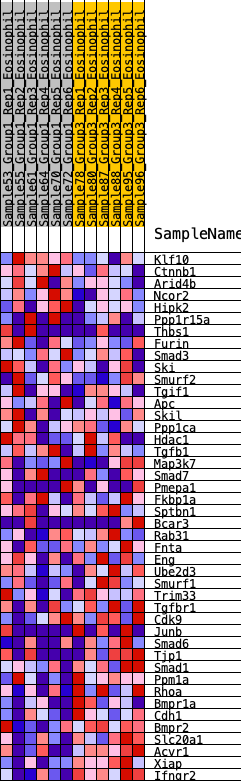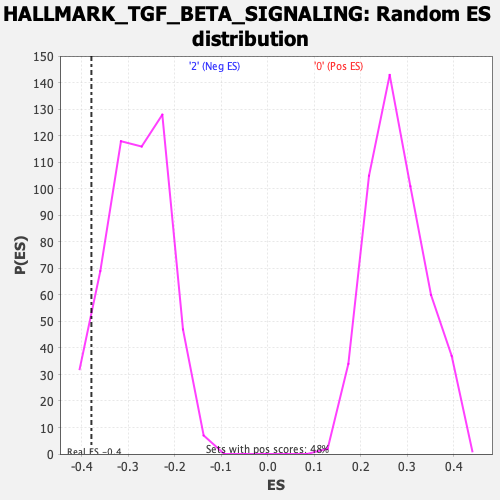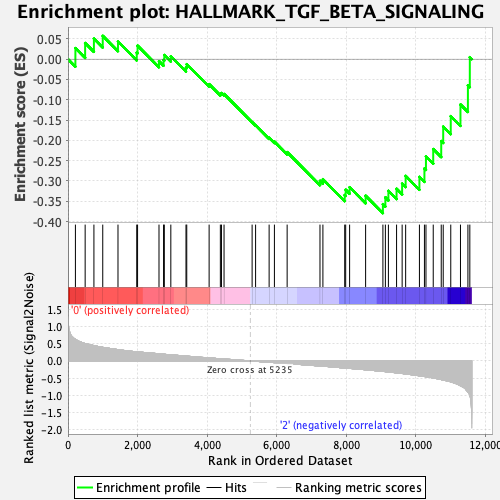
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Eosinophil.Eosinophil_Pheno.cls #Group1_versus_Group3.Eosinophil_Pheno.cls #Group1_versus_Group3_repos |
| Phenotype | Eosinophil_Pheno.cls#Group1_versus_Group3_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_TGF_BETA_SIGNALING |
| Enrichment Score (ES) | -0.3793951 |
| Normalized Enrichment Score (NES) | -1.3612161 |
| Nominal p-value | 0.061895553 |
| FDR q-value | 0.31547794 |
| FWER p-Value | 0.675 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Klf10 | 213 | 0.625 | 0.0273 | No |
| 2 | Ctnnb1 | 493 | 0.502 | 0.0398 | No |
| 3 | Arid4b | 745 | 0.444 | 0.0506 | No |
| 4 | Ncor2 | 999 | 0.393 | 0.0574 | No |
| 5 | Hipk2 | 1437 | 0.326 | 0.0435 | No |
| 6 | Ppp1r15a | 1976 | 0.263 | 0.0162 | No |
| 7 | Thbs1 | 2000 | 0.261 | 0.0333 | No |
| 8 | Furin | 2616 | 0.206 | -0.0048 | No |
| 9 | Smad3 | 2747 | 0.194 | -0.0019 | No |
| 10 | Ski | 2769 | 0.192 | 0.0103 | No |
| 11 | Smurf2 | 2956 | 0.174 | 0.0069 | No |
| 12 | Tgif1 | 3392 | 0.139 | -0.0206 | No |
| 13 | Apc | 3414 | 0.137 | -0.0124 | No |
| 14 | Skil | 4057 | 0.086 | -0.0616 | No |
| 15 | Ppp1ca | 4380 | 0.061 | -0.0850 | No |
| 16 | Hdac1 | 4412 | 0.059 | -0.0834 | No |
| 17 | Tgfb1 | 4486 | 0.054 | -0.0858 | No |
| 18 | Map3k7 | 5293 | -0.004 | -0.1552 | No |
| 19 | Smad7 | 5393 | -0.011 | -0.1630 | No |
| 20 | Pmepa1 | 5782 | -0.039 | -0.1937 | No |
| 21 | Fkbp1a | 5934 | -0.050 | -0.2031 | No |
| 22 | Sptbn1 | 6300 | -0.076 | -0.2291 | No |
| 23 | Bcar3 | 7244 | -0.148 | -0.2998 | No |
| 24 | Rab31 | 7328 | -0.154 | -0.2957 | No |
| 25 | Fnta | 7955 | -0.207 | -0.3347 | No |
| 26 | Eng | 7983 | -0.209 | -0.3218 | No |
| 27 | Ube2d3 | 8096 | -0.218 | -0.3155 | No |
| 28 | Smurf1 | 8556 | -0.256 | -0.3365 | No |
| 29 | Trim33 | 9053 | -0.303 | -0.3572 | Yes |
| 30 | Tgfbr1 | 9124 | -0.311 | -0.3405 | Yes |
| 31 | Cdk9 | 9211 | -0.321 | -0.3245 | Yes |
| 32 | Junb | 9446 | -0.347 | -0.3194 | Yes |
| 33 | Smad6 | 9606 | -0.364 | -0.3065 | Yes |
| 34 | Tjp1 | 9707 | -0.376 | -0.2877 | Yes |
| 35 | Smad1 | 10103 | -0.434 | -0.2901 | Yes |
| 36 | Ppm1a | 10247 | -0.453 | -0.2694 | Yes |
| 37 | Rhoa | 10291 | -0.459 | -0.2396 | Yes |
| 38 | Bmpr1a | 10500 | -0.493 | -0.2215 | Yes |
| 39 | Cdh1 | 10730 | -0.541 | -0.2018 | Yes |
| 40 | Bmpr2 | 10787 | -0.555 | -0.1661 | Yes |
| 41 | Slc20a1 | 11004 | -0.607 | -0.1404 | Yes |
| 42 | Acvr1 | 11283 | -0.721 | -0.1117 | Yes |
| 43 | Xiap | 11497 | -0.900 | -0.0644 | Yes |
| 44 | Ifngr2 | 11552 | -1.012 | 0.0049 | Yes |