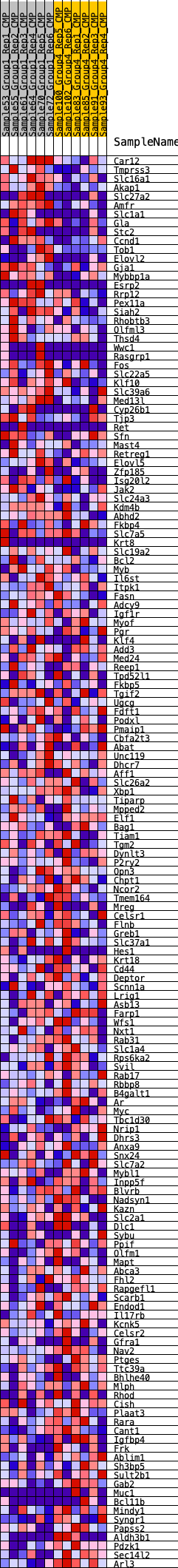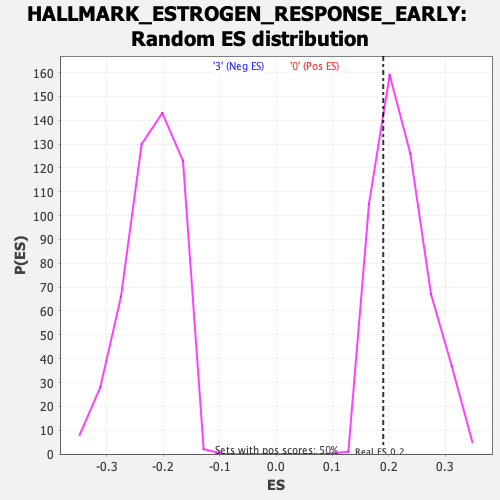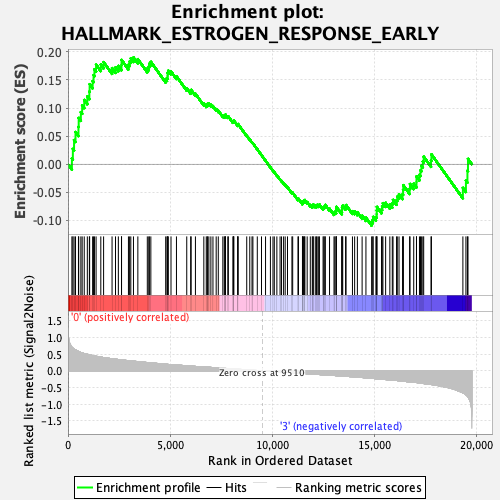
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | CMP.CMP_Pheno.cls#Group1_versus_Group4.CMP_Pheno.cls#Group1_versus_Group4_repos |
| Phenotype | CMP_Pheno.cls#Group1_versus_Group4_repos |
| Upregulated in class | 0 |
| GeneSet | HALLMARK_ESTROGEN_RESPONSE_EARLY |
| Enrichment Score (ES) | 0.19018243 |
| Normalized Enrichment Score (NES) | 0.85438514 |
| Nominal p-value | 0.71 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.999 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Car12 | 191 | 0.706 | 0.0104 | Yes |
| 2 | Tmprss3 | 238 | 0.681 | 0.0274 | Yes |
| 3 | Slc16a1 | 297 | 0.656 | 0.0431 | Yes |
| 4 | Akap1 | 368 | 0.625 | 0.0574 | Yes |
| 5 | Slc27a2 | 512 | 0.581 | 0.0666 | Yes |
| 6 | Amfr | 524 | 0.577 | 0.0825 | Yes |
| 7 | Slc1a1 | 630 | 0.550 | 0.0928 | Yes |
| 8 | Gla | 693 | 0.537 | 0.1049 | Yes |
| 9 | Stc2 | 795 | 0.517 | 0.1144 | Yes |
| 10 | Ccnd1 | 946 | 0.494 | 0.1209 | Yes |
| 11 | Tob1 | 1046 | 0.479 | 0.1295 | Yes |
| 12 | Elovl2 | 1059 | 0.478 | 0.1424 | Yes |
| 13 | Gja1 | 1206 | 0.457 | 0.1480 | Yes |
| 14 | Mybbp1a | 1251 | 0.451 | 0.1586 | Yes |
| 15 | Esrp2 | 1295 | 0.445 | 0.1691 | Yes |
| 16 | Rrp12 | 1375 | 0.436 | 0.1775 | Yes |
| 17 | Pex11a | 1606 | 0.411 | 0.1774 | Yes |
| 18 | Siah2 | 1743 | 0.397 | 0.1818 | Yes |
| 19 | Rhobtb3 | 2157 | 0.363 | 0.1711 | Yes |
| 20 | Olfml3 | 2326 | 0.350 | 0.1725 | Yes |
| 21 | Thsd4 | 2465 | 0.338 | 0.1751 | Yes |
| 22 | Wwc1 | 2619 | 0.325 | 0.1765 | Yes |
| 23 | Rasgrp1 | 2622 | 0.325 | 0.1857 | Yes |
| 24 | Fos | 2959 | 0.306 | 0.1773 | Yes |
| 25 | Slc22a5 | 3017 | 0.302 | 0.1830 | Yes |
| 26 | Klf10 | 3075 | 0.298 | 0.1885 | Yes |
| 27 | Slc39a6 | 3205 | 0.289 | 0.1902 | Yes |
| 28 | Med13l | 3420 | 0.275 | 0.1871 | No |
| 29 | Cyp26b1 | 3883 | 0.247 | 0.1706 | No |
| 30 | Tjp3 | 3955 | 0.242 | 0.1738 | No |
| 31 | Ret | 3984 | 0.240 | 0.1793 | No |
| 32 | Sfn | 4056 | 0.237 | 0.1824 | No |
| 33 | Mast4 | 4783 | 0.198 | 0.1510 | No |
| 34 | Retreg1 | 4845 | 0.194 | 0.1534 | No |
| 35 | Elovl5 | 4878 | 0.193 | 0.1573 | No |
| 36 | Zfp185 | 4879 | 0.193 | 0.1628 | No |
| 37 | Isg20l2 | 4910 | 0.191 | 0.1667 | No |
| 38 | Jak2 | 5043 | 0.184 | 0.1652 | No |
| 39 | Slc24a3 | 5310 | 0.173 | 0.1566 | No |
| 40 | Kdm4b | 5814 | 0.150 | 0.1352 | No |
| 41 | Abhd2 | 6007 | 0.142 | 0.1294 | No |
| 42 | Fkbp4 | 6028 | 0.141 | 0.1324 | No |
| 43 | Slc7a5 | 6239 | 0.132 | 0.1255 | No |
| 44 | Krt8 | 6650 | 0.114 | 0.1078 | No |
| 45 | Slc19a2 | 6769 | 0.112 | 0.1050 | No |
| 46 | Bcl2 | 6816 | 0.110 | 0.1058 | No |
| 47 | Myb | 6839 | 0.109 | 0.1077 | No |
| 48 | Il6st | 6897 | 0.107 | 0.1079 | No |
| 49 | Itpk1 | 6995 | 0.102 | 0.1059 | No |
| 50 | Fasn | 7097 | 0.098 | 0.1035 | No |
| 51 | Adcy9 | 7255 | 0.092 | 0.0981 | No |
| 52 | Igf1r | 7358 | 0.087 | 0.0954 | No |
| 53 | Myof | 7581 | 0.078 | 0.0863 | No |
| 54 | Pgr | 7651 | 0.076 | 0.0849 | No |
| 55 | Klf4 | 7685 | 0.073 | 0.0853 | No |
| 56 | Add3 | 7703 | 0.073 | 0.0865 | No |
| 57 | Med24 | 7707 | 0.073 | 0.0884 | No |
| 58 | Reep1 | 7826 | 0.067 | 0.0843 | No |
| 59 | Tpd52l1 | 7850 | 0.066 | 0.0850 | No |
| 60 | Fkbp5 | 8073 | 0.057 | 0.0753 | No |
| 61 | Tgif2 | 8101 | 0.056 | 0.0755 | No |
| 62 | Ugcg | 8107 | 0.055 | 0.0769 | No |
| 63 | Fdft1 | 8119 | 0.055 | 0.0779 | No |
| 64 | Podxl | 8312 | 0.047 | 0.0694 | No |
| 65 | Pmaip1 | 8316 | 0.047 | 0.0706 | No |
| 66 | Cbfa2t3 | 8323 | 0.047 | 0.0716 | No |
| 67 | Abat | 8751 | 0.030 | 0.0507 | No |
| 68 | Unc119 | 8900 | 0.024 | 0.0439 | No |
| 69 | Dhcr7 | 9009 | 0.021 | 0.0389 | No |
| 70 | Aff1 | 9053 | 0.018 | 0.0373 | No |
| 71 | Slc26a2 | 9268 | 0.009 | 0.0266 | No |
| 72 | Xbp1 | 9474 | 0.002 | 0.0162 | No |
| 73 | Tiparp | 9672 | -0.002 | 0.0062 | No |
| 74 | Mpped2 | 9910 | -0.011 | -0.0055 | No |
| 75 | Elf1 | 10036 | -0.016 | -0.0115 | No |
| 76 | Bag1 | 10101 | -0.018 | -0.0142 | No |
| 77 | Tiam1 | 10226 | -0.023 | -0.0199 | No |
| 78 | Tgm2 | 10383 | -0.029 | -0.0270 | No |
| 79 | Dynlt3 | 10455 | -0.032 | -0.0297 | No |
| 80 | P2ry2 | 10556 | -0.036 | -0.0338 | No |
| 81 | Opn3 | 10634 | -0.039 | -0.0366 | No |
| 82 | Chpt1 | 10747 | -0.044 | -0.0411 | No |
| 83 | Ncor2 | 10964 | -0.053 | -0.0506 | No |
| 84 | Tmem164 | 11001 | -0.054 | -0.0509 | No |
| 85 | Mreg | 11260 | -0.065 | -0.0622 | No |
| 86 | Celsr1 | 11284 | -0.066 | -0.0615 | No |
| 87 | Flnb | 11479 | -0.074 | -0.0693 | No |
| 88 | Greb1 | 11487 | -0.075 | -0.0675 | No |
| 89 | Slc37a1 | 11503 | -0.076 | -0.0661 | No |
| 90 | Hes1 | 11534 | -0.077 | -0.0655 | No |
| 91 | Krt18 | 11589 | -0.079 | -0.0659 | No |
| 92 | Cd44 | 11591 | -0.080 | -0.0637 | No |
| 93 | Deptor | 11710 | -0.084 | -0.0674 | No |
| 94 | Scnn1a | 11863 | -0.091 | -0.0725 | No |
| 95 | Lrig1 | 11960 | -0.095 | -0.0747 | No |
| 96 | Asb13 | 11992 | -0.096 | -0.0736 | No |
| 97 | Farp1 | 12009 | -0.097 | -0.0716 | No |
| 98 | Wfs1 | 12107 | -0.100 | -0.0737 | No |
| 99 | Nxt1 | 12170 | -0.103 | -0.0739 | No |
| 100 | Rab31 | 12197 | -0.105 | -0.0723 | No |
| 101 | Slc1a4 | 12273 | -0.108 | -0.0730 | No |
| 102 | Rps6ka2 | 12310 | -0.110 | -0.0717 | No |
| 103 | Svil | 12489 | -0.119 | -0.0774 | No |
| 104 | Rab17 | 12527 | -0.121 | -0.0759 | No |
| 105 | Rbbp8 | 12573 | -0.122 | -0.0747 | No |
| 106 | B4galt1 | 12598 | -0.123 | -0.0724 | No |
| 107 | Ar | 12805 | -0.128 | -0.0793 | No |
| 108 | Myc | 13024 | -0.139 | -0.0864 | No |
| 109 | Tbc1d30 | 13067 | -0.140 | -0.0846 | No |
| 110 | Nrip1 | 13129 | -0.143 | -0.0836 | No |
| 111 | Dhrs3 | 13134 | -0.143 | -0.0797 | No |
| 112 | Anxa9 | 13139 | -0.144 | -0.0758 | No |
| 113 | Snx24 | 13407 | -0.155 | -0.0850 | No |
| 114 | Slc7a2 | 13414 | -0.155 | -0.0809 | No |
| 115 | Mybl1 | 13419 | -0.155 | -0.0767 | No |
| 116 | Inpp5f | 13451 | -0.157 | -0.0738 | No |
| 117 | Blvrb | 13591 | -0.163 | -0.0763 | No |
| 118 | Nadsyn1 | 13613 | -0.164 | -0.0727 | No |
| 119 | Kazn | 13927 | -0.174 | -0.0837 | No |
| 120 | Slc2a1 | 14034 | -0.179 | -0.0840 | No |
| 121 | Dlc1 | 14168 | -0.186 | -0.0855 | No |
| 122 | Sybu | 14400 | -0.198 | -0.0916 | No |
| 123 | Ppif | 14587 | -0.209 | -0.0951 | No |
| 124 | Olfm1 | 14866 | -0.223 | -0.1030 | No |
| 125 | Mapt | 14920 | -0.226 | -0.0992 | No |
| 126 | Abca3 | 14940 | -0.227 | -0.0937 | No |
| 127 | Fhl2 | 15076 | -0.235 | -0.0939 | No |
| 128 | Rapgefl1 | 15113 | -0.237 | -0.0890 | No |
| 129 | Scarb1 | 15115 | -0.238 | -0.0823 | No |
| 130 | Endod1 | 15124 | -0.238 | -0.0759 | No |
| 131 | Il17rb | 15351 | -0.249 | -0.0804 | No |
| 132 | Kcnk5 | 15380 | -0.251 | -0.0747 | No |
| 133 | Celsr2 | 15418 | -0.253 | -0.0693 | No |
| 134 | Gfra1 | 15552 | -0.261 | -0.0687 | No |
| 135 | Nav2 | 15758 | -0.270 | -0.0715 | No |
| 136 | Ptges | 15877 | -0.276 | -0.0696 | No |
| 137 | Ttc39a | 15913 | -0.278 | -0.0635 | No |
| 138 | Bhlhe40 | 16086 | -0.288 | -0.0641 | No |
| 139 | Mlph | 16122 | -0.291 | -0.0576 | No |
| 140 | Rhod | 16207 | -0.296 | -0.0534 | No |
| 141 | Cish | 16381 | -0.309 | -0.0534 | No |
| 142 | Plaat3 | 16407 | -0.312 | -0.0458 | No |
| 143 | Rara | 16416 | -0.312 | -0.0373 | No |
| 144 | Cant1 | 16727 | -0.330 | -0.0438 | No |
| 145 | Igfbp4 | 16744 | -0.330 | -0.0352 | No |
| 146 | Frk | 16927 | -0.341 | -0.0347 | No |
| 147 | Ablim1 | 17062 | -0.349 | -0.0316 | No |
| 148 | Sh3bp5 | 17063 | -0.349 | -0.0217 | No |
| 149 | Sult2b1 | 17208 | -0.362 | -0.0187 | No |
| 150 | Gab2 | 17259 | -0.367 | -0.0108 | No |
| 151 | Muc1 | 17303 | -0.371 | -0.0024 | No |
| 152 | Bcl11b | 17373 | -0.376 | 0.0048 | No |
| 153 | Mindy1 | 17415 | -0.380 | 0.0135 | No |
| 154 | Syngr1 | 17774 | -0.409 | 0.0069 | No |
| 155 | Papss2 | 17794 | -0.411 | 0.0176 | No |
| 156 | Aldh3b1 | 19334 | -0.661 | -0.0421 | No |
| 157 | Pdzk1 | 19483 | -0.717 | -0.0292 | No |
| 158 | Sec14l2 | 19557 | -0.757 | -0.0114 | No |
| 159 | Arl3 | 19584 | -0.781 | 0.0095 | No |