 8674 | -0.159 | -0.2983 | No |
| 61 | Btbd3 | 8721 | -0.163 | -0.2969 | No |
| 62 | Gypc | 8763 | -0.167 | -0.2950 | No |
| 63 | Abcb1a | 8824 | -0.173 | -0.2944 | No |
| 64 | Trib1 | 9157 | -0.196 | -0.3152 | No |
| 65 | Lcp1 | 9218 | -0.201 | -0.3137 | No |
| 66 | Ctss | 9266 | -0.204 | -0.3111 | No |
| 67 | Pecam1 | 9438 | -0.219 | -0.3181 | No |
| 68 | Mycn | 9465 | -0.221 | -0.3132 | No |
| 69 | Inhba | 9510 | -0.225 | -0.3096 | No |
| 70 | Plaur | 9560 | -0.229 | -0.3063 | No |
| 71 | Laptm5 | 9609 | -0.233 | -0.3028 | No |
| 72 | Gprc5b | 9629 | -0.235 | -0.2969 | No |
| 73 | Cxcr4 | 9867 | -0.253 | -0.3082 | No |
| 74 | Hbegf | 9908 | -0.256 | -0.3034 | No |
| 75 | Il1rl2 | 10116 | -0.272 | -0.3116 | No |
| 76 | Gng11 | 10274 | -0.287 | -0.3153 | No |
| 77 | Epb41l3 | 10311 | -0.291 | -0.3090 | No |
| 78 | Gucy1a1 | 10564 | -0.317 | -0.3195 | Yes |
| 79 | Ets1 | 10567 | -0.318 | -0.3095 | Yes |
| 80 | Mtmr10 | 10694 | -0.331 | -0.3093 | Yes |
| 81 | Car2 | 10699 | -0.331 | -0.2991 | Yes |
| 82 | Yrdc | 10784 | -0.342 | -0.2951 | Yes |
| 83 | Cbr4 | 10898 | -0.356 | -0.2930 | Yes |
| 84 | Dnmbp | 10980 | -0.365 | -0.2880 | Yes |
| 85 | Rabgap1l | 10984 | -0.366 | -0.2766 | Yes |
| 86 | Bpgm | 11136 | -0.387 | -0.2766 | Yes |
| 87 | Il10ra | 11202 | -0.396 | -0.2693 | Yes |
| 88 | Btc | 11295 | -0.412 | -0.2637 | Yes |
| 89 | Birc3 | 11353 | -0.422 | -0.2550 | Yes |
| 90 | Atg10 | 11422 | -0.434 | -0.2468 | Yes |
| 91 | Mmd | 11433 | -0.436 | -0.2337 | Yes |
| 92 | Avl9 | 11486 | -0.444 | -0.2239 | Yes |
| 93 | Pdcd1lg2 | 11645 | -0.474 | -0.2217 | Yes |
| 94 | Sdccag8 | 11666 | -0.477 | -0.2081 | Yes |
| 95 | Tmem176b | 11880 | -0.539 | -0.2084 | Yes |
| 96 | Tmem176a | 11897 | -0.543 | -0.1924 | Yes |
| 97 | Klf4 | 11912 | -0.550 | -0.1761 | Yes |
| 98 | St6gal1 | 11970 | -0.571 | -0.1626 | Yes |
| 99 | Nin | 11992 | -0.576 | -0.1460 | Yes |
| 100 | Adgrl4 | 12070 | -0.614 | -0.1328 | Yes |
| 101 | Gadd45g | 12159 | -0.678 | -0.1184 | Yes |
| 102 | Crot | 12174 | -0.690 | -0.0977 | Yes |
| 103 | Satb1 | 12219 | -0.725 | -0.0782 | Yes |
| 104 | Spon1 | 12233 | -0.743 | -0.0557 | Yes |
| 105 | Ptbp2 | 12316 | -0.899 | -0.0338 | Yes |
| 106 | F13a1 | 12354 | -1.181 | 0.0007 | Yes |