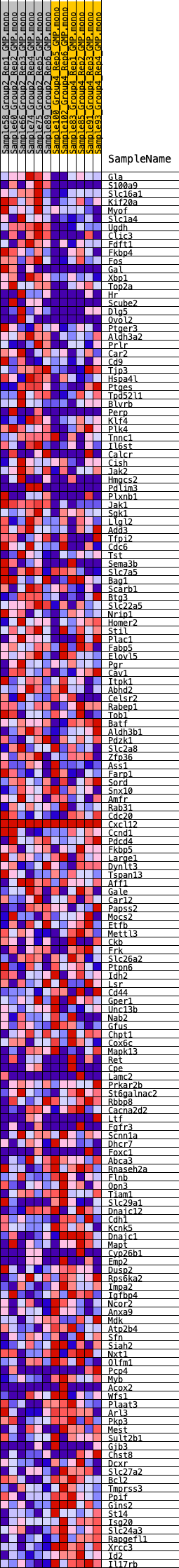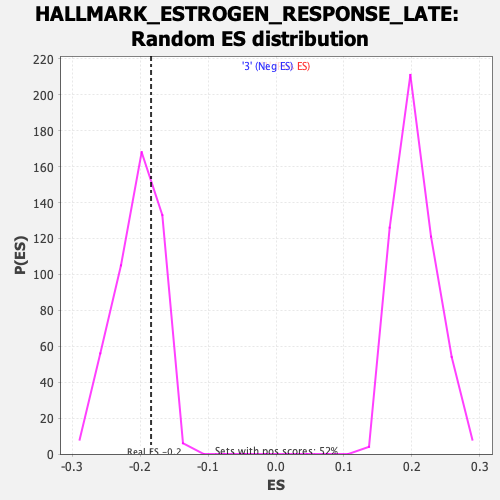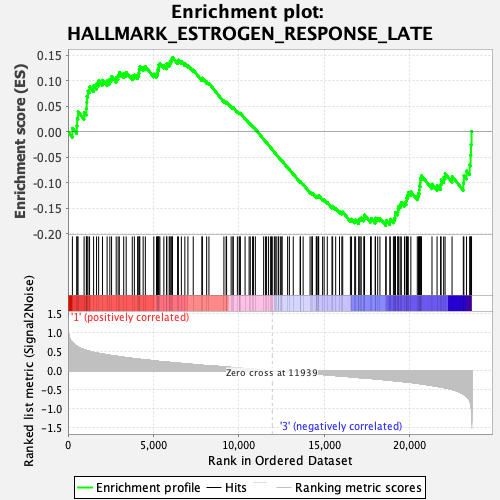
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | GMP.GMP.mono_Pheno.cls #Group2_versus_Group4.GMP.mono_Pheno.cls #Group2_versus_Group4_repos |
| Phenotype | GMP.mono_Pheno.cls#Group2_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_ESTROGEN_RESPONSE_LATE |
| Enrichment Score (ES) | -0.18417974 |
| Normalized Enrichment Score (NES) | -0.90125495 |
| Nominal p-value | 0.697479 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Gla | 257 | 0.740 | 0.0071 | No |
| 2 | S100a9 | 512 | 0.646 | 0.0120 | No |
| 3 | Slc16a1 | 537 | 0.637 | 0.0265 | No |
| 4 | Kif20a | 588 | 0.620 | 0.0395 | No |
| 5 | Myof | 941 | 0.555 | 0.0380 | No |
| 6 | Slc1a4 | 1073 | 0.535 | 0.0455 | No |
| 7 | Ugdh | 1093 | 0.530 | 0.0576 | No |
| 8 | Clic3 | 1108 | 0.528 | 0.0698 | No |
| 9 | Fdft1 | 1167 | 0.522 | 0.0801 | No |
| 10 | Fkbp4 | 1262 | 0.512 | 0.0886 | No |
| 11 | Fos | 1497 | 0.484 | 0.0904 | No |
| 12 | Gal | 1669 | 0.464 | 0.0944 | No |
| 13 | Xbp1 | 1792 | 0.456 | 0.1003 | No |
| 14 | Top2a | 2019 | 0.440 | 0.1014 | No |
| 15 | Hr | 2288 | 0.417 | 0.1001 | No |
| 16 | Scube2 | 2448 | 0.403 | 0.1032 | No |
| 17 | Dlg5 | 2549 | 0.398 | 0.1086 | No |
| 18 | Ovol2 | 2819 | 0.382 | 0.1064 | No |
| 19 | Ptger3 | 2931 | 0.374 | 0.1108 | No |
| 20 | Aldh3a2 | 3002 | 0.367 | 0.1168 | No |
| 21 | Prlr | 3259 | 0.352 | 0.1145 | No |
| 22 | Car2 | 3404 | 0.343 | 0.1167 | No |
| 23 | Cd9 | 3760 | 0.322 | 0.1094 | No |
| 24 | Tjp3 | 3883 | 0.313 | 0.1118 | No |
| 25 | Hspa4l | 4072 | 0.307 | 0.1113 | No |
| 26 | Ptges | 4148 | 0.302 | 0.1154 | No |
| 27 | Tpd52l1 | 4160 | 0.301 | 0.1223 | No |
| 28 | Blvrb | 4201 | 0.299 | 0.1279 | No |
| 29 | Perp | 4396 | 0.289 | 0.1267 | No |
| 30 | Klf4 | 4524 | 0.283 | 0.1282 | No |
| 31 | Plk4 | 5020 | 0.259 | 0.1134 | No |
| 32 | Tnnc1 | 5178 | 0.250 | 0.1128 | No |
| 33 | Il6st | 5239 | 0.247 | 0.1162 | No |
| 34 | Calcr | 5251 | 0.246 | 0.1218 | No |
| 35 | Cish | 5302 | 0.244 | 0.1256 | No |
| 36 | Jak2 | 5312 | 0.243 | 0.1311 | No |
| 37 | Hmgcs2 | 5375 | 0.240 | 0.1343 | No |
| 38 | Pdlim3 | 5608 | 0.229 | 0.1300 | No |
| 39 | Plxnb1 | 5769 | 0.227 | 0.1287 | No |
| 40 | Jak1 | 5780 | 0.226 | 0.1338 | No |
| 41 | Sgk1 | 5920 | 0.219 | 0.1332 | No |
| 42 | Llgl2 | 5964 | 0.217 | 0.1367 | No |
| 43 | Add3 | 6019 | 0.214 | 0.1396 | No |
| 44 | Tfpi2 | 6068 | 0.211 | 0.1427 | No |
| 45 | Cdc6 | 6115 | 0.208 | 0.1458 | No |
| 46 | Tst | 6411 | 0.199 | 0.1381 | No |
| 47 | Sema3b | 6459 | 0.197 | 0.1409 | No |
| 48 | Slc7a5 | 6632 | 0.189 | 0.1381 | No |
| 49 | Bag1 | 6836 | 0.182 | 0.1339 | No |
| 50 | Scarb1 | 7015 | 0.175 | 0.1306 | No |
| 51 | Btg3 | 7326 | 0.162 | 0.1213 | No |
| 52 | Slc22a5 | 7830 | 0.140 | 0.1033 | No |
| 53 | Nrip1 | 7863 | 0.139 | 0.1053 | No |
| 54 | Homer2 | 8112 | 0.130 | 0.0979 | No |
| 55 | Stil | 8245 | 0.125 | 0.0954 | No |
| 56 | Plac1 | 9121 | 0.098 | 0.0604 | No |
| 57 | Fabp5 | 9250 | 0.094 | 0.0573 | No |
| 58 | Elovl5 | 9279 | 0.093 | 0.0583 | No |
| 59 | Pgr | 9544 | 0.083 | 0.0491 | No |
| 60 | Cav1 | 9646 | 0.079 | 0.0467 | No |
| 61 | Itpk1 | 9668 | 0.078 | 0.0478 | No |
| 62 | Abhd2 | 9909 | 0.070 | 0.0392 | No |
| 63 | Celsr2 | 10012 | 0.066 | 0.0365 | No |
| 64 | Rabep1 | 10047 | 0.065 | 0.0366 | No |
| 65 | Tob1 | 10085 | 0.063 | 0.0366 | No |
| 66 | Batf | 10359 | 0.054 | 0.0262 | No |
| 67 | Aldh3b1 | 10595 | 0.046 | 0.0173 | No |
| 68 | Pdzk1 | 10664 | 0.043 | 0.0155 | No |
| 69 | Slc2a8 | 10798 | 0.039 | 0.0108 | No |
| 70 | Zfp36 | 10849 | 0.037 | 0.0095 | No |
| 71 | Ass1 | 10960 | 0.033 | 0.0057 | No |
| 72 | Farp1 | 11446 | 0.017 | -0.0146 | No |
| 73 | Sord | 11557 | 0.013 | -0.0190 | No |
| 74 | Snx10 | 11618 | 0.010 | -0.0213 | No |
| 75 | Amfr | 11734 | 0.007 | -0.0260 | No |
| 76 | Rab31 | 11854 | 0.003 | -0.0310 | No |
| 77 | Cdc20 | 11902 | 0.001 | -0.0330 | No |
| 78 | Cxcl12 | 11951 | 0.000 | -0.0351 | No |
| 79 | Ccnd1 | 12071 | -0.001 | -0.0401 | No |
| 80 | Pdcd4 | 12117 | -0.002 | -0.0420 | No |
| 81 | Fkbp5 | 12181 | -0.005 | -0.0445 | No |
| 82 | Large1 | 12296 | -0.009 | -0.0492 | No |
| 83 | Dynlt3 | 12322 | -0.010 | -0.0500 | No |
| 84 | Tspan13 | 12424 | -0.014 | -0.0540 | No |
| 85 | Aff1 | 12512 | -0.017 | -0.0573 | No |
| 86 | Gale | 12519 | -0.017 | -0.0571 | No |
| 87 | Car12 | 12848 | -0.028 | -0.0704 | No |
| 88 | Papss2 | 12951 | -0.032 | -0.0740 | No |
| 89 | Mocs2 | 13178 | -0.040 | -0.0827 | No |
| 90 | Etfb | 13580 | -0.054 | -0.0984 | No |
| 91 | Mettl3 | 13599 | -0.055 | -0.0979 | No |
| 92 | Ckb | 13756 | -0.059 | -0.1031 | No |
| 93 | Frk | 14162 | -0.073 | -0.1185 | No |
| 94 | Slc26a2 | 14262 | -0.077 | -0.1209 | No |
| 95 | Ptpn6 | 14299 | -0.079 | -0.1205 | No |
| 96 | Idh2 | 14522 | -0.088 | -0.1278 | No |
| 97 | Lsr | 14543 | -0.089 | -0.1265 | No |
| 98 | Cd44 | 14616 | -0.091 | -0.1274 | No |
| 99 | Gper1 | 14653 | -0.092 | -0.1267 | No |
| 100 | Unc13b | 14666 | -0.092 | -0.1249 | No |
| 101 | Nab2 | 14896 | -0.101 | -0.1322 | No |
| 102 | Gfus | 14987 | -0.104 | -0.1335 | No |
| 103 | Chpt1 | 15169 | -0.112 | -0.1385 | No |
| 104 | Cox6c | 15443 | -0.122 | -0.1472 | No |
| 105 | Mapk13 | 15483 | -0.124 | -0.1458 | No |
| 106 | Ret | 15660 | -0.129 | -0.1502 | No |
| 107 | Cpe | 15885 | -0.138 | -0.1564 | No |
| 108 | Lamc2 | 16020 | -0.143 | -0.1586 | No |
| 109 | Prkar2b | 16072 | -0.144 | -0.1573 | No |
| 110 | St6galnac2 | 16521 | -0.161 | -0.1724 | No |
| 111 | Rbbp8 | 16588 | -0.165 | -0.1712 | No |
| 112 | Cacna2d2 | 16772 | -0.172 | -0.1749 | No |
| 113 | Ltf | 16826 | -0.174 | -0.1729 | No |
| 114 | Fgfr3 | 17012 | -0.181 | -0.1764 | No |
| 115 | Scnn1a | 17022 | -0.182 | -0.1723 | No |
| 116 | Dhcr7 | 17088 | -0.184 | -0.1706 | No |
| 117 | Foxc1 | 17153 | -0.185 | -0.1688 | No |
| 118 | Abca3 | 17306 | -0.190 | -0.1706 | No |
| 119 | Rnaseh2a | 17341 | -0.191 | -0.1674 | No |
| 120 | Flnb | 17349 | -0.191 | -0.1631 | No |
| 121 | Opn3 | 17698 | -0.205 | -0.1729 | No |
| 122 | Tiam1 | 17744 | -0.206 | -0.1698 | No |
| 123 | Slc29a1 | 17975 | -0.216 | -0.1743 | No |
| 124 | Dnajc12 | 17981 | -0.216 | -0.1693 | No |
| 125 | Cdh1 | 18118 | -0.221 | -0.1697 | No |
| 126 | Kcnk5 | 18249 | -0.226 | -0.1697 | No |
| 127 | Dnajc1 | 18589 | -0.240 | -0.1783 | Yes |
| 128 | Mapt | 18614 | -0.241 | -0.1735 | Yes |
| 129 | Cyp26b1 | 18830 | -0.252 | -0.1765 | Yes |
| 130 | Emp2 | 18853 | -0.253 | -0.1713 | Yes |
| 131 | Dusp2 | 19037 | -0.263 | -0.1727 | Yes |
| 132 | Rps6ka2 | 19106 | -0.266 | -0.1691 | Yes |
| 133 | Impa2 | 19143 | -0.268 | -0.1641 | Yes |
| 134 | Igfbp4 | 19156 | -0.268 | -0.1581 | Yes |
| 135 | Ncor2 | 19291 | -0.275 | -0.1571 | Yes |
| 136 | Anxa9 | 19308 | -0.276 | -0.1510 | Yes |
| 137 | Mdk | 19350 | -0.278 | -0.1460 | Yes |
| 138 | Atp2b4 | 19436 | -0.280 | -0.1428 | Yes |
| 139 | Sfn | 19500 | -0.284 | -0.1386 | Yes |
| 140 | Siah2 | 19684 | -0.293 | -0.1392 | Yes |
| 141 | Nxt1 | 19783 | -0.299 | -0.1361 | Yes |
| 142 | Olfm1 | 19804 | -0.301 | -0.1296 | Yes |
| 143 | Pcp4 | 19858 | -0.305 | -0.1245 | Yes |
| 144 | Myb | 19908 | -0.307 | -0.1191 | Yes |
| 145 | Acox2 | 20058 | -0.313 | -0.1178 | Yes |
| 146 | Wfs1 | 20434 | -0.332 | -0.1257 | Yes |
| 147 | Plaat3 | 20504 | -0.336 | -0.1205 | Yes |
| 148 | Arl3 | 20555 | -0.339 | -0.1144 | Yes |
| 149 | Pkp3 | 20558 | -0.339 | -0.1062 | Yes |
| 150 | Mest | 20608 | -0.344 | -0.0999 | Yes |
| 151 | Sult2b1 | 20612 | -0.344 | -0.0916 | Yes |
| 152 | Gjb3 | 20680 | -0.348 | -0.0860 | Yes |
| 153 | Chst8 | 21294 | -0.391 | -0.1026 | Yes |
| 154 | Dcxr | 21591 | -0.413 | -0.1052 | Yes |
| 155 | Slc27a2 | 21798 | -0.433 | -0.1034 | Yes |
| 156 | Bcl2 | 21828 | -0.435 | -0.0940 | Yes |
| 157 | Tmprss3 | 21977 | -0.445 | -0.0895 | Yes |
| 158 | Ppif | 22060 | -0.454 | -0.0820 | Yes |
| 159 | Gins2 | 22468 | -0.496 | -0.0872 | Yes |
| 160 | St14 | 23127 | -0.616 | -0.1003 | Yes |
| 161 | Isg20 | 23159 | -0.624 | -0.0864 | Yes |
| 162 | Slc24a3 | 23315 | -0.677 | -0.0765 | Yes |
| 163 | Rapgefl1 | 23498 | -0.789 | -0.0650 | Yes |
| 164 | Xrcc3 | 23552 | -0.863 | -0.0462 | Yes |
| 165 | Id2 | 23574 | -0.922 | -0.0247 | Yes |
| 166 | Il17rb | 23608 | -1.099 | 0.0007 | Yes |