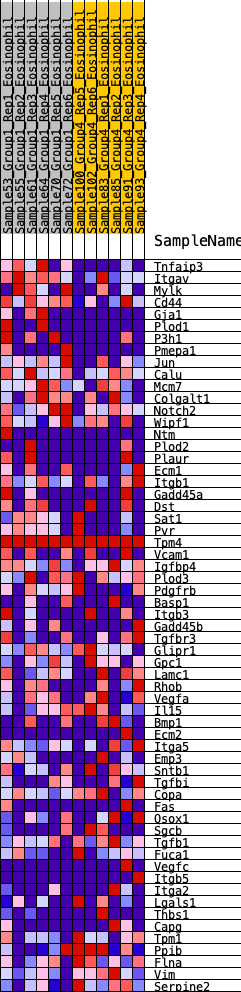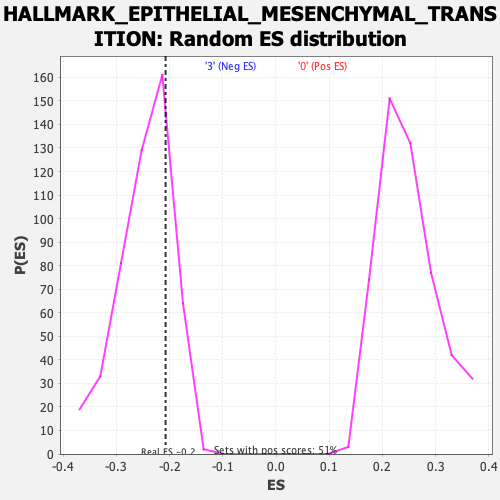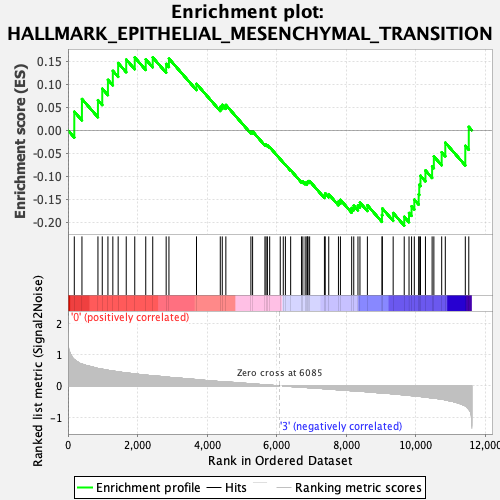
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Eosinophil.Eosinophil_Pheno.cls #Group1_versus_Group4.Eosinophil_Pheno.cls #Group1_versus_Group4_repos |
| Phenotype | Eosinophil_Pheno.cls#Group1_versus_Group4_repos |
| Upregulated in class | 3 |
| GeneSet | HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION |
| Enrichment Score (ES) | -0.20748796 |
| Normalized Enrichment Score (NES) | -0.846299 |
| Nominal p-value | 0.77096117 |
| FDR q-value | 0.85869586 |
| FWER p-Value | 1.0 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Tnfaip3 | 181 | 0.839 | 0.0405 | No |
| 2 | Itgav | 401 | 0.696 | 0.0681 | No |
| 3 | Mylk | 861 | 0.556 | 0.0656 | No |
| 4 | Cd44 | 985 | 0.532 | 0.0906 | No |
| 5 | Gja1 | 1148 | 0.497 | 0.1099 | No |
| 6 | Plod1 | 1289 | 0.473 | 0.1295 | No |
| 7 | P3h1 | 1442 | 0.451 | 0.1465 | No |
| 8 | Pmepa1 | 1674 | 0.412 | 0.1541 | No |
| 9 | Jun | 1919 | 0.383 | 0.1586 | No |
| 10 | Calu | 2235 | 0.344 | 0.1544 | No |
| 11 | Mcm7 | 2436 | 0.323 | 0.1587 | No |
| 12 | Colgalt1 | 2822 | 0.286 | 0.1445 | No |
| 13 | Notch2 | 2902 | 0.275 | 0.1561 | No |
| 14 | Wipf1 | 3694 | 0.202 | 0.1011 | No |
| 15 | Ntm | 4378 | 0.138 | 0.0512 | No |
| 16 | Plod2 | 4435 | 0.137 | 0.0555 | No |
| 17 | Plaur | 4538 | 0.128 | 0.0553 | No |
| 18 | Ecm1 | 5258 | 0.068 | -0.0025 | No |
| 19 | Itgb1 | 5307 | 0.062 | -0.0024 | No |
| 20 | Gadd45a | 5662 | 0.034 | -0.0308 | No |
| 21 | Dst | 5695 | 0.032 | -0.0315 | No |
| 22 | Sat1 | 5733 | 0.029 | -0.0327 | No |
| 23 | Pvr | 5798 | 0.024 | -0.0366 | No |
| 24 | Tpm4 | 6102 | 0.000 | -0.0629 | No |
| 25 | Vcam1 | 6189 | -0.001 | -0.0702 | No |
| 26 | Igfbp4 | 6250 | -0.006 | -0.0750 | No |
| 27 | Plod3 | 6401 | -0.018 | -0.0868 | No |
| 28 | Pdgfrb | 6714 | -0.038 | -0.1113 | No |
| 29 | Basp1 | 6752 | -0.041 | -0.1117 | No |
| 30 | Itgb3 | 6818 | -0.046 | -0.1143 | No |
| 31 | Gadd45b | 6868 | -0.051 | -0.1151 | No |
| 32 | Tgfbr3 | 6882 | -0.053 | -0.1127 | No |
| 33 | Glipr1 | 6901 | -0.054 | -0.1106 | No |
| 34 | Gpc1 | 6947 | -0.057 | -0.1107 | No |
| 35 | Lamc1 | 7372 | -0.089 | -0.1415 | No |
| 36 | Rhob | 7395 | -0.091 | -0.1373 | No |
| 37 | Vegfa | 7496 | -0.100 | -0.1392 | No |
| 38 | Il15 | 7775 | -0.122 | -0.1551 | No |
| 39 | Bmp1 | 7832 | -0.127 | -0.1514 | No |
| 40 | Ecm2 | 8155 | -0.151 | -0.1692 | No |
| 41 | Itga5 | 8215 | -0.153 | -0.1640 | No |
| 42 | Emp3 | 8336 | -0.161 | -0.1636 | No |
| 43 | Sntb1 | 8393 | -0.167 | -0.1573 | No |
| 44 | Tgfbi | 8607 | -0.187 | -0.1632 | No |
| 45 | Copa | 9026 | -0.224 | -0.1844 | No |
| 46 | Fas | 9037 | -0.225 | -0.1702 | No |
| 47 | Qsox1 | 9348 | -0.254 | -0.1800 | No |
| 48 | Sgcb | 9666 | -0.286 | -0.1883 | Yes |
| 49 | Tgfb1 | 9804 | -0.301 | -0.1800 | Yes |
| 50 | Fuca1 | 9877 | -0.310 | -0.1655 | Yes |
| 51 | Vegfc | 9956 | -0.318 | -0.1510 | Yes |
| 52 | Itgb5 | 10082 | -0.332 | -0.1395 | Yes |
| 53 | Itga2 | 10098 | -0.334 | -0.1185 | Yes |
| 54 | Lgals1 | 10134 | -0.338 | -0.0989 | Yes |
| 55 | Thbs1 | 10276 | -0.355 | -0.0873 | Yes |
| 56 | Capg | 10468 | -0.381 | -0.0784 | Yes |
| 57 | Tpm1 | 10518 | -0.385 | -0.0568 | Yes |
| 58 | Ppib | 10743 | -0.426 | -0.0477 | Yes |
| 59 | Flna | 10847 | -0.446 | -0.0268 | Yes |
| 60 | Vim | 11422 | -0.638 | -0.0338 | Yes |
| 61 | Serpine2 | 11522 | -0.745 | 0.0075 | Yes |