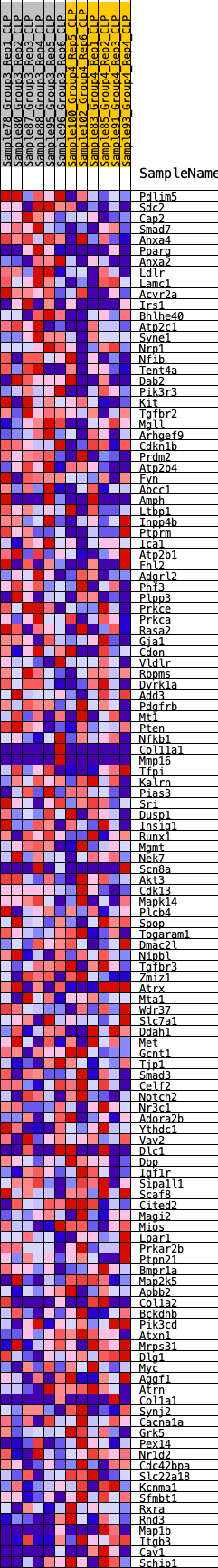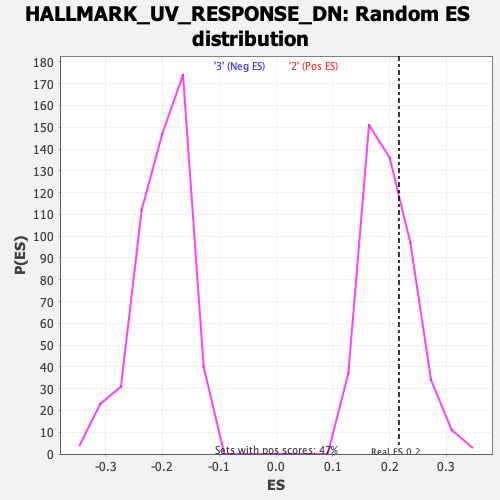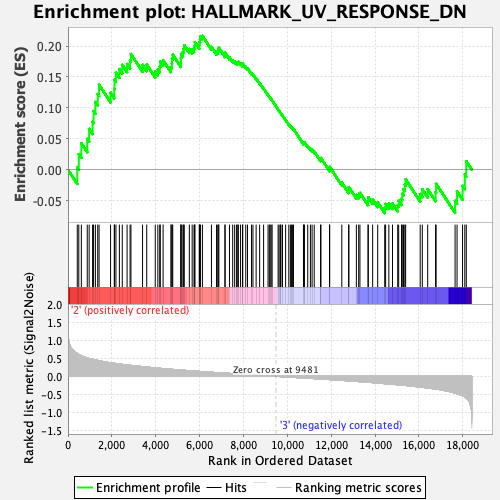
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | CLP.Basophil_Pheno.cls #Group3_versus_Group4.Basophil_Pheno.cls #Group3_versus_Group4_repos |
| Phenotype | Basophil_Pheno.cls#Group3_versus_Group4_repos |
| Upregulated in class | 2 |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | 0.21640736 |
| Normalized Enrichment Score (NES) | 1.0848067 |
| Nominal p-value | 0.3304904 |
| FDR q-value | 0.833239 |
| FWER p-Value | 0.972 |

| SYMBOL | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|
| 1 | Pdlim5 | 423 | 0.634 | 0.0036 | Yes |
| 2 | Sdc2 | 492 | 0.604 | 0.0253 | Yes |
| 3 | Cap2 | 608 | 0.570 | 0.0430 | Yes |
| 4 | Smad7 | 877 | 0.514 | 0.0499 | Yes |
| 5 | Anxa4 | 964 | 0.495 | 0.0661 | Yes |
| 6 | Pparg | 1121 | 0.475 | 0.0775 | Yes |
| 7 | Anxa2 | 1166 | 0.468 | 0.0948 | Yes |
| 8 | Ldlr | 1248 | 0.454 | 0.1095 | Yes |
| 9 | Lamc1 | 1355 | 0.443 | 0.1224 | Yes |
| 10 | Acvr2a | 1416 | 0.436 | 0.1374 | Yes |
| 11 | Irs1 | 1941 | 0.376 | 0.1246 | Yes |
| 12 | Bhlhe40 | 2110 | 0.363 | 0.1307 | Yes |
| 13 | Atp2c1 | 2119 | 0.362 | 0.1455 | Yes |
| 14 | Syne1 | 2185 | 0.355 | 0.1569 | Yes |
| 15 | Nrp1 | 2342 | 0.340 | 0.1627 | Yes |
| 16 | Nfib | 2474 | 0.333 | 0.1696 | Yes |
| 17 | Tent4a | 2693 | 0.316 | 0.1709 | Yes |
| 18 | Dab2 | 2830 | 0.307 | 0.1764 | Yes |
| 19 | Pik3r3 | 2869 | 0.305 | 0.1871 | Yes |
| 20 | Kit | 3399 | 0.270 | 0.1696 | Yes |
| 21 | Tgfbr2 | 3589 | 0.258 | 0.1701 | Yes |
| 22 | Mgll | 3977 | 0.236 | 0.1589 | Yes |
| 23 | Arhgef9 | 4096 | 0.229 | 0.1621 | Yes |
| 24 | Cdkn1b | 4183 | 0.225 | 0.1668 | Yes |
| 25 | Prdm2 | 4206 | 0.223 | 0.1750 | Yes |
| 26 | Atp2b4 | 4335 | 0.215 | 0.1771 | Yes |
| 27 | Fyn | 4689 | 0.198 | 0.1661 | Yes |
| 28 | Abcc1 | 4735 | 0.195 | 0.1718 | Yes |
| 29 | Amph | 4742 | 0.195 | 0.1797 | Yes |
| 30 | Ltbp1 | 4775 | 0.193 | 0.1861 | Yes |
| 31 | Inpp4b | 5144 | 0.176 | 0.1733 | Yes |
| 32 | Ptprm | 5149 | 0.175 | 0.1805 | Yes |
| 33 | Ica1 | 5159 | 0.175 | 0.1874 | Yes |
| 34 | Atp2b1 | 5237 | 0.171 | 0.1904 | Yes |
| 35 | Fhl2 | 5271 | 0.169 | 0.1957 | Yes |
| 36 | Adgrl2 | 5299 | 0.168 | 0.2013 | Yes |
| 37 | Phf3 | 5531 | 0.157 | 0.1952 | Yes |
| 38 | Plpp3 | 5660 | 0.151 | 0.1946 | Yes |
| 39 | Prkce | 5744 | 0.147 | 0.1962 | Yes |
| 40 | Prkca | 5775 | 0.146 | 0.2007 | Yes |
| 41 | Rasa2 | 5784 | 0.146 | 0.2064 | Yes |
| 42 | Gja1 | 5983 | 0.137 | 0.2014 | Yes |
| 43 | Cdon | 5993 | 0.137 | 0.2067 | Yes |
| 44 | Vldlr | 6020 | 0.136 | 0.2109 | Yes |
| 45 | Rbpms | 6034 | 0.135 | 0.2159 | Yes |
| 46 | Dyrk1a | 6127 | 0.131 | 0.2164 | Yes |
| 47 | Add3 | 6539 | 0.113 | 0.1987 | No |
| 48 | Pdgfrb | 6771 | 0.102 | 0.1904 | No |
| 49 | Mt1 | 6832 | 0.100 | 0.1913 | No |
| 50 | Pten | 6847 | 0.099 | 0.1947 | No |
| 51 | Nfkb1 | 6878 | 0.098 | 0.1972 | No |
| 52 | Col11a1 | 7157 | 0.087 | 0.1857 | No |
| 53 | Mmp16 | 7158 | 0.087 | 0.1893 | No |
| 54 | Tfpi | 7360 | 0.082 | 0.1818 | No |
| 55 | Kalrn | 7508 | 0.076 | 0.1770 | No |
| 56 | Pias3 | 7598 | 0.073 | 0.1752 | No |
| 57 | Sri | 7685 | 0.069 | 0.1734 | No |
| 58 | Dusp1 | 7739 | 0.067 | 0.1733 | No |
| 59 | Insig1 | 7764 | 0.066 | 0.1747 | No |
| 60 | Runx1 | 7866 | 0.062 | 0.1719 | No |
| 61 | Mgmt | 7956 | 0.060 | 0.1695 | No |
| 62 | Nek7 | 7966 | 0.059 | 0.1715 | No |
| 63 | Scn8a | 8103 | 0.054 | 0.1663 | No |
| 64 | Akt3 | 8184 | 0.050 | 0.1641 | No |
| 65 | Cdk13 | 8364 | 0.042 | 0.1561 | No |
| 66 | Mapk14 | 8431 | 0.039 | 0.1541 | No |
| 67 | Plcb4 | 8576 | 0.033 | 0.1476 | No |
| 68 | Spop | 8736 | 0.027 | 0.1400 | No |
| 69 | Togaram1 | 8917 | 0.020 | 0.1310 | No |
| 70 | Dmac2l | 9119 | 0.012 | 0.1206 | No |
| 71 | Nipbl | 9176 | 0.010 | 0.1179 | No |
| 72 | Tgfbr3 | 9215 | 0.009 | 0.1162 | No |
| 73 | Zmiz1 | 9260 | 0.008 | 0.1142 | No |
| 74 | Atrx | 9305 | 0.006 | 0.1120 | No |
| 75 | Mta1 | 9578 | -0.002 | 0.0972 | No |
| 76 | Wdr37 | 9626 | -0.003 | 0.0948 | No |
| 77 | Slc7a1 | 9691 | -0.006 | 0.0915 | No |
| 78 | Ddah1 | 9705 | -0.006 | 0.0911 | No |
| 79 | Met | 9777 | -0.009 | 0.0876 | No |
| 80 | Gcnt1 | 9921 | -0.014 | 0.0803 | No |
| 81 | Tjp1 | 10066 | -0.019 | 0.0733 | No |
| 82 | Smad3 | 10138 | -0.021 | 0.0703 | No |
| 83 | Celf2 | 10185 | -0.023 | 0.0687 | No |
| 84 | Notch2 | 10230 | -0.024 | 0.0674 | No |
| 85 | Nr3c1 | 10278 | -0.027 | 0.0659 | No |
| 86 | Adora2b | 10733 | -0.043 | 0.0429 | No |
| 87 | Ythdc1 | 10760 | -0.045 | 0.0434 | No |
| 88 | Vav2 | 10776 | -0.045 | 0.0445 | No |
| 89 | Dlc1 | 10929 | -0.051 | 0.0383 | No |
| 90 | Dbp | 11050 | -0.054 | 0.0340 | No |
| 91 | Igf1r | 11127 | -0.057 | 0.0322 | No |
| 92 | Sipa1l1 | 11224 | -0.061 | 0.0295 | No |
| 93 | Scaf8 | 11513 | -0.072 | 0.0168 | No |
| 94 | Cited2 | 11530 | -0.072 | 0.0190 | No |
| 95 | Magi2 | 11924 | -0.087 | 0.0012 | No |
| 96 | Mios | 11928 | -0.088 | 0.0047 | No |
| 97 | Lpar1 | 12482 | -0.110 | -0.0209 | No |
| 98 | Prkar2b | 12794 | -0.123 | -0.0328 | No |
| 99 | Ptpn21 | 12804 | -0.123 | -0.0281 | No |
| 100 | Bmpr1a | 13145 | -0.139 | -0.0408 | No |
| 101 | Map2k5 | 13251 | -0.143 | -0.0406 | No |
| 102 | Apbb2 | 13308 | -0.146 | -0.0375 | No |
| 103 | Col1a2 | 13676 | -0.163 | -0.0507 | No |
| 104 | Bckdhb | 13689 | -0.164 | -0.0445 | No |
| 105 | Pik3cd | 13888 | -0.173 | -0.0480 | No |
| 106 | Atxn1 | 14118 | -0.184 | -0.0528 | No |
| 107 | Mrps31 | 14438 | -0.201 | -0.0618 | No |
| 108 | Dlg1 | 14479 | -0.203 | -0.0555 | No |
| 109 | Myc | 14626 | -0.209 | -0.0547 | No |
| 110 | Aggf1 | 14792 | -0.218 | -0.0545 | No |
| 111 | Atrn | 15030 | -0.232 | -0.0577 | No |
| 112 | Col1a1 | 15074 | -0.235 | -0.0502 | No |
| 113 | Synj2 | 15204 | -0.240 | -0.0471 | No |
| 114 | Cacna1a | 15238 | -0.242 | -0.0388 | No |
| 115 | Grk5 | 15295 | -0.246 | -0.0315 | No |
| 116 | Pex14 | 15354 | -0.250 | -0.0241 | No |
| 117 | Nr1d2 | 15392 | -0.252 | -0.0155 | No |
| 118 | Cdc42bpa | 16055 | -0.297 | -0.0392 | No |
| 119 | Slc22a18 | 16151 | -0.304 | -0.0316 | No |
| 120 | Kcnma1 | 16397 | -0.321 | -0.0315 | No |
| 121 | Sfmbt1 | 16757 | -0.352 | -0.0363 | No |
| 122 | Rxra | 16780 | -0.353 | -0.0227 | No |
| 123 | Rnd3 | 17645 | -0.466 | -0.0503 | No |
| 124 | Map1b | 17735 | -0.481 | -0.0350 | No |
| 125 | Itgb3 | 17986 | -0.534 | -0.0262 | No |
| 126 | Cav1 | 18089 | -0.569 | -0.0078 | No |
| 127 | Schip1 | 18156 | -0.598 | 0.0138 | No |