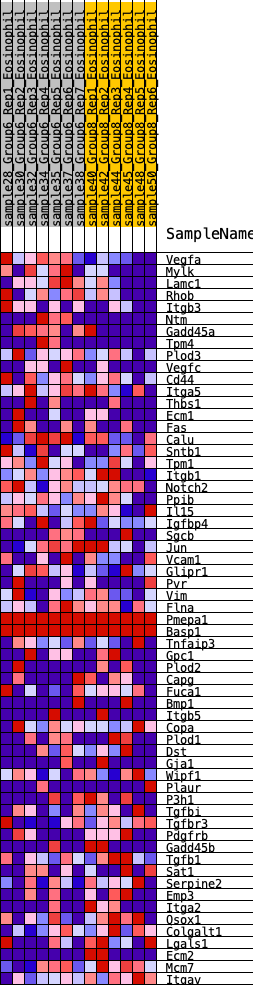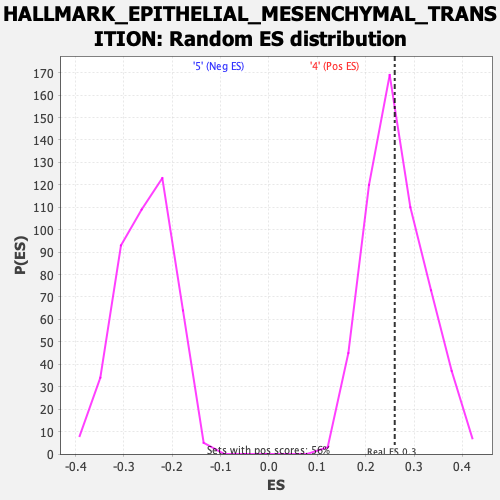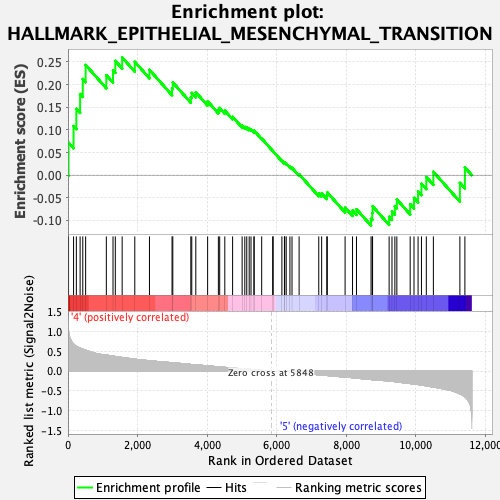
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Eosinophil.Eosinophil_Pheno.cls #Group6_versus_Group8.Eosinophil_Pheno.cls #Group6_versus_Group8_repos |
| Phenotype | Eosinophil_Pheno.cls#Group6_versus_Group8_repos |
| Upregulated in class | 4 |
| GeneSet | HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION |
| Enrichment Score (ES) | 0.26000005 |
| Normalized Enrichment Score (NES) | 0.9862162 |
| Nominal p-value | 0.46985817 |
| FDR q-value | 0.9713058 |
| FWER p-Value | 0.991 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Vegfa | 16 | 1.020 | 0.0717 | Yes |
| 2 | Mylk | 160 | 0.686 | 0.1085 | Yes |
| 3 | Lamc1 | 240 | 0.627 | 0.1467 | Yes |
| 4 | Rhob | 348 | 0.583 | 0.1792 | Yes |
| 5 | Itgb3 | 425 | 0.555 | 0.2124 | Yes |
| 6 | Ntm | 507 | 0.529 | 0.2433 | Yes |
| 7 | Gadd45a | 1102 | 0.403 | 0.2208 | Yes |
| 8 | Tpm4 | 1294 | 0.378 | 0.2314 | Yes |
| 9 | Plod3 | 1360 | 0.369 | 0.2523 | Yes |
| 10 | Vegfc | 1557 | 0.345 | 0.2600 | Yes |
| 11 | Cd44 | 1920 | 0.301 | 0.2503 | No |
| 12 | Itga5 | 2342 | 0.262 | 0.2326 | No |
| 13 | Thbs1 | 2991 | 0.211 | 0.1916 | No |
| 14 | Ecm1 | 3013 | 0.209 | 0.2047 | No |
| 15 | Fas | 3531 | 0.168 | 0.1720 | No |
| 16 | Calu | 3558 | 0.166 | 0.1816 | No |
| 17 | Sntb1 | 3674 | 0.158 | 0.1830 | No |
| 18 | Tpm1 | 4014 | 0.132 | 0.1631 | No |
| 19 | Itgb1 | 4323 | 0.108 | 0.1441 | No |
| 20 | Notch2 | 4361 | 0.105 | 0.1485 | No |
| 21 | Ppib | 4508 | 0.094 | 0.1426 | No |
| 22 | Il15 | 4732 | 0.078 | 0.1289 | No |
| 23 | Igfbp4 | 5006 | 0.058 | 0.1094 | No |
| 24 | Sgcb | 5083 | 0.052 | 0.1066 | No |
| 25 | Jun | 5139 | 0.050 | 0.1054 | No |
| 26 | Vcam1 | 5209 | 0.045 | 0.1026 | No |
| 27 | Glipr1 | 5261 | 0.041 | 0.1011 | No |
| 28 | Pvr | 5346 | 0.034 | 0.0963 | No |
| 29 | Vim | 5351 | 0.034 | 0.0984 | No |
| 30 | Flna | 5568 | 0.018 | 0.0810 | No |
| 31 | Pmepa1 | 5884 | 0.000 | 0.0537 | No |
| 32 | Basp1 | 5898 | 0.000 | 0.0526 | No |
| 33 | Tnfaip3 | 6149 | -0.013 | 0.0319 | No |
| 34 | Gpc1 | 6222 | -0.019 | 0.0270 | No |
| 35 | Plod2 | 6230 | -0.019 | 0.0278 | No |
| 36 | Capg | 6273 | -0.023 | 0.0258 | No |
| 37 | Fuca1 | 6380 | -0.031 | 0.0188 | No |
| 38 | Bmp1 | 6440 | -0.035 | 0.0162 | No |
| 39 | Itgb5 | 6644 | -0.050 | 0.0022 | No |
| 40 | Copa | 7209 | -0.093 | -0.0400 | No |
| 41 | Plod1 | 7292 | -0.099 | -0.0400 | No |
| 42 | Dst | 7440 | -0.109 | -0.0449 | No |
| 43 | Gja1 | 7454 | -0.111 | -0.0381 | No |
| 44 | Wipf1 | 7966 | -0.150 | -0.0715 | No |
| 45 | Plaur | 8180 | -0.167 | -0.0780 | No |
| 46 | P3h1 | 8293 | -0.176 | -0.0751 | No |
| 47 | Tgfbi | 8712 | -0.213 | -0.0960 | No |
| 48 | Tgfbr3 | 8748 | -0.217 | -0.0835 | No |
| 49 | Pdgfrb | 8756 | -0.218 | -0.0685 | No |
| 50 | Gadd45b | 9232 | -0.249 | -0.0918 | No |
| 51 | Tgfb1 | 9314 | -0.258 | -0.0804 | No |
| 52 | Sat1 | 9397 | -0.268 | -0.0683 | No |
| 53 | Serpine2 | 9454 | -0.273 | -0.0535 | No |
| 54 | Emp3 | 9837 | -0.315 | -0.0640 | No |
| 55 | Itga2 | 9947 | -0.327 | -0.0500 | No |
| 56 | Qsox1 | 10064 | -0.340 | -0.0357 | No |
| 57 | Colgalt1 | 10161 | -0.352 | -0.0187 | No |
| 58 | Lgals1 | 10301 | -0.373 | -0.0040 | No |
| 59 | Ecm2 | 10504 | -0.404 | 0.0075 | No |
| 60 | Mcm7 | 11266 | -0.578 | -0.0170 | No |
| 61 | Itgav | 11410 | -0.650 | 0.0172 | No |