![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | B188i_DC_0h_RNA_Seq.bam |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 19880761 |
| Filtered Sequences | 0 |
| Sequence length | 42 |
| %GC | 52 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality

![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores

![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution

![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels

![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GTCTGGAGTCTTGGAAGCTTGACTACCCTACGTTCTCCTACA | 26656 | 0.13407937452696103 | No Hit |
| CTGGAGTCTTGGAAGCTTGACTACCCTACGTTCTCCTACAAA | 26179 | 0.1316800699932965 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
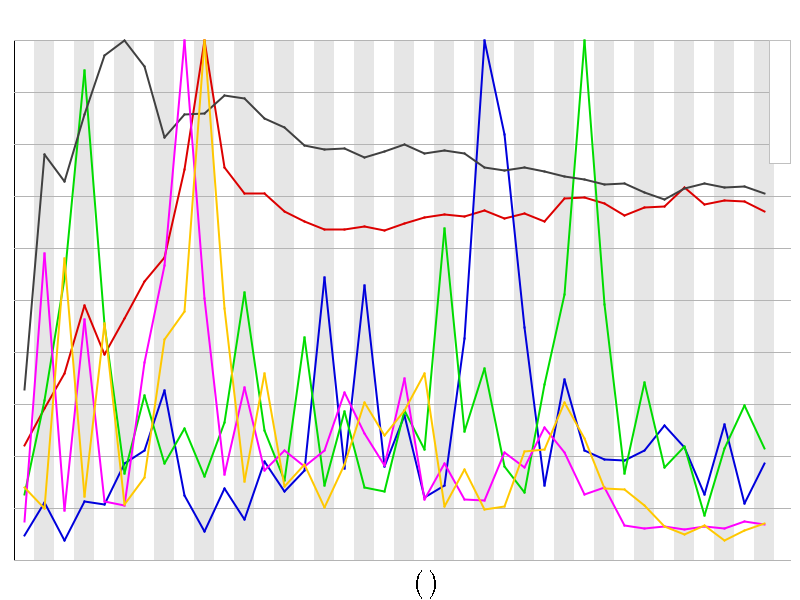
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2857460 | 7.499594 | 11.866362 | 10 |
| TTCAC | 4275375 | 5.2554803 | 21.684734 | 24 |
| CAGGC | 4120820 | 5.230162 | 16.63825 | 29 |
| TTTTT | 4000070 | 4.911603 | 6.2943053 | 6 |
| TGGAG | 2992215 | 4.4871316 | 20.092754 | 9 |
| GGAGT | 2983215 | 4.4736357 | 19.819012 | 10 |
| CACAG | 2927310 | 4.256317 | 19.277456 | 27 |
| ACAGG | 2529445 | 4.0298243 | 20.68014 | 28 |
| ATTCA | 2597710 | 4.0082817 | 19.83852 | 24 |
| CTGGA | 2911055 | 3.984106 | 17.604279 | 8 |
| TGCAG | 2869545 | 3.927295 | 17.842167 | 14 |
| TCACA | 2674205 | 3.8266025 | 18.398003 | 26 |
| CAGTG | 2775015 | 3.7979198 | 17.814089 | 16 |
| GCTAT | 2813090 | 3.7889378 | 16.624855 | 21 |
| AGTGG | 2507615 | 3.7604249 | 19.060099 | 17 |
| CCTCC | 3924350 | 3.563918 | 10.783061 | 13 |
| GAGTG | 2333165 | 3.4988194 | 18.721754 | 11 |
| GCAGT | 2531905 | 3.4651966 | 17.397615 | 15 |
| GTGCA | 2468325 | 3.3781803 | 17.356007 | 13 |
| TGGCT | 2799575 | 3.2914872 | 15.160633 | 19 |
| AGTGC | 2391510 | 3.2730503 | 17.274635 | 12 |
| GCTGG | 2693985 | 3.218414 | 15.417444 | 7 |
| AAAAT | 1413005 | 3.185818 | 5.721339 | 10 |
| TATTC | 2396685 | 3.17686 | 16.710243 | 23 |
| CTATT | 2363355 | 3.1326804 | 16.613277 | 22 |
| GTGGC | 2613710 | 3.1225119 | 15.202794 | 18 |
| GGCTG | 2608800 | 3.1166465 | 15.18452 | 6 |
| GGCTA | 2265335 | 3.1003656 | 16.869228 | 20 |
| AGGCT | 2211995 | 3.027364 | 16.057007 | 5 |
| GTTCA | 2221885 | 2.9926457 | 15.717446 | 23 |
| GCGCG | 2694755 | 2.9855003 | 14.295571 | 32 |
| GGGAG | 1901280 | 2.897135 | 6.0795646 | 19 |
| CCGCC | 3118200 | 2.8774688 | 10.733258 | 10 |
| AGGCG | 2056625 | 2.860107 | 17.537172 | 30 |
| TAAAA | 1242765 | 2.8019881 | 6.568469 | 9 |
| CTCCT | 2827795 | 2.7692158 | 10.848774 | 14 |
| CCGGG | 2493235 | 2.7622375 | 12.851645 | 31 |
| CGGGG | 2217430 | 2.6918035 | 13.149668 | 32 |
| CAGCA | 1824425 | 2.6527193 | 6.15442 | 34 |
| CCAGG | 2079710 | 2.639577 | 11.868018 | 3 |
| GCCGG | 2380395 | 2.6372228 | 11.7265835 | 30 |
| GGGGC | 2117370 | 2.5703373 | 12.999131 | 33 |
| CTCCC | 2817185 | 2.5584407 | 11.805729 | 17 |
| GGCGC | 2301430 | 2.5497382 | 13.987856 | 31 |
| TCACC | 2202995 | 2.5113244 | 13.395106 | 25 |
| TCCCA | 2194755 | 2.5019312 | 14.864011 | 38 |
| GCGAT | 1773660 | 2.4274533 | 16.749268 | 34 |
| TCCTC | 2478800 | 2.4274504 | 10.995356 | 15 |
| CGCGA | 1827755 | 2.3197944 | 15.553069 | 33 |
| GATCC | 1847005 | 2.3070285 | 15.688896 | 36 |
| CCGTT | 2125345 | 2.2805202 | 11.8976965 | 21 |
| GGCGG | 1858290 | 2.2558327 | 12.664156 | 35 |
| TTCTT | 1953495 | 2.2244315 | 5.5098467 | 6 |
| CCCCG | 2403930 | 2.2183416 | 10.015166 | 19 |
| GGGCG | 1820230 | 2.2096305 | 12.615254 | 34 |
| CGATC | 1763585 | 2.2028315 | 15.51094 | 35 |
| ATCCC | 1888755 | 2.1531036 | 14.3862095 | 37 |
| CCCAG | 1825590 | 2.1146536 | 9.253869 | 2 |
| CTCAG | 1690940 | 2.1120932 | 9.312626 | 2 |
| ACTGA | 1338555 | 2.0987003 | 9.374615 | 38 |
| CGTTC | 1949245 | 2.091563 | 11.30939 | 22 |
| ACCGC | 1774115 | 2.055028 | 11.855709 | 27 |
| TCAGG | 1501225 | 2.054595 | 7.635273 | 3 |
| GGTTC | 1709065 | 2.0093641 | 13.337297 | 5 |
| TCCCC | 2211735 | 2.0085983 | 9.767593 | 18 |
| GCCTC | 2007245 | 1.9973581 | 10.782504 | 12 |
| ATCAG | 1255265 | 1.9681113 | 5.628912 | 32 |
| TCTTC | 1858220 | 1.9622526 | 5.1208034 | 7 |
| CCACT | 1717485 | 1.9578629 | 6.9231386 | 33 |
| ATGTT | 1346165 | 1.9551543 | 6.26702 | 7 |
| CCCAC | 1834650 | 1.9395171 | 6.66916 | 38 |
| CACCG | 1661990 | 1.9251492 | 11.77182 | 26 |
| CGCCG | 1900615 | 1.9217478 | 10.680178 | 29 |
| GCGGC | 1710730 | 1.8953059 | 11.318866 | 36 |
| GGCTC | 1738035 | 1.8950021 | 11.32901 | 38 |
| TGTTG | 1442515 | 1.8288131 | 5.3903103 | 8 |
| CCCGT | 1799915 | 1.7910494 | 10.333525 | 20 |
| CCCGC | 1916290 | 1.7683486 | 9.721844 | 9 |
| AGCAC | 1207210 | 1.7552868 | 5.123901 | 35 |
| GATCA | 1107685 | 1.7367228 | 5.3584714 | 31 |
| ACTAC | 1211485 | 1.7335513 | 8.038903 | 35 |
| CACTA | 1179080 | 1.6871818 | 8.031555 | 34 |
| CTACT | 1357780 | 1.6690435 | 7.078636 | 36 |
| TACTG | 1227575 | 1.653415 | 7.7959566 | 37 |
| TCCAG | 1274580 | 1.5920327 | 6.932506 | 2 |
| CGGCT | 1447550 | 1.5782825 | 10.829936 | 37 |
| GCCCA | 1342530 | 1.5551059 | 8.239181 | 1 |
| GCTCA | 1203875 | 1.5037175 | 7.0833435 | 1 |
| CTCCA | 1316615 | 1.5008874 | 7.0022326 | 1 |
| TTCCC | 1531105 | 1.4993874 | 10.532534 | 7 |
| TATGT | 1007595 | 1.4634192 | 5.7718387 | 6 |
| GTTCC | 1360465 | 1.4597949 | 11.503308 | 6 |
| CGCCT | 1454335 | 1.4471714 | 10.136064 | 11 |
| TCCCG | 1452900 | 1.4457436 | 10.361562 | 8 |
| CTCGT | 1281135 | 1.3746729 | 9.068487 | 32 |
| CGGTT | 1167990 | 1.3732173 | 12.223493 | 4 |
| CCGCT | 1295820 | 1.2894373 | 7.933394 | 37 |
| CGTCC | 1274420 | 1.2681429 | 7.875044 | 34 |
| GTCCG | 1149715 | 1.2535491 | 8.399076 | 35 |
| GCTCG | 1130845 | 1.2329749 | 8.540543 | 31 |
| CTATG | 915300 | 1.232813 | 5.2305593 | 5 |
| TCGTC | 1109480 | 1.1904851 | 8.268044 | 33 |
| TTCGG | 932400 | 1.0962316 | 12.303671 | 2 |
| TCGGT | 928500 | 1.0916464 | 12.23399 | 3 |
| TCCGC | 1056840 | 1.0516344 | 7.6894264 | 36 |
| CGCTC | 990140 | 0.98526293 | 7.7811284 | 38 |
| CTTCG | 860925 | 0.9237825 | 11.244692 | 1 |