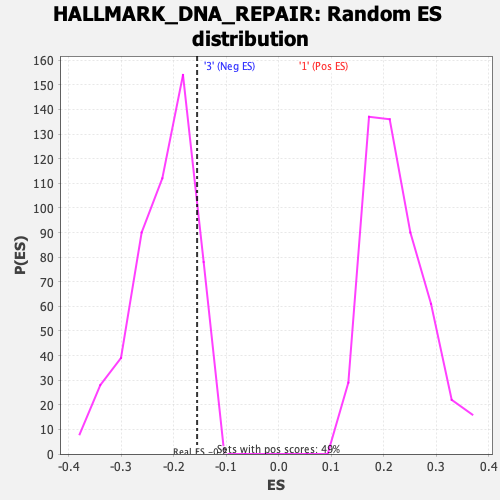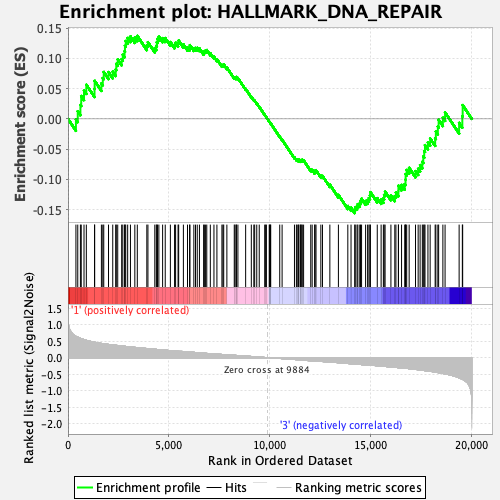
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | GMP.GMP.neu_Pheno.cls #Group2_versus_Group4.GMP.neu_Pheno.cls #Group2_versus_Group4_repos |
| Phenotype | GMP.neu_Pheno.cls#Group2_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_DNA_REPAIR |
| Enrichment Score (ES) | -0.15572955 |
| Normalized Enrichment Score (NES) | -0.70764476 |
| Nominal p-value | 0.90176815 |
| FDR q-value | 0.88131654 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Ercc1 | 393 | 0.645 | -0.0010 | No |
| 2 | Nelfcd | 484 | 0.620 | 0.0125 | No |
| 3 | Pnp | 615 | 0.585 | 0.0230 | No |
| 4 | Vps37b | 656 | 0.576 | 0.0378 | No |
| 5 | Srsf6 | 794 | 0.548 | 0.0469 | No |
| 6 | Polr2d | 909 | 0.526 | 0.0564 | No |
| 7 | Edf1 | 1315 | 0.466 | 0.0496 | No |
| 8 | Polr3c | 1323 | 0.465 | 0.0628 | No |
| 9 | Polr2e | 1656 | 0.431 | 0.0587 | No |
| 10 | Nt5c | 1726 | 0.424 | 0.0675 | No |
| 11 | Ada | 1773 | 0.421 | 0.0775 | No |
| 12 | Mrpl40 | 2009 | 0.401 | 0.0773 | No |
| 13 | Gtf2f1 | 2219 | 0.385 | 0.0780 | No |
| 14 | Ercc2 | 2366 | 0.373 | 0.0816 | No |
| 15 | Ssrp1 | 2394 | 0.372 | 0.0910 | No |
| 16 | Polr2a | 2466 | 0.368 | 0.0982 | No |
| 17 | Sf3a3 | 2662 | 0.353 | 0.0986 | No |
| 18 | Taf10 | 2718 | 0.349 | 0.1060 | No |
| 19 | Taf9 | 2806 | 0.344 | 0.1117 | No |
| 20 | Alyref | 2825 | 0.343 | 0.1207 | No |
| 21 | Ak3 | 2863 | 0.339 | 0.1288 | No |
| 22 | Ddb1 | 2959 | 0.334 | 0.1337 | No |
| 23 | Hcls1 | 3097 | 0.323 | 0.1362 | No |
| 24 | Eif1b | 3314 | 0.311 | 0.1344 | No |
| 25 | Polr1c | 3437 | 0.304 | 0.1371 | No |
| 26 | Gtf2h1 | 3909 | 0.276 | 0.1215 | No |
| 27 | Cda | 3964 | 0.273 | 0.1267 | No |
| 28 | Nme3 | 4305 | 0.254 | 0.1170 | No |
| 29 | Ercc3 | 4379 | 0.250 | 0.1206 | No |
| 30 | Pde6g | 4406 | 0.249 | 0.1266 | No |
| 31 | Tarbp2 | 4437 | 0.246 | 0.1322 | No |
| 32 | Nfx1 | 4506 | 0.242 | 0.1359 | No |
| 33 | Pcna | 4692 | 0.235 | 0.1334 | No |
| 34 | Poll | 4822 | 0.227 | 0.1335 | No |
| 35 | Nelfb | 5077 | 0.213 | 0.1269 | No |
| 36 | Polr2j | 5285 | 0.203 | 0.1225 | No |
| 37 | Rfc4 | 5336 | 0.200 | 0.1258 | No |
| 38 | Polh | 5459 | 0.193 | 0.1253 | No |
| 39 | Impdh2 | 5488 | 0.191 | 0.1294 | No |
| 40 | Pold3 | 5722 | 0.184 | 0.1231 | No |
| 41 | Umps | 5926 | 0.174 | 0.1179 | No |
| 42 | Fen1 | 6030 | 0.170 | 0.1177 | No |
| 43 | Polr1d | 6047 | 0.169 | 0.1218 | No |
| 44 | Rad51 | 6241 | 0.161 | 0.1168 | No |
| 45 | Snapc4 | 6324 | 0.156 | 0.1172 | No |
| 46 | Adcy6 | 6410 | 0.152 | 0.1173 | No |
| 47 | Npr2 | 6518 | 0.149 | 0.1163 | No |
| 48 | Lig1 | 6726 | 0.137 | 0.1099 | No |
| 49 | Sec61a1 | 6763 | 0.136 | 0.1120 | No |
| 50 | Cmpk2 | 6824 | 0.132 | 0.1128 | No |
| 51 | Taf6 | 6882 | 0.129 | 0.1137 | No |
| 52 | Ercc4 | 7058 | 0.120 | 0.1084 | No |
| 53 | Nudt21 | 7238 | 0.112 | 0.1027 | No |
| 54 | Supt5 | 7385 | 0.108 | 0.0985 | No |
| 55 | Cetn2 | 7633 | 0.096 | 0.0889 | No |
| 56 | Taf13 | 7683 | 0.095 | 0.0891 | No |
| 57 | Surf1 | 7722 | 0.092 | 0.0899 | No |
| 58 | Dad1 | 7879 | 0.089 | 0.0847 | No |
| 59 | Aprt | 8245 | 0.073 | 0.0684 | No |
| 60 | Gtf2b | 8283 | 0.071 | 0.0686 | No |
| 61 | Bcap31 | 8345 | 0.068 | 0.0675 | No |
| 62 | Gtf2a2 | 8350 | 0.067 | 0.0693 | No |
| 63 | Dgcr8 | 8426 | 0.064 | 0.0674 | No |
| 64 | Gtf3c5 | 8810 | 0.048 | 0.0495 | No |
| 65 | Cant1 | 9091 | 0.036 | 0.0365 | No |
| 66 | Ercc5 | 9216 | 0.031 | 0.0312 | No |
| 67 | Eloa | 9258 | 0.030 | 0.0300 | No |
| 68 | Tk2 | 9361 | 0.024 | 0.0255 | No |
| 69 | Smad5 | 9484 | 0.019 | 0.0199 | No |
| 70 | Polr2f | 9755 | 0.006 | 0.0065 | No |
| 71 | Arl6ip1 | 9800 | 0.005 | 0.0045 | No |
| 72 | Brf2 | 9839 | 0.002 | 0.0026 | No |
| 73 | Ccno | 9983 | -0.001 | -0.0045 | No |
| 74 | Polr2k | 9988 | -0.001 | -0.0047 | No |
| 75 | Clp1 | 10044 | -0.004 | -0.0073 | No |
| 76 | Dctn4 | 10052 | -0.005 | -0.0076 | No |
| 77 | Sdcbp | 10500 | -0.023 | -0.0294 | No |
| 78 | Dguok | 10618 | -0.028 | -0.0344 | No |
| 79 | Gtf2h5 | 11231 | -0.054 | -0.0637 | No |
| 80 | Cstf3 | 11326 | -0.058 | -0.0667 | No |
| 81 | Xpc | 11367 | -0.060 | -0.0670 | No |
| 82 | Nme1 | 11437 | -0.063 | -0.0686 | No |
| 83 | Ago4 | 11448 | -0.064 | -0.0672 | No |
| 84 | Rrm2b | 11525 | -0.067 | -0.0691 | No |
| 85 | Cox17 | 11545 | -0.069 | -0.0680 | No |
| 86 | Gtf2h3 | 11592 | -0.071 | -0.0683 | No |
| 87 | Pole4 | 11616 | -0.072 | -0.0673 | No |
| 88 | Rae1 | 11684 | -0.075 | -0.0685 | No |
| 89 | Sac3d1 | 12039 | -0.092 | -0.0836 | No |
| 90 | Mpc2 | 12108 | -0.095 | -0.0843 | No |
| 91 | Dut | 12231 | -0.101 | -0.0875 | No |
| 92 | Adrm1 | 12251 | -0.102 | -0.0855 | No |
| 93 | Pold1 | 12299 | -0.104 | -0.0848 | No |
| 94 | Polb | 12539 | -0.113 | -0.0935 | No |
| 95 | Pom121 | 12620 | -0.115 | -0.0942 | No |
| 96 | Rnmt | 12986 | -0.132 | -0.1087 | No |
| 97 | Snapc5 | 13410 | -0.150 | -0.1256 | No |
| 98 | Rbx1 | 13877 | -0.175 | -0.1440 | No |
| 99 | Taf1c | 14036 | -0.183 | -0.1466 | No |
| 100 | Ak1 | 14219 | -0.191 | -0.1502 | Yes |
| 101 | Gpx4 | 14245 | -0.192 | -0.1458 | Yes |
| 102 | Polr2h | 14333 | -0.197 | -0.1445 | Yes |
| 103 | Nelfe | 14375 | -0.199 | -0.1407 | Yes |
| 104 | Pola1 | 14470 | -0.203 | -0.1396 | Yes |
| 105 | Rala | 14505 | -0.205 | -0.1353 | Yes |
| 106 | Nudt9 | 14559 | -0.208 | -0.1319 | Yes |
| 107 | Zwint | 14759 | -0.217 | -0.1356 | Yes |
| 108 | Usp11 | 14858 | -0.223 | -0.1340 | Yes |
| 109 | Polr2i | 14924 | -0.226 | -0.1307 | Yes |
| 110 | Rfc5 | 14974 | -0.228 | -0.1266 | Yes |
| 111 | Polr1h | 14997 | -0.229 | -0.1210 | Yes |
| 112 | Ddb2 | 15337 | -0.243 | -0.1310 | Yes |
| 113 | Gmpr2 | 15528 | -0.253 | -0.1332 | Yes |
| 114 | Rad52 | 15640 | -0.259 | -0.1312 | Yes |
| 115 | Nt5c3 | 15679 | -0.261 | -0.1255 | Yes |
| 116 | Aaas | 15725 | -0.264 | -0.1201 | Yes |
| 117 | Polr2c | 16014 | -0.280 | -0.1264 | Yes |
| 118 | Zfp707 | 16207 | -0.291 | -0.1276 | Yes |
| 119 | Ncbp2 | 16262 | -0.293 | -0.1218 | Yes |
| 120 | Tsg101 | 16386 | -0.302 | -0.1192 | Yes |
| 121 | Mpg | 16387 | -0.302 | -0.1104 | Yes |
| 122 | Prim1 | 16542 | -0.310 | -0.1091 | Yes |
| 123 | Polr3gl | 16699 | -0.318 | -0.1077 | Yes |
| 124 | Pold4 | 16739 | -0.321 | -0.1003 | Yes |
| 125 | Rev3l | 16744 | -0.321 | -0.0912 | Yes |
| 126 | Gsdme | 16796 | -0.324 | -0.0843 | Yes |
| 127 | Taf12 | 16919 | -0.332 | -0.0808 | Yes |
| 128 | Tyms | 17234 | -0.354 | -0.0863 | Yes |
| 129 | Stx3 | 17365 | -0.363 | -0.0823 | Yes |
| 130 | Ercc8 | 17462 | -0.370 | -0.0763 | Yes |
| 131 | Pola2 | 17572 | -0.380 | -0.0707 | Yes |
| 132 | Upf3b | 17620 | -0.384 | -0.0619 | Yes |
| 133 | Pde4b | 17666 | -0.388 | -0.0529 | Yes |
| 134 | Supt4a | 17704 | -0.391 | -0.0434 | Yes |
| 135 | Trp53 | 17850 | -0.406 | -0.0388 | Yes |
| 136 | Nme4 | 17960 | -0.413 | -0.0323 | Yes |
| 137 | Rpa2 | 18208 | -0.437 | -0.0320 | Yes |
| 138 | Polr2g | 18243 | -0.440 | -0.0209 | Yes |
| 139 | Rfc2 | 18342 | -0.450 | -0.0128 | Yes |
| 140 | Itpa | 18381 | -0.453 | -0.0015 | Yes |
| 141 | Vps28 | 18591 | -0.478 | 0.0019 | Yes |
| 142 | Rpa3 | 18700 | -0.487 | 0.0107 | Yes |
| 143 | Ell | 19397 | -0.611 | -0.0066 | Yes |
| 144 | Guk1 | 19556 | -0.652 | 0.0045 | Yes |
| 145 | Rfc3 | 19573 | -0.657 | 0.0228 | Yes |