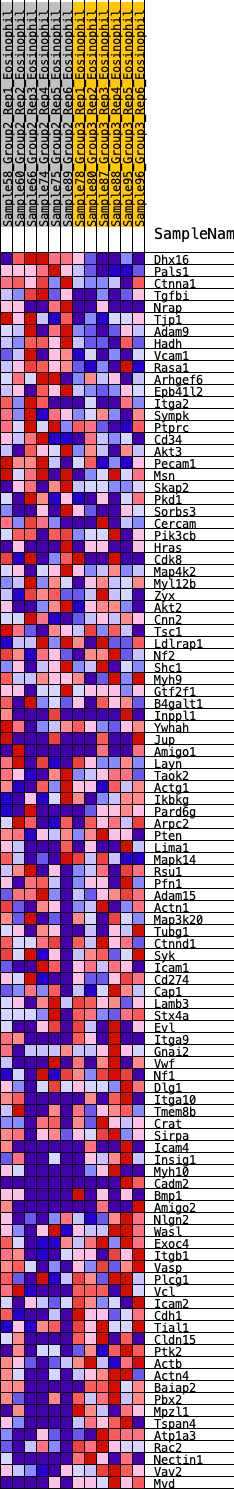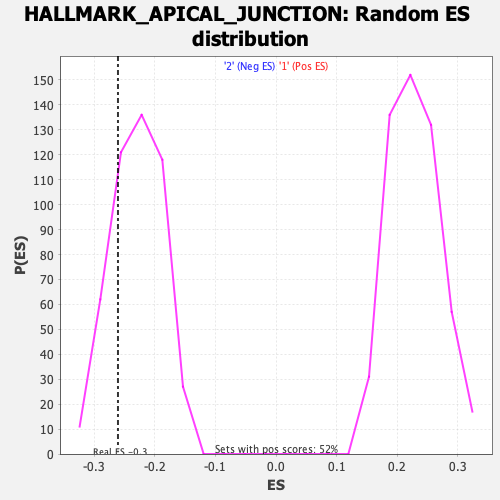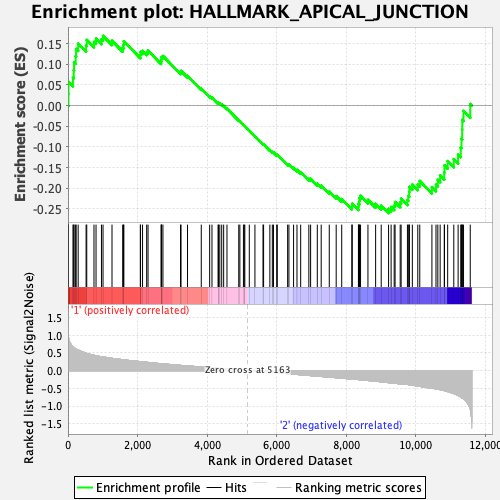
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Eosinophil.Eosinophil_Pheno.cls #Group2_versus_Group3.Eosinophil_Pheno.cls #Group2_versus_Group3_repos |
| Phenotype | Eosinophil_Pheno.cls#Group2_versus_Group3_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_APICAL_JUNCTION |
| Enrichment Score (ES) | -0.26058182 |
| Normalized Enrichment Score (NES) | -1.1395795 |
| Nominal p-value | 0.25473684 |
| FDR q-value | 0.6646725 |
| FWER p-Value | 0.982 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Dhx16 | 13 | 0.960 | 0.0285 | No |
| 2 | Pals1 | 15 | 0.950 | 0.0577 | No |
| 3 | Ctnna1 | 143 | 0.676 | 0.0675 | No |
| 4 | Tgfbi | 165 | 0.653 | 0.0858 | No |
| 5 | Nrap | 174 | 0.640 | 0.1049 | No |
| 6 | Tjp1 | 220 | 0.610 | 0.1198 | No |
| 7 | Adam9 | 233 | 0.604 | 0.1374 | No |
| 8 | Hadh | 292 | 0.579 | 0.1502 | No |
| 9 | Vcam1 | 520 | 0.493 | 0.1457 | No |
| 10 | Rasa1 | 539 | 0.485 | 0.1591 | No |
| 11 | Arhgef6 | 747 | 0.433 | 0.1545 | No |
| 12 | Epb41l2 | 806 | 0.418 | 0.1623 | No |
| 13 | Itga2 | 962 | 0.390 | 0.1609 | No |
| 14 | Sympk | 1007 | 0.385 | 0.1690 | No |
| 15 | Ptprc | 1265 | 0.344 | 0.1573 | No |
| 16 | Cd34 | 1572 | 0.305 | 0.1401 | No |
| 17 | Akt3 | 1597 | 0.302 | 0.1473 | No |
| 18 | Pecam1 | 1605 | 0.302 | 0.1560 | No |
| 19 | Msn | 2079 | 0.253 | 0.1227 | No |
| 20 | Skap2 | 2085 | 0.251 | 0.1300 | No |
| 21 | Pkd1 | 2143 | 0.246 | 0.1326 | No |
| 22 | Sorbs3 | 2262 | 0.233 | 0.1296 | No |
| 23 | Cercam | 2301 | 0.230 | 0.1334 | No |
| 24 | Pik3cb | 2676 | 0.193 | 0.1068 | No |
| 25 | Hras | 2685 | 0.193 | 0.1121 | No |
| 26 | Cdk8 | 2690 | 0.193 | 0.1177 | No |
| 27 | Map4k2 | 2729 | 0.190 | 0.1202 | No |
| 28 | Myl12b | 3235 | 0.146 | 0.0809 | No |
| 29 | Zyx | 3249 | 0.145 | 0.0842 | No |
| 30 | Akt2 | 3436 | 0.130 | 0.0721 | No |
| 31 | Cnn2 | 3831 | 0.101 | 0.0409 | No |
| 32 | Tsc1 | 4072 | 0.083 | 0.0227 | No |
| 33 | Ldlrap1 | 4139 | 0.078 | 0.0193 | No |
| 34 | Nf2 | 4314 | 0.063 | 0.0061 | No |
| 35 | Shc1 | 4328 | 0.062 | 0.0069 | No |
| 36 | Myh9 | 4364 | 0.059 | 0.0057 | No |
| 37 | Gtf2f1 | 4419 | 0.055 | 0.0027 | No |
| 38 | B4galt1 | 4475 | 0.052 | -0.0005 | No |
| 39 | Inppl1 | 4570 | 0.044 | -0.0073 | No |
| 40 | Ywhah | 4909 | 0.018 | -0.0361 | No |
| 41 | Jup | 4941 | 0.015 | -0.0384 | No |
| 42 | Amigo1 | 5043 | 0.008 | -0.0469 | No |
| 43 | Layn | 5065 | 0.007 | -0.0485 | No |
| 44 | Taok2 | 5083 | 0.005 | -0.0498 | No |
| 45 | Actg1 | 5212 | -0.003 | -0.0609 | No |
| 46 | Ikbkg | 5374 | -0.015 | -0.0744 | No |
| 47 | Pard6g | 5606 | -0.033 | -0.0934 | No |
| 48 | Arpc2 | 5623 | -0.035 | -0.0937 | No |
| 49 | Pten | 5803 | -0.048 | -0.1078 | No |
| 50 | Lima1 | 5874 | -0.053 | -0.1123 | No |
| 51 | Mapk14 | 5907 | -0.055 | -0.1134 | No |
| 52 | Rsu1 | 6001 | -0.062 | -0.1195 | No |
| 53 | Pfn1 | 6016 | -0.063 | -0.1188 | No |
| 54 | Adam15 | 6314 | -0.086 | -0.1420 | No |
| 55 | Actn1 | 6351 | -0.090 | -0.1423 | No |
| 56 | Map3k20 | 6485 | -0.101 | -0.1508 | No |
| 57 | Tubg1 | 6583 | -0.109 | -0.1558 | No |
| 58 | Ctnnd1 | 6689 | -0.118 | -0.1613 | No |
| 59 | Syk | 6924 | -0.136 | -0.1775 | No |
| 60 | Icam1 | 6975 | -0.141 | -0.1775 | No |
| 61 | Cd274 | 7167 | -0.156 | -0.1892 | No |
| 62 | Cap1 | 7279 | -0.165 | -0.1938 | No |
| 63 | Lamb3 | 7511 | -0.183 | -0.2082 | No |
| 64 | Stx4a | 7710 | -0.201 | -0.2192 | No |
| 65 | Evl | 7869 | -0.215 | -0.2263 | No |
| 66 | Itga9 | 8160 | -0.239 | -0.2442 | No |
| 67 | Gnai2 | 8171 | -0.239 | -0.2377 | No |
| 68 | Vwf | 8349 | -0.253 | -0.2452 | No |
| 69 | Nf1 | 8352 | -0.253 | -0.2376 | No |
| 70 | Dlg1 | 8376 | -0.257 | -0.2317 | No |
| 71 | Itga10 | 8380 | -0.257 | -0.2240 | No |
| 72 | Tmem8b | 8410 | -0.260 | -0.2185 | No |
| 73 | Crat | 8623 | -0.279 | -0.2283 | No |
| 74 | Sirpa | 8844 | -0.300 | -0.2382 | No |
| 75 | Icam4 | 9004 | -0.316 | -0.2422 | No |
| 76 | Insig1 | 9216 | -0.340 | -0.2501 | Yes |
| 77 | Myh10 | 9289 | -0.345 | -0.2457 | Yes |
| 78 | Cadm2 | 9377 | -0.352 | -0.2424 | Yes |
| 79 | Bmp1 | 9403 | -0.356 | -0.2336 | Yes |
| 80 | Amigo2 | 9550 | -0.374 | -0.2347 | Yes |
| 81 | Nlgn2 | 9577 | -0.375 | -0.2254 | Yes |
| 82 | Wasl | 9758 | -0.391 | -0.2290 | Yes |
| 83 | Exoc4 | 9782 | -0.395 | -0.2188 | Yes |
| 84 | Itgb1 | 9810 | -0.400 | -0.2088 | Yes |
| 85 | Vasp | 9814 | -0.400 | -0.1967 | Yes |
| 86 | Plcg1 | 9899 | -0.413 | -0.1913 | Yes |
| 87 | Vcl | 10056 | -0.436 | -0.1914 | Yes |
| 88 | Icam2 | 10113 | -0.445 | -0.1826 | Yes |
| 89 | Cdh1 | 10462 | -0.498 | -0.1974 | Yes |
| 90 | Tial1 | 10579 | -0.516 | -0.1916 | Yes |
| 91 | Cldn15 | 10632 | -0.524 | -0.1800 | Yes |
| 92 | Ptk2 | 10699 | -0.538 | -0.1691 | Yes |
| 93 | Actb | 10817 | -0.565 | -0.1618 | Yes |
| 94 | Actn4 | 10824 | -0.567 | -0.1449 | Yes |
| 95 | Baiap2 | 10915 | -0.591 | -0.1344 | Yes |
| 96 | Pbx2 | 11091 | -0.649 | -0.1296 | Yes |
| 97 | Mpzl1 | 11215 | -0.704 | -0.1186 | Yes |
| 98 | Tspan4 | 11291 | -0.747 | -0.1020 | Yes |
| 99 | Atp1a3 | 11312 | -0.757 | -0.0804 | Yes |
| 100 | Rac2 | 11333 | -0.773 | -0.0583 | Yes |
| 101 | Nectin1 | 11335 | -0.774 | -0.0345 | Yes |
| 102 | Vav2 | 11371 | -0.797 | -0.0130 | Yes |
| 103 | Mvd | 11564 | -1.088 | 0.0039 | Yes |